

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
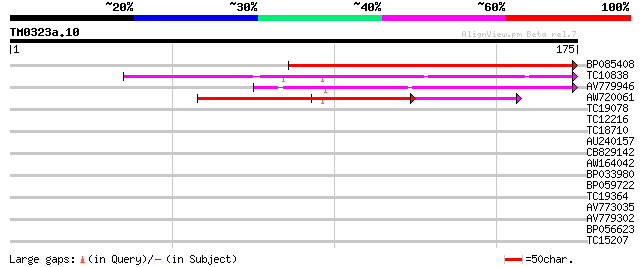
Query= TM0323a.10
(175 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP085408 201 8e-53
TC10838 weakly similar to PIR|T04232|T04232 pathogenesis-related... 100 1e-22
AV779946 92 4e-20
AW720061 77 2e-15
TC19078 similar to UP|Q85FM6 (Q85FM6) Photosystem II protein M (... 28 0.88
TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Ar... 27 1.5
TC18710 26 4.4
AU240157 26 4.4
CB829142 25 5.7
AW164042 25 5.7
BP033980 25 7.4
BP059722 25 7.4
TC19364 weakly similar to UP|Q8H146 (Q8H146) Putatative tyrosine... 25 7.4
AV773035 25 7.4
AV779302 25 9.7
BP056623 25 9.7
TC15207 similar to UP|Q9FGP9 (Q9FGP9) Gb|AAF31706.1 (AT5g22790/K... 25 9.7
>BP085408
Length = 350
Score = 201 bits (510), Expect = 8e-53
Identities = 89/89 (100%), Positives = 89/89 (100%)
Frame = -1
Query: 87 NKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGC 146
NKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGC
Sbjct: 350 NKYGSNQDLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGC 171
Query: 147 AQLTCVVDKLILTICFYDPPGNINGESPY 175
AQLTCVVDKLILTICFYDPPGNINGESPY
Sbjct: 170 AQLTCVVDKLILTICFYDPPGNINGESPY 84
>TC10838 weakly similar to PIR|T04232|T04232 pathogenesis-related protein
homolog F14M19.60 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (78%)
Length = 914
Score = 100 bits (249), Expect = 1e-22
Identities = 55/144 (38%), Positives = 81/144 (56%), Gaps = 4/144 (2%)
Frame = +3
Query: 36 EAKEFLDAHNWARAEVGVEPLEWSEALAKDAGLFARYQRDRLGCKLAN-FSHNKYGSNQD 94
E+ EFL HN RA PL W L A +A ++ CK+ + F N + ++
Sbjct: 363 ESLEFLFRHNMVRAAKWESPLMWDFQLQSYARWWAGQRKP--DCKVEHSFPENDFKLGEN 536
Query: 95 L---GGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTC 151
+ G WTP+ AV+ W +E+++Y+Y TN+C C HYTQ+VW+ +K+VGCA++ C
Sbjct: 537 IFWGSGSAWTPTDAVKAWADEEKYYTYATNTCEEGQM-CGHYTQIVWKNTKRVGCARVVC 713
Query: 152 VVDKLILTICFYDPPGNINGESPY 175
+ +T C YDP GN GE PY
Sbjct: 714 DDGDVFMT-CNYDPVGNYVGERPY 782
>AV779946
Length = 533
Score = 92.4 bits (228), Expect = 4e-20
Identities = 48/101 (47%), Positives = 60/101 (58%), Gaps = 1/101 (0%)
Frame = -3
Query: 76 RLGCKLANFSHNKYGSNQDLG-GMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYT 134
R C L + S+ YG N G G GW PS AV WV E+++Y+Y NSC A C HYT
Sbjct: 501 RNDCALEH-SNGPYGENIFWGSGTGWKPSPAVDAWVEERQWYNYWHNSC-ANGEMCGHYT 328
Query: 135 QVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGNINGESPY 175
Q+VW +++VGCA +TC + C YDPPGN GE PY
Sbjct: 327 QIVWGDTRKVGCASVTCSGGQGTFMTCNYDPPGNYYGERPY 205
>AW720061
Length = 304
Score = 76.6 bits (187), Expect = 2e-15
Identities = 36/68 (52%), Positives = 46/68 (66%), Gaps = 1/68 (1%)
Frame = +1
Query: 59 SEALAKDAGLFARYQRDRLGCKLANFSHNKYGSNQDL-GGMGWTPSVAVQDWVNEKEFYS 117
SE L KDA LF RYQR++ GC AN + ++YG NQ + G + P VAV +WV EK+FY
Sbjct: 4 SEKLGKDASLFVRYQRNKFGCGFANLTDSEYGGNQIMAGSVSVAPRVAVAEWVKEKDFYI 183
Query: 118 YRTNSCVA 125
+ NSCVA
Sbjct: 184 HANNSCVA 207
Score = 44.7 bits (104), Expect = 9e-06
Identities = 26/65 (40%), Positives = 33/65 (50%)
Frame = +3
Query: 94 DLGGMGWTPSVAVQDWVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVV 153
D GG G + V E E + + V C YTQVVWR SK+VGC+Q +CV
Sbjct: 108 DHGGFGVGGAARGGGGVGEGEGFLHPRE*LVRGGT*CGVYTQVVWRDSKEVGCSQASCVK 287
Query: 154 DKLIL 158
+K L
Sbjct: 288 EKASL 302
>TC19078 similar to UP|Q85FM6 (Q85FM6) Photosystem II protein M (Fragment),
partial (37%)
Length = 603
Score = 28.1 bits (61), Expect = 0.88
Identities = 9/28 (32%), Positives = 15/28 (53%)
Frame = +1
Query: 96 GGMGWTPSVAVQDWVNEKEFYSYRTNSC 123
GG W P ++DW+ ++ R N+C
Sbjct: 463 GGFVWGPVQVLRDWLGASAYHFARVNAC 546
>TC12216 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;}, partial (30%)
