

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
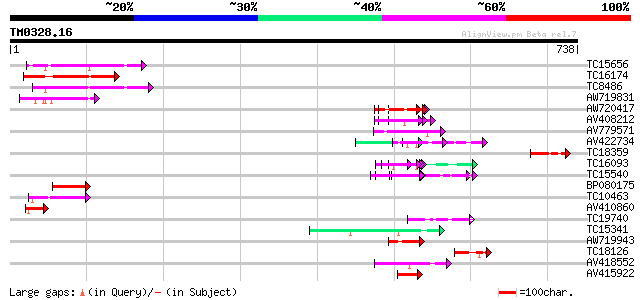
Query= TM0328.16
(738 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC15656 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Ar... 125 3e-29
TC16174 similar to UP|Q940N7 (Q940N7) AT4g05150/C17L7_70, partia... 124 7e-29
TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Ara... 123 1e-28
AW719831 74 6e-14
AW720417 62 4e-10
AV408212 60 1e-09
AV779571 53 2e-07
AV422734 52 3e-07
TC18359 52 4e-07
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 51 8e-07
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 50 1e-06
BP080175 50 1e-06
TC10463 similar to UP|Q9LRV7 (Q9LRV7) Gb|AAF04880.1, partial (19%) 49 2e-06
AV410860 49 4e-06
TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, part... 47 1e-05
TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pro... 47 1e-05
AW719943 45 4e-05
TC18126 44 7e-05
AV418552 43 2e-04
AV415922 42 3e-04
>TC15656 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;}, partial (23%)
Length = 846
Score = 125 bits (313), Expect = 3e-29
Identities = 76/166 (45%), Positives = 97/166 (57%), Gaps = 9/166 (5%)
Frame = +3
Query: 22 PPQPPPHLNYADSVDSSPRSRNTD-----SWDDPFPPTSKLRLMCSYGGHIVPRPHDKSL 76
P PP +Y DS +SSPRSR D WDD K++ MCSYGG I PR HD L
Sbjct: 69 PAYPP---SYPDSGESSPRSREIDFENPPPWDDQ---NYKVKFMCSYGGKIQPRTHDNQL 230
Query: 77 CYVGGDTRIVVVERATSLADISTRLS---KTFLDGRPFTLKYQLPNEDLDSLISVTTDED 133
YVGGDT+I+ V+R + ++LS + T KYQLP EDLD+LISVT D+D
Sbjct: 231 SYVGGDTKILAVDRNIKFPALLSKLSALCNNCDNPADLTFKYQLPGEDLDALISVTNDDD 410
Query: 134 LENMIDEYDRTTTTTASSIKPSRIRLFLF-PTKPESAHSIPPQILD 178
L++M+ EYDR +A +PSR+RLFLF P P + P + D
Sbjct: 411 LDHMMHEYDRLYRASA---RPSRMRLFLFLPENPLPSALAPAPVSD 539
>TC16174 similar to UP|Q940N7 (Q940N7) AT4g05150/C17L7_70, partial (20%)
Length = 491
Score = 124 bits (310), Expect = 7e-29
Identities = 74/126 (58%), Positives = 87/126 (68%), Gaps = 1/126 (0%)
Frame = +1
Query: 19 VAPPPQPPPHLNYADSVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCY 78
+A P Q PP L+ DSV SSPRS D+ PP ++R M S+ G I+PRPHD L Y
Sbjct: 121 MATPSQYPPELD-TDSVASSPRSDFHDA-----PP--RVRFMVSFRGKILPRPHDNQLRY 276
Query: 79 VGGDTRIVVVERATSLADISTRLSKTFLDGRP-FTLKYQLPNEDLDSLISVTTDEDLENM 137
VGGDTRIV V R+ S + +LSK L G TLKYQLPNE+LD+LISVTTDED+ENM
Sbjct: 277 VGGDTRIVAVNRSASFPALLLKLSK--LSGMTKITLKYQLPNEELDALISVTTDEDVENM 450
Query: 138 IDEYDR 143
IDEYDR
Sbjct: 451 IDEYDR 468
>TC8486 similar to GB|AAP68317.1|31711922|BT008878 At5g64430 {Arabidopsis
thaliana;}, partial (25%)
Length = 856
Score = 123 bits (308), Expect = 1e-28
Identities = 75/166 (45%), Positives = 93/166 (55%), Gaps = 8/166 (4%)
Frame = +1
Query: 30 NYADSVDSSPRSRNTD-----SWDDPFPPTS-KLRLMCSYGGHIVPRPHDKSLCYVGGDT 83
