

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
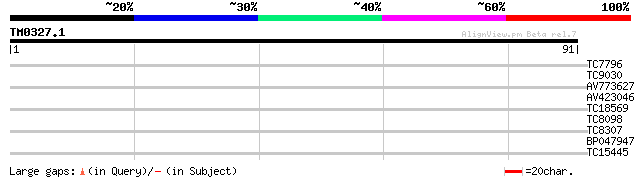
Query= TM0327.1
(91 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC7796 homologue to gb|AF036495.1|AF036495 Hamamelis virginiana ... 28 0.21
TC9030 similar to GB|AAM63479.1|21554372|AY086477 phospholipase-... 26 1.4
AV773627 25 2.3
AV423046 25 3.1
TC18569 weakly similar to UP|Q9LWT2 (Q9LWT2) ESTs AU082419(E6174... 24 4.0
TC8098 similar to UP|Q94FT5 (Q94FT5) Pectate lyase (Fragment), p... 23 6.8
TC8307 weakly similar to UP|Q9SP47 (Q9SP47) Homeodomain-leucine ... 23 6.8
BP047947 23 6.8
TC15445 similar to UP|Q8S937 (Q8S937) BLE1 protein, partial (80%) 23 8.9
>TC7796 homologue to gb|AF036495.1|AF036495 Hamamelis virginiana large
subunit 26S ribosomal RNA gene, partial sequence,
complete
Length = 6006
Score = 28.5 bits (62), Expect = 0.21
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = -2
Query: 33 GGSTNPVILRESCWRESTFQALSAKGTPFDP 63
GG+T P+ R + E T Q + +G P DP
Sbjct: 905 GGTTRPIKARSASPAEGTSQPVHTRGGPIDP 813
>TC9030 similar to GB|AAM63479.1|21554372|AY086477 phospholipase-like
protein {Arabidopsis thaliana;}, partial (53%)
Length = 596
Score = 25.8 bits (55), Expect = 1.4
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = -1
Query: 3 *ILLVDLVLMQLRVDGVLIRLRDTRMHCVFGGSTN 37
*++ V+LV ++ VL+RLRD HC + N
Sbjct: 134 *VVSVELVRSEVAEGVVLVRLRDWVHHCCYPAPKN 30
>AV773627
Length = 465
Score = 25.0 bits (53), Expect = 2.3
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = +2
Query: 35 STNPVILRESCWRESTFQALSAKGTPFDPAVY 66
+TNPV+L +S F ALS++ T F ++
Sbjct: 17 TTNPVVLSSILHLQSLFSALSSQKTWFHVVLF 112
>AV423046
Length = 427
Score = 24.6 bits (52), Expect = 3.1
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = -1
Query: 9 LVLMQLRVDGVLIRLRDTRMHCVFGGS 35
L+ Q + G+ RLR T++ C+FG S
Sbjct: 415 LLCYQHLLSGLCSRLRRTQVFCLFGSS 335
>TC18569 weakly similar to UP|Q9LWT2 (Q9LWT2) ESTs AU082419(E61744), partial
(95%)
Length = 538
Score = 24.3 bits (51), Expect = 4.0
Identities = 17/55 (30%), Positives = 26/55 (46%)
Frame = +1
Query: 31 VFGGSTNPVILRESCWRESTFQALSAKGTPFDPAVYNDPSIISQRLPIVKQMTQN 85
+FGG+T I W E+ + S T P V N IS++ IVK +++
Sbjct: 157 IFGGTTPGTITNTGWWEETDKKFQSWPRTAGPPVVMNP---ISRQNFIVKSRSES 312
>TC8098 similar to UP|Q94FT5 (Q94FT5) Pectate lyase (Fragment), partial
(56%)
Length = 1349
Score = 23.5 bits (49), Expect = 6.8
Identities = 9/26 (34%), Positives = 14/26 (53%)
Frame = +1
Query: 32 FGGSTNPVILRESCWRESTFQALSAK 57
F +T+P +R SCW T ++K
Sbjct: 124 FPTTTSPTTMRRSCWATVTLTLETSK 201
>TC8307 weakly similar to UP|Q9SP47 (Q9SP47) Homeodomain-leucine zipper
protein 57, partial (51%)
Length = 1629
Score = 23.5 bits (49), Expect = 6.8
Identities = 9/25 (36%), Positives = 14/25 (56%)
Frame = -2
Query: 33 GGSTNPVILRESCWRESTFQALSAK 57
GGSTNP + +C++ S S +
Sbjct: 989 GGSTNPFVSLSTCFKFSLASTFSVR 915
>BP047947
Length = 541
Score = 23.5 bits (49), Expect = 6.8
Identities = 13/41 (31%), Positives = 19/41 (45%)
Frame = +3
Query: 5 LLVDLVLMQLRVDGVLIRLRDTRMHCVFGGSTNPVILRESC 45
+L L L+ L +LIR + F + NPVI+ C
Sbjct: 153 ILSSLHLLNLSKSVLLIRATAAALRSYFPFAINPVIMPSQC 275
>TC15445 similar to UP|Q8S937 (Q8S937) BLE1 protein, partial (80%)
Length = 580
Score = 23.1 bits (48), Expect = 8.9
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = +3
Query: 46 WRESTFQALSAKGTPFDPAVYNDPS 70
W E T +A + +PF P PS
Sbjct: 60 WEEDTAEARRTRASPFTPLNAGTPS 134
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.331 0.144 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,395,829
Number of Sequences: 28460
Number of extensions: 14707
Number of successful extensions: 111
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 111
length of query: 91
length of database: 4,897,600
effective HSP length: 67
effective length of query: 24
effective length of database: 2,990,780
effective search space: 71778720
effective search space used: 71778720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0327.1


