

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0322.2
(856 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
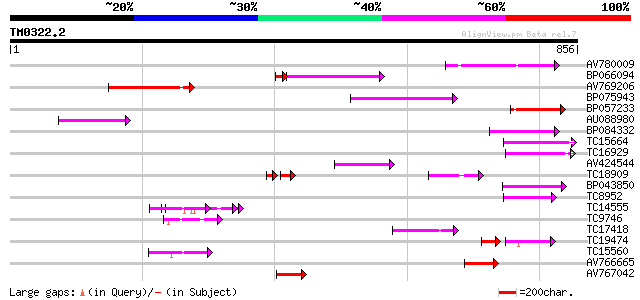
Score E
Sequences producing significant alignments: (bits) Value
AV780009 137 9e-33
BP066094 112 5e-27
AV769206 107 6e-24
BP075943 93 2e-19
BP057233 86 2e-17
AU088980 82 5e-16
BP084332 76 2e-14
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 76 3e-14
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 74 1e-13
AV424544 65 3e-11
TC18909 63 2e-10
BP043850 61 9e-10
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 57 9e-09
TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precu... 52 3e-07
TC9746 49 3e-06
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 49 3e-06
TC19474 weakly similar to UP|Q9FJA1 (Q9FJA1) Similarity to retro... 35 4e-06
TC15560 similar to SP|Q03173|NDPP_MOUSE NPC derived proline rich... 47 1e-05
AV766665 47 1e-05
AV767042 47 1e-05
>AV780009
Length = 529
Score = 137 bits (344), Expect = 9e-33
Identities = 78/173 (45%), Positives = 103/173 (59%), Gaps = 2/173 (1%)
Frame = -1
Query: 659 AVKRILRYLKGTVDFGLHLKPCSLHQPLSLIAFCDADWAADPDDHRSTSGSLVFLGPNLV 718
A R+LRY+KG GL S PL L A+ D+DWA PD RS +G +FLG +L+
Sbjct: 526 AATRVLRYVKGAPAQGLFF---SADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLI 356
Query: 719 SWCSKKQSLVARSSTEAEYRNLANTVAELLWVQSLLQ--ELHIPHKTPLILCDNMSTVAL 776
SW +KKQ+ V+RSS+EAEYR LA TV E+ W+ L Q +L++P PL CDN S + +
Sbjct: 355 SWRTKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKLNVPLPVPL-FCDNQSALHI 179
Query: 777 THNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRF 829
HNP H RTKH+ELD VR K+ + + + + Q DI TK L F
Sbjct: 178 AHNPTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKPCPHLCF 20
>BP066094
Length = 532
Score = 112 bits (279), Expect(2) = 5e-27
Identities = 59/148 (39%), Positives = 90/148 (59%), Gaps = 1/148 (0%)
Frame = +2
Query: 419 KPITVRIILSLAVSKGWHIHQLDVNNAFLNGKLEEEVYMVQPPGFE-AKDRSLVCRLNKA 477
K +R+++S +V+ +HQ++V +AFLNG + EEVY+ QPPG E K+ + +L K+
Sbjct: 89 KTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKKS 268
Query: 478 LYGLKQAPRAWFERLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLP 537
LYGLKQAPRAW+ERL + LL+ + +LF + V +YVDDII +
Sbjct: 269 LYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANPS 448
Query: 538 VLQHLIAKLNAEFSLKQLGTLDYFLGVQ 565
+ + + AEF ++ +G L YFLG+Q
Sbjct: 449 LCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
Score = 26.9 bits (58), Expect(2) = 5e-27
Identities = 11/18 (61%), Positives = 13/18 (72%)
Frame = +3
Query: 402 YSQVEGFDYKETFSPVVK 419
YSQ G DY ETF+PV +
Sbjct: 39 YSQQ*GIDYTETFAPVAR 92
>AV769206
Length = 536
Score = 107 bits (268), Expect = 6e-24
Identities = 60/130 (46%), Positives = 82/130 (62%), Gaps = 1/130 (0%)
Frame = -3
Query: 150 YLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLDPTGRLYISKDVLFNEFRFPYKELFGCA 209
+LRPYN+ KLSLHSKECVF+GYS HK YKCLD +GR+YISKDVLF+E RFPY LF
Sbjct: 534 FLRPYNSTKLSLHSKECVFIGYSINHKGYKCLDQSGRIYISKDVLFHEHRFPYPSLFPDE 355
Query: 210 SSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHS-PNTTLPESSSPASQSSTPILPEPTSP 268