+Y DS DSSPRSR D W+D P + K + +CSYGG I PR HD L YVGGDT
Sbjct: 55 SYPDSGDSSPRSREIDFENPTPWEDQQSPHNYKAKFLCSYGGKIQPRTHDNQLSYVGGDT 234
Query: 84 RIVVVERATSLADISTRLSKTFLDGRP--FTLKYQLPNEDLDSLISVTTDEDLENMIDEY 141
+I+ V+R T ++L+ D P T KYQLP EDLD+LISVT D+DLE+++ EY
Sbjct: 235 KILAVDRNTKFPTFLSKLA-ALCDAAPQELTFKYQLPGEDLDALISVTNDDDLEHLMHEY 411
Query: 142 DRTTTTTASSIKPSRIRLFLFPTKPESAHSIPPQILDSSAKSDDWF 187
DR + KP R+RLFLF S P L D F
Sbjct: 412 DRLYRPAS---KPVRMRLFLFSAPNPGPLSQQPDPLKPQPNVDFLF 540
>AW719831
Length = 436
Score = 74.3 bits (181), Expect = 6e-14
Identities = 50/127 (39%), Positives = 67/127 (52%), Gaps = 23/127 (18%)
Frame = +3
Query: 14 PPLSSVAPPPQPPPHLNYA---DSVDSSPRSR--------NTD-------SWDDPFPP-- 53
P + + P + NY+ +S DSSPRSR N D S+D+P PP
Sbjct: 57 PKIPNSISPTMENNNNNYSSHPNSRDSSPRSRSDVIDTTTNNDTNNHHHSSFDEPPPPPP 236
Query: 54 ---TSKLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTFLDGRP 110
K++LMCSYGG I PRPHD +L YV GDT+I+ V+R L +L+ + +
Sbjct: 237 SAAACKVKLMCSYGGKIQPRPHDNNLTYVSGDTKILAVDRFIKLPAFIAKLT-SLTNSSD 413
Query: 111 FTLKYQL 117
F LKYQL
Sbjct: 414 FCLKYQL 434
>AW720417
Length = 554
Score = 61.6 bits (148), Expect = 4e-10
Identities = 35/66 (53%), Positives = 44/66 (66%), Gaps = 2/66 (3%)
Frame = +3
Query: 481 QVQQHHEQGYLLQQQFDQQQHLL--QQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQ 538
Q + H +Q QQ Q+H++ QQQ+ QQQHQQ +Q Q HQQ QH QQQQQQQQQQ
Sbjct: 18 QPRPHQQQQ---QQHQYVQRHMMPNQQQYHQQQHQQ--NQPQYHQQHQHNQQQQQQQQQQ 182
Query: 539 YIHGTQ 544
++ TQ
Sbjct: 183 WLRRTQ 200
Score = 47.8 bits (112), Expect = 6e-06
Identities = 31/64 (48%), Positives = 39/64 (60%), Gaps = 4/64 (6%)
Frame = +3
Query: 476 PQSRIQVQQHHE--QGYLL--QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQ 531
P+ Q QQ H+ Q +++ QQQ+ QQQH QQ Q HQQ QH QQQ QQQQ QQ
Sbjct: 21 PRPHQQQQQQHQYVQRHMMPNQQQYHQQQH---QQNQPQYHQQHQHNQQQ-QQQQQQQWL 188
Query: 532 QQQQ 535
++ Q
Sbjct: 189 RRTQ 200
Score = 41.6 bits (96), Expect = 5e-04
Identities = 24/55 (43%), Positives = 32/55 (57%), Gaps = 1/55 (1%)
Frame = +3
Query: 493 QQQFDQQQHLLQQQF-DQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFI 546
QQQ Q Q++ + +QQQ+ QQQHQQ Q Q+ QQ Q QQQQ Q++
Sbjct: 33 QQQQQQHQYVQRHMMPNQQQYHQQQHQQNQ---PQYHQQHQHNQQQQQQQQQQWL 188
Score = 33.9 bits (76), Expect = 0.096
Identities = 19/40 (47%), Positives = 20/40 (49%), Gaps = 1/40 (2%)
Frame = +3
Query: 511 QHQQQQHQQ-QQHQQQQHQQQQQQQQQQQYIHGTQFIHHN 549
Q QQQQHQ Q+H QQ QQQ QQ Q HN
Sbjct: 33 QQQQQQHQYVQRHMMPNQQQYHQQQHQQNQPQYHQQHQHN 152
Score = 28.9 bits (63), Expect = 3.1
Identities = 21/60 (35%), Positives = 26/60 (43%), Gaps = 2/60 (3%)
Frame = +3
Query: 516 QHQQQQHQQQQHQ--QQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHPQHAQ 573