Length = 570
Score = 27.3 bits (59), Expect = 1.5
Identities = 16/67 (23%), Positives = 27/67 (39%)
Frame = +2
Query: 109 WVNEKEFYSYRTNSCVAPSFPCWHYTQVVWRKSKQVGCAQLTCVVDKLILTICFYDPPGN 168
W N++ F YR +SC A + WRK + + +V I+ Y
Sbjct: 110 WSNDQGFLCYRCDSCKAGVLAS---LKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNKR 280
Query: 169 INGESPY 175
++ + PY
Sbjct: 281 MDNDEPY 301
>TC18710
Length = 843
Score = 25.8 bits (55), Expect = 4.4
Identities = 14/33 (42%), Positives = 18/33 (54%), Gaps = 4/33 (12%)
Frame = +1
Query: 113 KEFYSYRTNSC----VAPSFPCWHYTQVVWRKS 141
K YS +T+S A SFPC+ T W+KS
Sbjct: 460 KWLYSRKTDSR*FTDCAGSFPCYKQTGCTWQKS 558
>AU240157
Length = 300
Score = 25.8 bits (55), Expect = 4.4
Identities = 9/20 (45%), Positives = 14/20 (70%)
Frame = -3
Query: 124 VAPSFPCWHYTQVVWRKSKQ 143
V+ SF C+HY +WRK ++
Sbjct: 166 VSNSFNCFHYHLRIWRKRRR 107
>CB829142
Length = 534
Score = 25.4 bits (54), Expect = 5.7
Identities = 9/25 (36%), Positives = 12/25 (48%)
Frame = +1
Query: 99 GWTPSVAVQDWVNEKEFYSYRTNSC 123
GW P V +NE EF+ + C
Sbjct: 202 GWDPETVVDYRINEDEFHKFSLLDC 276
>AW164042
Length = 404
Score = 25.4 bits (54), Expect = 5.7
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +1
Query: 144 VGCAQLTCVVDKLILTICFYDP 165
VGC TCVVD TI ++P
Sbjct: 178 VGCPLSTCVVDTFATTIGTHEP 243
>BP033980
Length = 615
Score = 25.0 bits (53), Expect = 7.4
Identities = 8/20 (40%), Positives = 12/20 (60%)
Frame = +1
Query: 113 KEFYSYRTNSCVAPSFPCWH 132
K +Y + +C +P FP WH
Sbjct: 313 KIYYGWC*QTCGSPGFPVWH 372
>BP059722
Length = 473
Score = 25.0 bits (53), Expect = 7.4
Identities = 8/27 (29%), Positives = 15/27 (54%)
Frame = -1
Query: 129 PCWHYTQVVWRKSKQVGCAQLTCVVDK 155
PC+H+ +VW +G A++ + K
Sbjct: 260 PCFHFPVIVWSCHPNLGLAEVLIQIKK 180
>TC19364 weakly similar to UP|Q8H146 (Q8H146) Putatative tyrosine
aminotransferase, partial (14%)
Length = 387
Score = 25.0 bits (53), Expect = 7.4
Identities = 10/30 (33%), Positives = 15/30 (49%)
Frame = -3
Query: 45 NWARAEVGVEPLEWSEALAKDAGLFARYQR 74
NW R + V+P E E L++ RY +
Sbjct: 304 NWLRISLAVDPSELEEGLSRIKAFSLRYAK 215
>AV773035
Length = 459
Score = 25.0 bits (53), Expect = 7.4
Identities = 14/46 (30%), Positives = 24/46 (51%), Gaps = 5/46 (10%)
Frame = -1
Query: 120 TNSCVAPSF-----PCWHYTQVVWRKSKQVGCAQLTCVVDKLILTI 160
+ +CV SF PC+ + ++ R S ++ C LTC + K L +
Sbjct: 459 SKACVYRSFWWWL*PCFIFGHILSRDSCKLRCKILTCPLKKQSLLL 322
>AV779302
Length = 509
Score = 24.6 bits (52), Expect = 9.7
Identities = 9/23 (39%), Positives = 15/23 (65%)
Frame = +2
Query: 100 WTPSVAVQDWVNEKEFYSYRTNS 122
+ P+ + WV ++ F SYRT+S
Sbjct: 425 YVPTPSQYPWVGQQPFNSYRTSS 493
>BP056623
Length = 451
Score = 24.6 bits (52), Expect = 9.7
Identities = 10/27 (37%), Positives = 14/27 (51%)
Frame = -3
Query: 97 GMGWTPSVAVQDWVNEKEFYSYRTNSC 123
G GW P V + D + + RT+SC
Sbjct: 404 GPGWVPPVLLLD*MGDMSVSPLRTSSC 324
>TC15207 similar to UP|Q9FGP9 (Q9FGP9) Gb|AAF31706.1 (AT5g22790/K8E10_2),
partial (55%)
Length = 1195
Score = 24.6 bits (52), Expect = 9.7
Identities = 8/27 (29%), Positives = 15/27 (54%)
Frame = +3
Query: 129 PCWHYTQVVWRKSKQVGCAQLTCVVDK 155
PC+H+ +VW +G A++ + K
Sbjct: 759 PCFHFPVIVWSFHPNLGIAKVLIQIKK 839
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,718,715
Number of Sequences: 28460
Number of extensions: 56593
Number of successful extensions: 266
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 261
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 261
length of query: 175
length of database: 4,897,600
effective HSP length: 84
effective length of query: 91
effective length of database: 2,506,960
effective search space: 228133360
effective search space used: 228133360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0323a.10