++ S+ S+IP+ S L+S S + TLP S +P ++ + L E P
Sbjct: 354 TTSSTSAEFFPLSTIPIVSHTTPPALASSGFSSLGSATLPSSFNPTAKLT---LTEW*QP 184
Query: 269 VPQSSPAPIS 278
+ + + P++
Sbjct: 183 LTELTAVPLT 154
>BP075943
Length = 547
Score = 92.8 bits (229), Expect = 2e-19
Identities = 53/162 (32%), Positives = 89/162 (54%), Gaps = 1/162 (0%)
Frame = -3
Query: 515 SSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEFSLKQLGTLDYFLGVQVQHLHDKSL 574
SSG +L+YVDD+++ GN + +Q + + L+ +F +K LG YFLG+++ +
Sbjct: 542 SSGSFTALLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIAR-STSGI 366
Query: 575 LLTQSKYIKDLL-ERASMAYAKGISTPMISNQKLTKHGANYMADPSLYRSIVGALQYVTI 633
+L Q KY L+ + + + ++Q L + + D YR IVG L Y+
Sbjct: 365 VLNQRKYALQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNT 186
Query: 634 TRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGL 675
TRP+I+++VN++ QFLS P + H + L+YL G+ GL
Sbjct: 185 TRPDITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGL 60
>BP057233
Length = 473
Score = 85.9 bits (211), Expect = 2e-17
Identities = 40/83 (48%), Positives = 58/83 (69%)
Frame = -2
Query: 756 ELHIPHKTPLILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLIIQHVPSLDQ 815
+L +P K P++ CDN+S AL NPVLH R+KH+E+D+ ++R++VL +++ +VP+ DQ
Sbjct: 472 KLPMPRK-PILWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQ 296
Query: 816 RDDILTKALSPLRFEDLRTKLNV 838
D LTK LS RF LR KL V
Sbjct: 295 IADCLTKPLSHTRFSQLRDKLGV 227
>AU088980
Length = 360
Score = 81.6 bits (200), Expect = 5e-16
Identities = 41/109 (37%), Positives = 60/109 (54%)
Frame = +2
Query: 74 NGIVERKHRHIVEMGLALLAHAHISLEYWDHAFLTVVYLINRLPTVVLDNESPFTKLFQV 133
N IVERKHRHI+ + AL+ H+++ W A YLIN LP+ +L + P+ L
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 134 SFDVKFLRIFGCACYPYLRPYNAHKLSLHSKECVFLGYSTTHKSYKCLD 182
S + L++FG CY +N KL +++CVFLG+ T Y +D
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMD 328
>BP084332
Length = 368
Score = 76.3 bits (186), Expect = 2e-14
Identities = 39/105 (37%), Positives = 63/105 (59%)
Frame = +2
Query: 725 QSLVARSSTEAEYRNLANTVAELLWVQSLLQELHIPHKTPLILCDNMSTVALTHNPVLHT 784
QS +A S+ EAEY + A ++LW++ L++ I I CDN + ++L+ NP+LH+
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS 211
Query: 785 RTKHMELDIFFVREKVLNKSLIIQHVPSLDQRDDILTKALSPLRF 829
R KH+E+ F+R+ V L+++ V + Q DI TK L+ RF
Sbjct: 212 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 346
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 75.9 bits (185), Expect = 3e-14
Identities = 44/111 (39%), Positives = 64/111 (57%), Gaps = 1/111 (0%)
Frame = +1
Query: 746 ELLWVQSLLQELHIPHKTP-LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKS 804
EL W+ +LQ+L +P +P L+ CD+ S + N V H RTKH+++D VREK+ K
Sbjct: 4 EL*WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKL 183
Query: 805 LIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVTDKFTLAQPP*VCGGIL 855
+ + S+DQ DILTK L F L +KL V + ++ A CGG+L
Sbjct: 184 FHLLPISSVDQTADILTKPLESGPFSHLVSKLGVLNIYSSA-----CGGVL 321
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 73.6 bits (179), Expect = 1e-13