Q + Q QQQQHQ Q+ QQQY H Q + P Y+ + QQ Q Q
Sbjct: 18 QPRPHQQQQQQHQYVQRHMMPNQQQY-HQQQHQQNQ-----PQYHQQHQHNQQQQQQQQQ 179
>AV408212
Length = 362
Score = 60.1 bits (144), Expect = 1e-09
Identities = 32/68 (47%), Positives = 40/68 (58%)
Frame = +3
Query: 476 PQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
PQ QQ +Q +QQ Q+ LQQQ QQQ QQQQ QQQQ QQQQ Q QQQ +
Sbjct: 6 PQQSPHQQQQQQQHMQMQQILLQRHAALQQQQQQQQQQQQQQQQQQSQQQQPQSQQQSRD 185
Query: 536 QQQYIHGT 543
+ ++G+
Sbjct: 186 RAHLLNGS 209
Score = 59.3 bits (142), Expect = 2e-09
Identities = 33/58 (56%), Positives = 34/58 (57%)
Frame = +3
Query: 481 QVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQ 538
Q QQ Q QQ QQ LLQ+ QQ QQQQ QQQQ QQQQ QQQQ Q QQQ
Sbjct: 3 QPQQSPHQQQQQQQHMQMQQILLQRHAALQQQQQQQQQQQQQQQQQQSQQQQPQSQQQ 176
Score = 56.6 bits (135), Expect = 1e-08
Identities = 32/66 (48%), Positives = 37/66 (55%), Gaps = 5/66 (7%)
Frame = +3
Query: 494 QQFDQQQHLLQQQFDQQQH-----QQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHH 548
QQ QQQH+ QQ Q+H QQQQ QQQQ QQQQ Q QQQQ Q QQ + +
Sbjct: 24 QQQQQQQHMQMQQILLQRHAALQQQQQQQQQQQQQQQQQQSQQQQPQSQQQSRDRAHLLN 203
Query: 549 NPANAI 554
AN +
Sbjct: 204 GSANGL 221
Score = 32.0 bits (71), Expect = 0.36
Identities = 22/61 (36%), Positives = 33/61 (54%), Gaps = 2/61 (3%)
Frame = +3
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDL--PSPDSVSSDSSLANAMSRQKPVIYQEQ 450
QQQ QQQ Q QSQ Q QQ+S L S + + +S ANA++ + +Y+E+
Sbjct: 105 QQQQQQQQQQQQQSQQQQPQSQQQSRDRAHLLNGSANGLVGNSGTANALATK---MYEER 275
Query: 451 V 451
+
Sbjct: 276 L 278
>AV779571
Length = 538
Score = 53.1 bits (126), Expect = 2e-07
Identities = 38/98 (38%), Positives = 45/98 (45%), Gaps = 4/98 (4%)
Frame = +1
Query: 474 SDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQ 533
S PQ Q Q ++Q LLQ Q QQHL Q Q Q H QQQ QQQQ Q QQQ
Sbjct: 136 SQPQQVFQNNQENQQ--LLQSQSHLQQHLQHQHSFNGQTQNHNHNQQQQQQQQQQTQQQV 309
Query: 534 QQQQQYIHG----TQFIHHNPANAIPAYYPVYPSQQQT 567
QQ + +QF+ + P SQQQ+
Sbjct: 310 VDNQQIGNAGSTMSQFVSAPQPQSSPTQAQSSLSQQQS 423
>AV422734
Length = 448
Score = 52.0 bits (123), Expect = 3e-07
Identities = 43/129 (33%), Positives = 51/129 (39%), Gaps = 41/129 (31%)
Frame = +1
Query: 451 VLIQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQH-LLQQQFDQ 509
V+ S + H+ + K Q + Q Q H+Q +LQQQ QQQH LL QQ +
Sbjct: 25 VVSVSSDCSISWHTKMEEAKALHQQHQHQQQQHQQHQQQLMLQQQQQQQQHFLLLQQLQK 204
Query: 510 QQHQQQQ--------------------------------------HQ--QQQHQQQQHQQ 529
QQ QQQQ HQ H QQQ QQ
Sbjct: 205 QQQQQQQAAAISRFPSSIDAHLRPIRPINLQQNPNPNPNNPILNLHQNPNSNHLQQQQQQ 384
Query: 530 QQQQQQQQQ 538
QQ QQQQQQ
Sbjct: 385 QQPQQQQQQ 411
Score = 50.8 bits (120), Expect = 8e-07
Identities = 35/76 (46%), Positives = 42/76 (55%), Gaps = 5/76 (6%)
