

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
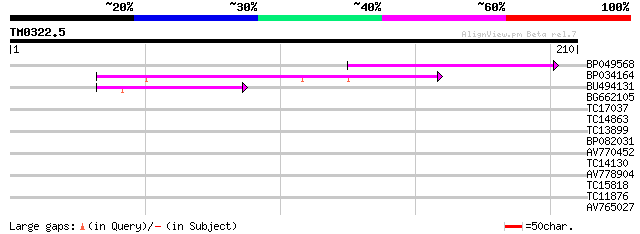
Query= TM0322.5
(210 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP049568 49 6e-07
BP034164 40 2e-04
BU494131 39 7e-04
BG662105 37 0.002
TC17037 similar to UP|Q8LRK7 (Q8LRK7) HDA2 (Fragment), partial (... 29 0.69
TC14863 28 0.90
TC13899 similar to UP|O82188 (O82188) At2g19930 protein, partial... 28 1.5
BP082031 27 2.0
AV770452 25 7.6
TC14130 similar to UP|ATPG_PEA (P28552) ATP synthase gamma chain... 25 7.6
AV778904 25 9.9
TC15818 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyge... 25 9.9
TC11876 25 9.9
AV765027 25 9.9
>BP049568
Length = 598
Score = 48.9 bits (115), Expect = 6e-07
Identities = 23/78 (29%), Positives = 46/78 (58%)
Frame = -1
Query: 126 DLAKDEHLDPINVKIAELREALESIITEQKYLKARDNRHRYTNESTKTRVIFYTVLEYIL 185
++AK E ++ + +++ +L ++ +I+ KYLK R+ R +E T TRV ++++ +
Sbjct: 592 EVAKKEKIEGVELELRKLEGSVTAILENLKYLKGREAEMRSVSEKTNTRVATFSIMSLGI 413
Query: 186 LAVTSILQVVYIRRLFSK 203
V S LQ+ ++R F K
Sbjct: 412 CIVVSALQLWRLKRYFQK 359
>BP034164
Length = 515
Score = 40.4 bits (93), Expect = 2e-04
Identities = 33/134 (24%), Positives = 61/134 (44%), Gaps = 6/134 (4%)
Frame = +1
Query: 33 ECVSEHAQSGDSVTGNF-VVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFK 91
+C++E +S G + +V ++ + D I V++P GN+ H+ F F
Sbjct: 112 KCITEDIKSNAMSVGKYSIVNPNEGYPIPDTHKITVKVSSPYGNSYHNGDQVDSGDFAFT 291
Query: 92 APQNGIYKFCFHNPMS----TPETVSFYIHVGHIPSE-HDLAKDEHLDPINVKIAELREA 146
A ++G Y CF P S + T+ F G + +AK ++ + ++ +L +
Sbjct: 292 AAEDGDYTACFWIPDSRAGVSIVTIEFDWRTGVAAKDWSKVAKKGQVEVMEFELKKLYDT 471
Query: 147 LESIITEQKYLKAR 160
+ SI E YL+ R
Sbjct: 472 VASIHEEMFYLRER 513
>BU494131
Length = 381
Score = 38.9 bits (89), Expect = 7e-04
Identities = 19/57 (33%), Positives = 28/57 (48%), Gaps = 1/57 (1%)
Frame = +2
Query: 33 ECVSEHAQ-SGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKF 88
EC S + GD+V +FVV+ D W G+D V P G +H + + +KF
Sbjct: 209 ECFSHDVKYEGDTVHISFVVIKADSPWHYGDEGVDLVVKGPAGEQIHDFRDKTSEKF 379
>BG662105
Length = 342
Score = 37.4 bits (85), Expect = 0.002
Identities = 27/102 (26%), Positives = 47/102 (45%), Gaps = 6/102 (5%)
Frame = +3
Query: 63 PGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQNGIYKFCF-----HNPMSTPETVSFYIH 117
P + VT+P GN +H + + + F A ++G Y CF H + +S
Sbjct: 36 PTVSAKVTSPYGNNLHHNENVTQGQSAFTATESGNYMACFWIDGKHQEAAGGTILSLEWK 215
Query: 118 VGHIPSEHD-LAKDEHLDPINVKIAELREALESIITEQKYLK 158
G + D +AK E L+ + +++ +L +E+I YLK
Sbjct: 216 TGISAKDWDSVAKKEKLEGVXLELRKLEGXVEAIRDNLIYLK 341
>TC17037 similar to UP|Q8LRK7 (Q8LRK7) HDA2 (Fragment), partial (39%)
Length = 774
Score = 28.9 bits (63), Expect = 0.69
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = -1
Query: 177 FYTVLEYILLAVTSILQVVYIRRLFS 202
FY+ Y +L T ILQ++Y++ FS
Sbjct: 618 FYSYFMYHILTFTDILQLIYVKHXFS 541
>TC14863
Length = 1803
Score = 28.5 bits (62), Expect = 0.90
Identities = 13/29 (44%), Positives = 15/29 (50%)
Frame = -3
Query: 89 EFKAPQNGIYKFCFHNPMSTPETVSFYIH 117
E K P G+ K+C NP TP F IH
Sbjct: 727 EMKKPSTGLIKYCARNP--TPTVHGFRIH 647