Identities = 41/106 (38%), Positives = 58/106 (54%), Gaps = 1/106 (0%)
Frame = +1
Query: 749 WVQSLLQELHIPHKTP-LILCDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKSLII 807
W+ LLQ+L +P + P L+ CDN S + NPV H RTKH+E+D VRE++ + +
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 808 QHVPSLDQRDDILTKALSPLRFEDLRTKLNVTDKFTLAQPP*VCGG 853
+ S + DI TKALSP F + KL + + P CGG
Sbjct: 184 LPISSSEPLADIYTKALSPQNFHQICAKLGL---INICSP--ACGG 306
>AV424544
Length = 276
Score = 65.5 bits (158), Expect = 3e-11
Identities = 37/91 (40%), Positives = 53/91 (57%)
Frame = +3
Query: 491 RLKAALLKFGFTASCCNPSLFTLYSSGCSVYVLVYVDDIIITGNSLPVLQHLIAKLNAEF 550
+L + L G+ S + SLFT + +LVYVDD+I+ GN L +Q + KL+ +F
Sbjct: 6 KLSSYLHILGYIQSAHDHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQF 185
Query: 551 SLKQLGTLDYFLGVQVQHLHDKSLLLTQSKY 581
+K LGTL YFLG++V L L+Q KY
Sbjct: 186 RIKDLGTLKYFLGLEVAR-SSCGLFLSQRKY 275
>TC18909
Length = 621
Score = 62.8 bits (151), Expect = 2e-10
Identities = 40/85 (47%), Positives = 50/85 (58%), Gaps = 2/85 (2%)
Frame = +2
Query: 633 ITRPEISYSVNKVCQFLSQPLEEHWAAVKRILRYL-KGTVDFGLHLKPCSLHQPLSLIAF 691
+T P+IS+SVNKVC FLS+ LEEH A K IL K G ++ + AF
Sbjct: 323 LTSPQISFSVNKVC*FLSESLEEHGTAGKCILNVT*KAL*TMGFFIQ-------TNFPAF 481
Query: 692 CDADWAADPDDHRSTSGSLVF-LGP 715
DADWA++ DD RST +VF LGP
Sbjct: 482 SDADWASNVDDRRSTFWEIVFTLGP 556
Score = 41.2 bits (95), Expect(2) = 1e-07
Identities = 19/23 (82%), Positives = 21/23 (90%)
Frame = +2
Query: 409 DYKETFSPVVKPITVRIILSLAV 431
DY ET SPVVKP+TVRI+LSLAV
Sbjct: 62 DYTETVSPVVKPVTVRILLSLAV 130
Score = 32.3 bits (72), Expect(2) = 1e-07
Identities = 13/17 (76%), Positives = 16/17 (93%)
Frame = +1
Query: 388 GSVNRYKARLVAKGYSQ 404
GS+N+YKARLVAKG+ Q
Sbjct: 1 GSINKYKARLVAKGFHQ 51
>BP043850
Length = 515
Score = 60.8 bits (146), Expect = 9e-10
Identities = 35/97 (36%), Positives = 54/97 (55%), Gaps = 1/97 (1%)
Frame = -1
Query: 745 AELLWVQSLLQELHIPHKTPLIL-CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNK 803
+E++W++ LL EL P L DN S + + NPV H T+H+E+D VRE +
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 804 SLIIQHVPSLDQRDDILTKALSPLRFEDLRTKLNVTD 840
+ + HV + Q DILTK+L+ R L +KL + +
Sbjct: 335 VITLPHVSTSVQIADILTKSLTRQRHNFLVSKLMLVE 225
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 57.4 bits (137), Expect = 9e-09
Identities = 33/81 (40%), Positives = 46/81 (56%), Gaps = 1/81 (1%)
Frame = +3
Query: 746 ELLWVQSLLQELHIPHKTPLIL-CDNMSTVALTHNPVLHTRTKHMELDIFFVREKVLNKS 804
E LW+ L +L I +++ CDN S + L N V H RT+++E+D V KVL
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 805 LIIQHVPSLDQRDDILTKALS 825
L + HVPS DQ D+ TK +S
Sbjct: 201 LHLLHVPSSDQVADVFTKTIS 263
>TC14555 similar to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (43%)
Length = 1038
Score = 52.4 bits (124), Expect = 3e-07
Identities = 44/125 (35%), Positives = 55/125 (43%), Gaps = 11/125 (8%)
Frame = +2
Query: 230 PQLTTLSSPAPHSPNTTLP---ESSSPASQSSTPIL-----PEPTSPVPQSSPAPIS--- 278
P + +SPAP +P T P +SS P +QSS P + P T P QSSP P+S
Sbjct: 191 PSNSPTTSPAPPTPTTPQPPAAQSSPPPAQSSPPPVQSSPPPASTPPPAQSSPPPVSSPP 370
Query: 279 PALSSPAPQSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTSAKL 338