Frame = +1
Query: 499 QQHLLQQQFDQQQHQQQQHQQQQHQQQQH-----QQQQQQQQQQQYIHGTQFIHHNPANA 553
QQH QQQ QQHQQQ QQQ QQQQH Q Q+QQQQQQQ ++F P++
Sbjct: 94 QQHQHQQQ-QHQQHQQQLMLQQQQQQQQHFLLLQQLQKQQQQQQQAAAISRF----PSSI 258
Query: 554 IPAYYPVYPSQQQTHP 569
P+ P Q +P
Sbjct: 259 DAHLRPIRPINLQQNP 306
Score = 48.5 bits (114), Expect = 4e-06
Identities = 39/113 (34%), Positives = 48/113 (41%), Gaps = 2/113 (1%)
Frame = +1
Query: 512 HQQQQHQQQQHQQQQHQ--QQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQQTHP 569
HQQ QHQQQQHQQ Q Q QQQQQQQQ ++ Q QQQ
Sbjct: 91 HQQHQHQQQQHQQHQQQLMLQQQQQQQQHFLLLQQL----------------QKQQQQQQ 222
Query: 570 QHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTLVPQTAAYNNPMRN 622
Q A + P+ A+ ++ N+ Q PNP P + NP N
Sbjct: 223 QAAAISRFPS-SIDAHLRPIRPINL------QQNPNPNPNNPILNLHQNPNSN 360
Score = 35.8 bits (81), Expect = 0.025
Identities = 39/128 (30%), Positives = 47/128 (36%)
Frame = +1
Query: 393 QQQAQQQLHTQTQSQSHPAQFQQRSSGGTDLPSPDSVSSDSSLANAMSRQKPVIYQEQVL 452
Q Q QQQ H Q Q Q Q QQ+ L A A+SR I
Sbjct: 97 QHQHQQQQHQQHQQQLMLQQQQQQQQHFLLLQQLQKQQQQQQQAAAISRFPSSIDAHLRP 276
Query: 453 IQSGTTRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQH 512
I+ + + NP +P LN+ Q HL QQQ QQQ
Sbjct: 277 IRPINLQQNPNPNPNNPILNL--------------------HQNPNSNHLQQQQ-QQQQP 393
Query: 513 QQQQHQQQ 520
QQQQ QQ+
Sbjct: 394 QQQQQQQK 417
>TC18359
Length = 475
Score = 51.6 bits (122), Expect = 4e-07
Identities = 26/52 (50%), Positives = 35/52 (67%)
Frame = +2
Query: 678 PPANYAYDYVDPAHAQIYYSQPLAPTMPSQYQTMTAAAMVLPEGSSAQLPSD 729
PP+NY YD+ DPA+AQ+ Y+QPLAP PSQ+ +LP+ S+ LP D
Sbjct: 2 PPSNYVYDFADPAYAQMLYTQPLAP--PSQHTNRP*C--LLPDDSAQLLPLD 145
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 50.8 bits (120), Expect = 8e-07
Identities = 26/52 (50%), Positives = 30/52 (57%)
Frame = +1
Query: 484 QHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQ 535
Q +Q + Q+ Q QH QQ QQQ QQQQ QQ QQ Q QQQ+Q QQ
Sbjct: 85 QQFQQLQIHPQKHQQHQHQFQQPIQQQQQQQQQPIQQPQQQMQQQQQEQHQQ 240
Score = 50.1 bits (118), Expect = 1e-06
Identities = 42/122 (34%), Positives = 48/122 (38%), Gaps = 7/122 (5%)
Frame = +1
Query: 494 QQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQ-----QQQQQQQQQQYIHGTQFIHH 548
QQF H + QQ QQ Q Q+HQQ QHQ QQQQQQQQQ
Sbjct: 37 QQFYPHNHSMHTLLAAQQFQQLQIHPQKHQQHQHQFQQPIQQQQQQQQQ----------- 183
Query: 549 NPANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPP--NP 606
P P QQQ QH Q+ +L M+ A S K P NP
Sbjct: 184 ------PIQQPQQQMQQQQQEQH----------QQSGDLKMRSATSSPSPKDNDSPDLNP 315
Query: 607 TT 608
+T
Sbjct: 316 ST 321
Score = 48.5 bits (114), Expect = 4e-06
Identities = 29/60 (48%), Positives = 34/60 (56%), Gaps = 5/60 (8%)
Frame = +1
Query: 485 HHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQH-----QQQQQQQQQQQY 539