>TC13899 similar to UP|O82188 (O82188) At2g19930 protein, partial (10%)
Length = 626
Score = 27.7 bits (60), Expect = 1.5
Identities = 16/76 (21%), Positives = 28/76 (36%)
Frame = +2
Query: 34 CVSEHAQSGDSVTGNFVVMDHDIFWSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAP 93
CV + + + G+F D D++W S HP + ++ + +
Sbjct: 269 CVGPRSIADEIAGGDF---DGDLYWVSKHPQVSISILVQKSSCTYQ-------------- 397
Query: 94 QNGIYKFCFHNPMSTP 109
I +C H P S P
Sbjct: 398 ---I*SYCIHTPFSVP 436
>BP082031
Length = 208
Score = 27.3 bits (59), Expect = 2.0
Identities = 12/35 (34%), Positives = 20/35 (56%)
Frame = +1
Query: 60 SDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQ 94
SD P + +T+T PG N V + +G+ +K+ Q
Sbjct: 94 SDQPKVSYTLTRPGSNQV-QINLATGNTLSYKSVQ 195
>AV770452
Length = 237
Score = 25.4 bits (54), Expect = 7.6
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +1
Query: 134 DPINVKIAELREALESIITEQKYLKARDNRHRY 166
DP+++KI + L I +Y++A HRY
Sbjct: 52 DPVSIKILQAYSYL*GCINISRYVEASHIAHRY 150
>TC14130 similar to UP|ATPG_PEA (P28552) ATP synthase gamma chain,
chloroplast precursor , partial (94%)
Length = 1527
Score = 25.4 bits (54), Expect = 7.6
Identities = 12/47 (25%), Positives = 24/47 (50%)
Frame = +3
Query: 113 SFYIHVGHIPSEHDLAKDEHLDPINVKIAELREALESIITEQKYLKA 159
S +++ +PS++ + P+ + ELR +ES+ QK +A
Sbjct: 174 SIHLNPFQLPSQNSPPSRISVSPVQCGLRELRTRIESVKNTQKITEA 314
>AV778904
Length = 541
Score = 25.0 bits (53), Expect = 9.9
Identities = 9/21 (42%), Positives = 12/21 (56%)
Frame = -1
Query: 3 WFSKICTLVIFFLGTLIYNVF 23
W +C FF+GT+I VF
Sbjct: 394 WLFYLCHYATFFIGTIICGVF 332
>TC15818 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (37%)
Length = 618
Score = 25.0 bits (53), Expect = 9.9
Identities = 9/27 (33%), Positives = 17/27 (62%)
Frame = +1
Query: 15 LGTLIYNVFSLSISLNDVECVSEHAQS 41
+G +YN++ L +S V+C S H ++
Sbjct: 487 IGIFLYNMYGLKLSCFRVQCNSLHGKN 567
>TC11876
Length = 541
Score = 25.0 bits (53), Expect = 9.9
Identities = 21/81 (25%), Positives = 34/81 (41%), Gaps = 1/81 (1%)
Frame = -1
Query: 34 CVSEHAQSGDSVTGNFVVMDHDIF-WSSDHPGIDFTVTNPGGNAVHSLKGTSGDKFEFKA 92
C E+ G + N+ +MD + W+ H + V + G N + +
Sbjct: 229 CACENDLGGSTNDINWYIMD*QLLNWNIIH---QWAVMH*GNNLC*IIDSSH-------V 80
Query: 93 PQNGIYKFCFHNPMSTPETVS 113
PQ+ I+KFC +S P T S
Sbjct: 79 PQHTIFKFCKSLSLSQPSTSS 17
>AV765027
Length = 491
Score = 25.0 bits (53), Expect = 9.9
Identities = 11/34 (32%), Positives = 17/34 (49%)
Frame = -2
Query: 62 HPGIDFTVTNPGGNAVHSLKGTSGDKFEFKAPQN 95
HPG+ V GG+ SL D + F+ P++
Sbjct: 445 HPGVTKIVVRDGGSVELSLDHLELDMWRFRLPES 344
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,659,928
Number of Sequences: 28460
Number of extensions: 48598
Number of successful extensions: 264
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 264
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 264
length of query: 210
length of database: 4,897,600
effective HSP length: 86
effective length of query: 124
effective length of database: 2,450,040
effective search space: 303804960
effective search space used: 303804960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0322.5