P S+P P +ST +S P P T P L P S A P AK+
Sbjct: 371 PVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATP----PPALTPVPATSPA-PAPAKV 535
Query: 339 ALKDP 343
K P
Sbjct: 536 KSKSP 550
Score = 46.6 bits (109), Expect = 2e-05
Identities = 36/108 (33%), Positives = 51/108 (46%), Gaps = 16/108 (14%)
Frame = +2
Query: 212 PQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQ 271
P SS V S P+ S P S+P P SP P ++P ++TP P +PVP
Sbjct: 332 PAQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAATP--PPALTPVPA 505
Query: 272 SSPAPI-------------SPALS---SPAPQSSSTDSSSSVSPIVNP 303
+SPAP +PAL+ S +P + S SS+SP ++P
Sbjct: 506 TSPAPAPAKVKSKSPAPAPAPALAPVVSLSPSEAPGPSLSSLSPALSP 649
Score = 43.9 bits (102), Expect = 1e-04
Identities = 35/123 (28%), Positives = 53/123 (42%), Gaps = 6/123 (4%)
Frame = +2
Query: 236 SSPAPHSPNTTLPESSSPASQSSTPILPEPTSPVPQSS-----PAPISPALSSPAPQSSS 290
SSP P S T P SSP SS P + P P S+ P P SP ++P P +++
Sbjct: 302 SSPPPAS--TPPPAQSSPPPVSSPPPVQSTPPPAPASTPPPASPPPFSPPPATPPPPAAT 475
Query: 291 TDSSSSVSPIVNPHNTHSMV-TRAKDGIVKPRLVPTLLLSQAEPTSAKLALKDPKWYSAM 349
+ + P +P + V +++ P L P + LS +E L+ P A
Sbjct: 476 PPPALTPVPATSPAPAPAKVKSKSPAPAPAPALAPVVSLSPSEAPGPSLSSLSPALSPAA 655
Query: 350 QDE 352
D+
Sbjct: 656 SDD 664
>TC9746
Length = 637
Score = 49.3 bits (116), Expect = 3e-06
Identities = 36/95 (37%), Positives = 50/95 (51%), Gaps = 6/95 (6%)
Frame = +2
Query: 233 TTLSSP----APHSPNTTLPESSSPASQSSTPILPEPTSPVPQS--SPAPISPALSSPAP 286
TTL+SP +PHS + L SS+ QSS P P P SP P+ SP+P SP +SP+P
Sbjct: 338 TTLTSPPHSSSPHSTTSPLASSSASPCQSS-P*SPSPPSPSPKHPHSPSPRSPPPASPSP 514
Query: 287 QSSSTDSSSSVSPIVNPHNTHSMVTRAKDGIVKPR 321
SSS P+ P +T + + ++ PR
Sbjct: 515 PY----FSSSAPPVSCPSSTATSLCQSSAAFSSPR 607
Score = 32.7 bits (73), Expect = 0.25
Identities = 33/90 (36%), Positives = 42/90 (46%), Gaps = 10/90 (11%)
Frame = +2
Query: 210 SSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPAS------QSSTPILP 263
+SP SS S+ P S P + SP+P P++ P S PAS SS P +
Sbjct: 383 TSPLASS-----SASPCQSSP*SPSPPSPSPKHPHSPSPRSPPPASPSPPYFSSSAPPVS 547
Query: 264 EPTSPVP---QSSPAPISP-ALSSPAPQSS 289
P+S QSS A SP +SP P SS
Sbjct: 548 CPSSTATSLCQSSAAFSSPRGSTSPCPPSS 637
Score = 27.7 bits (60), Expect = 8.0
Identities = 19/73 (26%), Positives = 32/73 (43%)
Frame = +2
Query: 208 CASSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSPNTTLPESSSPASQSSTPILPEPTS 267
C SSP S S P P+ +SP+P +++ P S P+S +++
Sbjct: 416 CQSSP*SPSPPSPSPKHPHSPSPRSPPPASPSPPYFSSSAPPVSCPSSTATSLCQSSAAF 595
Query: 268 PVPQSSPAPISPA 280
P+ S +P P+
Sbjct: 596 SSPRGSTSPCPPS 634
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 49.3 bits (116), Expect = 3e-06
Identities = 28/101 (27%), Positives = 55/101 (53%), Gaps = 1/101 (0%)
Frame = -1
Query: 578 QSKYIKDLLERASMAYAKGISTPMISNQ-KLTKHGANYMADPSLYRSIVGALQYVTITRP 636
Q Y+ D+L++ M +K IST + + + + D + Y+S++G+++Y+ R
Sbjct: 739 QKIYVDDILKKFKMTNSKYISTTIGGKEIEAGRRNGGKRVDSTYYKSLIGSVRYLNTVRS 560
Query: 637 EISYSVNKVCQFLSQPLEEHWAAVKRILRYLKGTVDFGLHL 677
+I V +F+ +P + H +R LRY+KGT+ G+ +
Sbjct: 559 DIVCGVGLRSRFM-EP*DCH*QGAQRSLRYIKGTLKDGIFM 440