+H LL Q QQ + Q+ Q QHQ QQ QQQ QQQQ QQQ QQQQQ+Q+
Sbjct: 55 NHSMHTLLAAQQFQQLQIHPQKHQQHQHQFQQPIQQQQQQQQQPIQQPQQQMQQQQQEQH 234
Score = 47.4 bits (111), Expect = 8e-06
Identities = 30/63 (47%), Positives = 32/63 (50%), Gaps = 5/63 (7%)
Frame = +1
Query: 485 HHEQGYLLQQQFDQ-----QQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQY 539
H L QQF Q Q+H Q QQ QQQQ QQQQ QQ QQ QQQQQ+Q
Sbjct: 58 HSMHTLLAAQQFQQLQIHPQKHQQHQHQFQQPIQQQQQQQQQPIQQPQQQMQQQQQEQHQ 237
Query: 540 IHG 542
G
Sbjct: 238 QSG 246
Score = 46.2 bits (108), Expect = 2e-05
Identities = 22/48 (45%), Positives = 27/48 (55%)
Frame = +1
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQ 524
Q +I Q+H + + QQ QQQ QQ Q Q Q QQ QQ+QHQQ
Sbjct: 97 QLQIHPQKHQQHQHQFQQPIQQQQQQQQQPIQQPQQQMQQQQQEQHQQ 240
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 50.4 bits (119), Expect = 1e-06
Identities = 41/112 (36%), Positives = 47/112 (41%)
Frame = +2
Query: 498 QQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAY 557
QQQ LQQQ Q QQQ QQQ Q QQ Q Q QQQQ + Q P +P
Sbjct: 20 QQQQQLQQQPPQLTQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQQQQLSQLQPQQQLPQL 199
Query: 558 YPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQTPPNPTTL 609
+ Q Q PQ Q V A QAY ++ M G Q PT +
Sbjct: 200 QQL---QHQQLPQQQQ-QMVGAGMGQAYVQGPGRSQMVSQG--QVSQGPTNI 337
Score = 49.3 bits (116), Expect = 2e-06
Identities = 31/72 (43%), Positives = 38/72 (52%), Gaps = 1/72 (1%)
Frame = +2
Query: 470 KLNVSDPQ-SRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQ 528
+L PQ +++Q QQ + + QQQ Q Q Q QQQ Q Q QQQ Q QQ Q
Sbjct: 32 QLQQQPPQLTQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQQQQLSQLQPQQQLPQLQQLQ 211
Query: 529 QQQQQQQQQQYI 540
QQ QQQQQ +
Sbjct: 212 HQQLPQQQQQMV 247
Score = 48.5 bits (114), Expect = 4e-06
Identities = 41/111 (36%), Positives = 46/111 (40%), Gaps = 4/111 (3%)
Frame = +2
Query: 495 QFDQQQHLLQQ--QFDQQQHQQQQHQQQQHQQQQHQQQQQQ--QQQQQYIHGTQFIHHNP 550
Q QQQ L QQ Q Q Q QQQQ Q QQQ Q QQQQ Q QQQ + Q
Sbjct: 14 QLQQQQQLQQQPPQLTQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQQQQLSQLQ------ 175
Query: 551 ANAIPAYYPVYPSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGESGKPQ 601
P QQ Q Q +P +Q Q M QA + G+ Q
Sbjct: 176 -----------PQQQLPQLQQLQHQQLPQQQQQMVGAGMGQAYVQGPGRSQ 295
Score = 46.6 bits (109), Expect = 1e-05
Identities = 31/65 (47%), Positives = 34/65 (51%)
Frame = +2
Query: 477 QSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQ 536
Q + Q+QQ Q LQQQ QQ +QQQ Q QQQ Q QQ Q Q Q QQQ Q
Sbjct: 20 QQQQQLQQQPPQLTQLQQQQQQQLPQVQQQQLSQMQQQQLPQLQQQQLSQLQPQQQLPQL 199
Query: 537 QQYIH 541
QQ H
Sbjct: 200 QQLQH 214
Score = 31.2 bits (69), Expect = 0.62