>TC19474 weakly similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement pol
polyprotein, partial (3%)
Length = 517
Score = 34.7 bits (78), Expect(2) = 4e-06
Identities = 16/29 (55%), Positives = 21/29 (72%)
Frame = -1
Query: 713 LGPNLVSWCSKKQSLVARSSTEAEYRNLA 741
L NL+SW SK+ +VARS EAE+R +A
Sbjct: 517 LSDNLISWKSKETIIVARSRAEAEFRVMA 431
Score = 33.5 bits (75), Expect(2) = 4e-06
Identities = 23/81 (28%), Positives = 37/81 (45%), Gaps = 6/81 (7%)
Frame = -2
Query: 749 WVQSLLQELHIPHKTPLI----LCDNMSTVALTHNPVL--HTRTKHMELDIFFVREKVLN 802
W+Q L Q +P L+ + A H P + R KH+E+D F+ ++ L
Sbjct: 435 WLQPLXQSSLLPKPRDLVDVSLITHT*DNKAALHMPKI*FSMRAKHIEIDFHFL*DRRLY 256
Query: 803 KSLIIQHVPSLDQRDDILTKA 823
+ + V S DQ D+ TK+
Sbjct: 255 LDISTRFVNSNDQLTDVFTKS 193
>TC15560 similar to SP|Q03173|NDPP_MOUSE NPC derived proline rich protein 1
(NDPP-1). {Mus musculus;}, partial (8%)
Length = 1134
Score = 47.4 bits (111), Expect = 1e-05
Identities = 34/101 (33%), Positives = 45/101 (43%), Gaps = 4/101 (3%)
Frame = +3
Query: 210 SSPQLSSVGSVHSSIPLFSFPQLTTLSSPAPHSP----NTTLPESSSPASQSSTPILPEP 265
+S SS S SS P S P + P P +P + PE SS S+ P P+P
Sbjct: 126 ASASSSSSSSPSSS*PSPSLPPPSPPPLPPPQAPPLPSHLPSPEPSSTMPPSTPP--PQP 299
Query: 266 TSPVPQSSPAPISPALSSPAPQSSSTDSSSSVSPIVNPHNT 306
T P P P P S+P +S+ +S SP PH+T
Sbjct: 300 TKPCPLPKSTPSQPPSSAPHRVPTSSSLASPTSPSSGPHST 422
Score = 38.1 bits (87), Expect = 0.006
Identities = 34/99 (34%), Positives = 44/99 (44%), Gaps = 4/99 (4%)
Frame = +3
Query: 241 HSPNTTLPESSSPASQSSTPILPEPTSPVPQSS--PAPISPALSS--PAPQSSSTDSSSS 296
H P TT +SS +S S + P P+ P P P P +P L S P+P+ SST S+
Sbjct: 105 HYPLTTTASASSSSSSSPSSS*PSPSLPPPSPPPLPPPQAPPLPSHLPSPEPSSTMPPST 284
Query: 297 VSPIVNPHNTHSMVTRAKDGIVKPRLVPTLLLSQAEPTS 335
P T ++ P VPT S A PTS
Sbjct: 285 PPPQPTKPCPLPKSTPSQPPSSAPHRVPT-SSSLASPTS 398
>AV766665
Length = 601
Score = 47.4 bits (111), Expect = 1e-05
Identities = 25/52 (48%), Positives = 33/52 (63%)
Frame = +3
Query: 687 SLIAFCDADWAADPDDHRSTSGSLVFLGPNLVSWCSKKQSLVARSSTEAEYR 738
S+ F AD+ D ST G +FL NLV+W SK+Q+++ARSS EAE R
Sbjct: 438 SMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTWRSKQQNIIARSSGEAELR 593
>AV767042
Length = 444
Score = 47.4 bits (111), Expect = 1e-05
Identities = 20/44 (45%), Positives = 30/44 (67%)
Frame = -3
Query: 404 QVEGFDYKETFSPVVKPITVRIILSLAVSKGWHIHQLDVNNAFL 447
Q G D +TFSPVVKP +R + ++A+S+ W I QL+V N ++
Sbjct: 349 QTAGVDCNKTFSPVVKPAPIRTVFTIALSRSWPIPQLNVQNLWI 218
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.135 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,611,197
Number of Sequences: 28460
Number of extensions: 290636
Number of successful extensions: 6164
Number of sequences better than 10.0: 845
Number of HSP's better than 10.0 without gapping: 3641
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4908
length of query: 856
length of database: 4,897,600
effective HSP length: 98
effective length of query: 758
effective length of database: 2,108,520
effective search space: 1598258160
effective search space used: 1598258160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0322.2