Identities = 27/84 (32%), Positives = 37/84 (43%), Gaps = 3/84 (3%)
Frame = +2
Query: 441 RQKPVIYQEQVLIQSGT--TRVPTHSNPVDPKLNVSDPQSRIQVQQHHEQGYLLQQQFDQ 498
+Q P + Q+Q L Q T++ P++ Q ++ Q + L QQQ Q
Sbjct: 2 QQLPQLQQQQQLQQQPPQLTQLQQQQQQQLPQVQ----QQQLSQMQQQQLPQLQQQQLSQ 169
Query: 499 QQHLLQ-QQFDQQQHQQQQHQQQQ 521
Q Q Q Q QHQQ QQQQ
Sbjct: 170 LQPQQQLPQLQQLQHQQLPQQQQQ 241
Score = 28.1 bits (61), Expect = 5.3
Identities = 23/71 (32%), Positives = 28/71 (39%), Gaps = 6/71 (8%)
Frame = +2
Query: 470 KLNVSDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQ------QQHQQQQHQ 523
+L+ Q Q+QQ QQQ Q Q L QQ QQQ Q Q + Q +
Sbjct: 110 QLSQMQQQQLPQLQQQQLSQLQPQQQLPQLQQLQHQQLPQQQQQMVGAGMGQAYVQGPGR 289
Query: 524 QQQHQQQQQQQ 534
Q Q Q Q
Sbjct: 290 SQMVSQGQVSQ 322
>BP080175
Length = 421
Score = 50.4 bits (119), Expect = 1e-06
Identities = 20/50 (40%), Positives = 35/50 (70%)
Frame = -2
Query: 56 KLRLMCSYGGHIVPRPHDKSLCYVGGDTRIVVVERATSLADISTRLSKTF 105
K++++CS+GG I+PRP D L YVGG+TRI+ + + S ++ ++S +
Sbjct: 177 KMKVLCSFGGKILPRPGDGKLRYVGGETRIISIMKDISWQELMQKMSSIY 28
>TC10463 similar to UP|Q9LRV7 (Q9LRV7) Gb|AAF04880.1, partial (19%)
Length = 326
Score = 49.3 bits (116), Expect = 2e-06
Identities = 30/86 (34%), Positives = 42/86 (47%), Gaps = 5/86 (5%)
Frame = +3
Query: 25 PPPH-----LNYADSVDSSPRSRNTDSWDDPFPPTSKLRLMCSYGGHIVPRPHDKSLCYV 79
P PH L Y +S P S P S ++ +CSYGG I+PR D L Y+
Sbjct: 45 PFPHFLHSKLGYLPQPESKPEPNMFGSSQPA--PKSSIKFLCSYGGKILPRYPDGKLRYL 218
Query: 80 GGDTRIVVVERATSLADISTRLSKTF 105
GG TR++ V+R+ + S L+ F
Sbjct: 219 GGHTRVLAVDRSIPFSG*SLSLTVNF 296
>AV410860
Length = 110
Score = 48.5 bits (114), Expect = 4e-06
Identities = 23/33 (69%), Positives = 24/33 (72%), Gaps = 3/33 (9%)
Frame = +3
Query: 21 PPP---QPPPHLNYADSVDSSPRSRNTDSWDDP 50
PPP QP P Y DS +SSPRSRNTDSWDDP
Sbjct: 12 PPPMEAQPTPPPIYPDSTNSSPRSRNTDSWDDP 110
>TC19740 similar to UP|FKH_DROME (P14734) Fork head protein, partial (4%)
Length = 622
Score = 47.0 bits (110), Expect = 1e-05
Identities = 34/90 (37%), Positives = 43/90 (47%), Gaps = 2/90 (2%)
Frame = +3
Query: 518 QQQQHQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYY--PVYPSQQQTHPQHAQVY 575
Q QQHQ QQHQ Q QQQQQ Q+ H H P P ++ P P QQ HP
Sbjct: 117 QHQQHQHQQHQHQFQQQQQPQHFH-----HQQPG---PEFHRGPPPPPPQQPHPPP---- 260
Query: 576 YVPARQAQAYNLSMQQANMGESGKPQTPPN 605
+ A + NL+ + + G G P PP+
Sbjct: 261 MMRQPSASSTNLAPEFHHPGPGGPP--PPH 344
Score = 35.0 bits (79), Expect = 0.043
Identities = 41/156 (26%), Positives = 57/156 (36%), Gaps = 17/156 (10%)
Frame = +3
Query: 494 QQFDQQQHLLQQQFDQQQHQQQQHQQQQ----HQ-------QQQHQQQQQQQQQQQYIH- 541
QQ QQH Q QF QQQ Q H QQ H+ QQ H +Q +
Sbjct: 123 QQHQHQQH--QHQFQQQQQPQHFHHQQPGPEFHRGPPPPPPQQPHPPPMMRQPSASSTNL 296
Query: 542 GTQFIHHNPANAIPAYYPVY-----PSQQQTHPQHAQVYYVPARQAQAYNLSMQQANMGE 596
+F H P P +Y V+ + + H Q V Y + + + M Q + +
Sbjct: 297 APEFHHPGPGGPPPPHYDVHSDSHGAKRMRKHTQRKAVDYT-STVVRYMQIRMWQHDSSD 473
Query: 597 SGKPQTPPNPTTLVPQTAAYNNPMRNAPLPKTEMTA 632
T L P TAA + + A P T+
Sbjct: 474 R---------TVLQPTTAAAIDMLPAAGYPDNPSTS 554
Score = 28.1 bits (61), Expect = 5.3
Identities = 25/77 (32%), Positives = 33/77 (42%), Gaps = 5/77 (6%)
Frame = +3
Query: 381 PVGYRKPPTPTPQQQAQQQLHTQTQSQSH--PAQFQQRSSGGTDLPSPDSVSSDSSLANA 438
P +R PP P PQQ + Q + S +F GG P D V SDS A
Sbjct: 204 PEFHRGPPPPPPQQPHPPPMMRQPSASSTNLAPEFHHPGPGGPPPPHYD-VHSDSHGAKR 380
Query: 439 M---SRQKPVIYQEQVL 452
M +++K V Y V+
Sbjct: 381 MRKHTQRKAVDYTSTVV 431
>TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (33%)
Length = 1002
Score = 46.6 bits (109), Expect = 1e-05
Identities = 57/201 (28%), Positives = 79/201 (38%), Gaps = 26/201 (12%)
Frame = +1
Query: 391 TPQQQAQQQL-HTQTQSQSHPAQFQQRSSGGTDLPSPDSVS--SDSSLANAMSRQ----- 442
T ++ A+ QL ++ S SH + +S TD D+ S S+ LA A+ Q
Sbjct: 418 TQKELAKLQLAQKESSSSSHSQSNEDKSPSTTDSKKADNASDASNQQLALALPNQIAPQP 597
Query: 443 KPVIYQEQVLIQSGTT--RVPTHSNPVDPKLNVSDPQSRIQVQQH--HEQGYLLQQQFDQ 498
+P Q Q + T + P + P P + +++ Q+ +Q Y Q D
Sbjct: 598 QPAAAQPQAPAPNVTQAPQQPAYYIPSTPLPGNTQTVAQLPQNQYLPPDQQYRTPQMQDM 777
Query: 499 QQHLLQ--------------QQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQ 544
+ Q QQ+ Q Q QQQ QQQQ QQQ QQ QQ Q TQ
Sbjct: 778 SRVAPQPCTNXSGXSPPPPVQQYPQYQQPQQQQQQQQQQQQWPQQLPQQVQP------TQ 939
Query: 545 FIHHNPANAIPAYYPVYPSQQ 565
P PVYP Q
Sbjct: 940 PPSMQSPQIRPPSSPVYPPYQ 1002
>AW719943
Length = 508
Score = 45.1 bits (105), Expect = 4e-05
Identities = 27/48 (56%), Positives = 29/48 (60%)
Frame = +2
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQYI 540
QQQ QQQ L Q Q QQ QQ+ QQQQ QQQQQQQQQQ +
Sbjct: 92 QQQPQQQQQLFQPQ---QQSPFQQNSLFNSQQQQFQQQQQQQQQQHQL 226
Score = 41.6 bits (96), Expect = 5e-04
Identities = 25/47 (53%), Positives = 27/47 (57%)
Frame = +2
Query: 493 QQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQQY 539
QQQ QQQ QQ F QQ Q + QQQ QQQQQQQQQQ+
Sbjct: 89 QQQQPQQQ---QQLFQPQQQSPFQQNSLFNSQQQQFQQQQQQQQQQH 220
Score = 38.5 bits (88), Expect = 0.004
Identities = 26/56 (46%), Positives = 27/56 (47%)
Frame = +2
Query: 474 SDPQSRIQVQQHHEQGYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQ 529
S PQ + Q QQ QQ F QQ QQ QQQQ QQQQ QQQQ Q
Sbjct: 77 STPQQQQQPQQQ-------QQLFQPQQQSPFQQNSLFNSQQQQFQQQQQQQQQQHQ 223
>TC18126
Length = 518
Score = 44.3 bits (103), Expect = 7e-05
Identities = 28/53 (52%), Positives = 34/53 (63%), Gaps = 4/53 (7%)
Frame = -1
Query: 579 ARQAQAYNLSMQQANMGESGKPQTPPNPTTL----VPQTAAYNNPMRNAPLPK 627
ARQAQAYNLS Q AN+GES TT+ P ++AY N +RNAPLP+
Sbjct: 134 ARQAQAYNLSAQHANVGESA--------TTIS*SAAPSSSAY-NLLRNAPLPQ 3
Score = 34.7 bits (78), Expect = 0.056
Identities = 17/27 (62%), Positives = 20/27 (73%)
Frame = -3
Query: 431 SDSSLANAMSRQKPVIYQEQVLIQSGT 457
S+ L A+S KPVIYQ+QV IQSGT
Sbjct: 360 SNKCLLTAVSYPKPVIYQDQVQIQSGT 280
Score = 28.5 bits (62), Expect = 4.0
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -1
Query: 611 PQTAAYNNPMRNAPLPK 627
P +A YNN +RN PLP+
Sbjct: 233 PSSAVYNNSLRNVPLPQ 183
>AV418552
Length = 386
Score = 43.1 bits (100), Expect = 2e-04
Identities = 42/114 (36%), Positives = 49/114 (42%), Gaps = 14/114 (12%)
Frame = +2
Query: 475 DPQSRIQ--VQQHHEQGYLLQQQFDQQQHL-LQQQFDQQQHQQQQHQ----------QQQ 521
+P S+ Q +QQH + Q Q HL LQQQ QQQ Q Q Q QQQ
Sbjct: 41 EPSSQEQQLMQQHQQMLSYFQPQQQLLPHLQLQQQLQQQQLAQLQQQLSQLQLTLLQQQQ 220
Query: 522 HQQQQHQQQQQQQQQQQYIHGTQFIHHNPANAIPAYYPVYPSQQ-QTHPQHAQV 574
Q Q QQ Q QQQ + TQ H + P QQ Q PQH Q+
Sbjct: 221 LTQLQQQQLAQLQQQLAQLQLTQHQH------TQLQQQLLPQQQFQQLPQHQQM 364
Score = 39.3 bits (90), Expect = 0.002
Identities = 31/68 (45%), Positives = 35/68 (50%), Gaps = 5/68 (7%)
Frame = +2
Query: 478 SRIQVQQHHEQGYLLQQQFD---QQQHL--LQQQFDQQQHQQQQHQQQQHQQQQHQQQQQ 532
+++Q Q Q LLQQQ QQQ L LQQQ Q Q Q QH Q Q Q QQ QQ
Sbjct: 164 AQLQQQLSQLQLTLLQQQQLTQLQQQQLAQLQQQLAQLQLTQHQHTQLQQQLLPQQQFQQ 343
Query: 533 QQQQQQYI 540
Q QQ +
Sbjct: 344 LPQHQQMV 367
>AV415922
Length = 422
Score = 42.4 bits (98), Expect = 3e-04
Identities = 20/32 (62%), Positives = 21/32 (65%)
Frame = +3
Query: 506 QFDQQQHQQQQHQQQQHQQQQHQQQQQQQQQQ 537
QF QHQQ+ QQQQ QQQQ QQQQQ Q
Sbjct: 153 QFQGNQHQQR*QQQQQQQQQQQQQQQQPSNSQ 248
Score = 38.5 bits (88), Expect = 0.004
Identities = 20/32 (62%), Positives = 22/32 (68%)
Frame = +3
Query: 513 QQQQHQQQQHQQQQHQQQQQQQQQQQYIHGTQ 544
Q Q +Q QQ QQQ QQQQQQQQQQQ +Q
Sbjct: 153 QFQGNQHQQR*QQQQQQQQQQQQQQQQPSNSQ 248
Score = 36.6 bits (83), Expect = 0.015
Identities = 21/41 (51%), Positives = 23/41 (55%)
Frame = +3
Query: 489 GYLLQQQFDQQQHLLQQQFDQQQHQQQQHQQQQHQQQQHQQ 529
G+ QF QH QQ QQQ QQQQ QQQQ QQ + Q
Sbjct: 135 GFRYPIQFQGNQH---QQR*QQQQQQQQQQQQQQQQPSNSQ 248
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.127 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,060,155
Number of Sequences: 28460
Number of extensions: 287059
Number of successful extensions: 10619
Number of sequences better than 10.0: 668
Number of HSP's better than 10.0 without gapping: 4272
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7992
length of query: 738
length of database: 4,897,600
effective HSP length: 97
effective length of query: 641
effective length of database: 2,136,980
effective search space: 1369804180
effective search space used: 1369804180
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0328.16


