

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
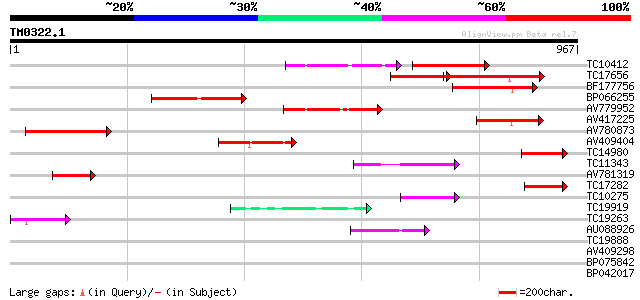
Query= TM0322.1
(967 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC10412 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATP... 199 5e-88
TC17656 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transport... 253 1e-67
BF177756 173 1e-43
BP066255 144 9e-35
AV779952 125 2e-29
AV417225 115 4e-26
AV780873 112 3e-25
AV409404 106 2e-23
TC14980 similar to UP|ACA9_ARATH (Q9LU41) Potential calcium-tran... 91 1e-18
TC11343 similar to PIR|B96660|B96660 protein F2K11.18 [imported]... 86 3e-17
AV781319 64 9e-11
TC17282 similar to UP|Q8L8A0 (Q8L8A0) Type IIB calcium ATPase MC... 63 2e-10
TC10275 similar to UP|Q7Y051 (Q7Y051) Paa2 P-type ATPase, partia... 58 8e-09
TC19919 homologue to UP|Q7Y068 (Q7Y068) Plasma membrane H+-ATPas... 49 5e-06
TC19263 similar to UP|Q8L8A0 (Q8L8A0) Type IIB calcium ATPase MC... 48 6e-06
AU088926 48 8e-06
TC19888 weakly similar to PIR|T08551|T08551 Ca2+-transporting AT... 40 0.001
AV409298 40 0.002
BP075842 39 0.004
BP042017 36 0.033
>TC10412 homologue to UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase,
partial (33%)
Length = 1061
Score = 199 bits (506), Expect(2) = 5e-88
Identities = 95/131 (72%), Positives = 113/131 (85%)
Frame = +3
Query: 687 GHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCV 746
G VVAVTGDGTNDAPAL EADIGL+MGI GTEVAKES+D++ILDDNF+++ TV +WGR V
Sbjct: 648 GEVVAVTGDGTNDAPALHEADIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSV 827
Query: 747 YNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPT 806
Y NIQKF+QFQLTVNV AL++NF +A +G PLT VQLLWVN+IMDTLGALALATE P
Sbjct: 828 YINIQKFVQFQLTVNVVALIVNFTSACLTGTAPLTAVQLLWVNMIMDTLGALALATEPPK 1007
Query: 807 KELMQKKPIGR 817
+LM++ P+GR
Sbjct: 1008DDLMKRSPVGR 1040
Score = 143 bits (361), Expect(2) = 5e-88
Identities = 86/199 (43%), Positives = 117/199 (58%), Gaps = 1/199 (0%)
Frame = +2
Query: 471 DMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGT 530
D +Q ++ VE FNS KKR VAV + H KGA+E+VLA C ++SNG
Sbjct: 2 DFQGERQACNLVKVEPFNSTKKRMSVAVELPGGG-LRAHCKGASEIVLAACDKVLNSNGE 178
Query: 531 QKSLDEER-SKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTLLGI 589
LDEE + + I A+ +LR + AYME+ E G E P G T +G+
Sbjct: 179 VVPLDEESINHLNSTINQFASEALRTLCLAYMEL-ENGFSAEDPIPLS-----GFTCIGV 340
Query: 590 VGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVVEGVE 649
VG+KDP RP VK++V C+ AG+ ++M+TGDNI TAKAIA ECGIL G+ +EG E
Sbjct: 341 VGIKDPVRPGVKESVAVCRSAGITVRMVTGDNINTAKAIARECGIL---TDDGIAIEGPE 511
Query: 650 FRNYTEEERMEKVDKIRVM 668
FR + EE +E + KI+V+
Sbjct: 512 FREKSLEELLELIPKIQVL 568
>TC17656 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transporting
ATPase-like protein {Arabidopsis thaliana;} , partial
(25%)
Length = 821
Score = 253 bits (646), Expect = 1e-67
Identities = 124/180 (68%), Positives = 150/180 (82%), Gaps = 7/180 (3%)
Frame = +2
Query: 740 LRWGRCVYNNIQKFIQFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALA 799
+RWGR VY NIQKFIQFQLTVNVAALVIN +AA+SSGDVPL VQLLWVNLIMDTLGALA
Sbjct: 278 VRWGRSVYANIQKFIQFQLTVNVAALVINVVAAISSGDVPLNAVQLLWVNLIMDTLGALA 457
Query: 800 LATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSI-------F 852
LATE PT LM + P+GR EPLIT IMWRNLL QA+YQ++VLLV F G+SI F
Sbjct: 458 LATEPPTDHLMDRSPVGRREPLITNIMWRNLLIQAMYQVSVLLVLNFRGRSILGLTHDKF 637
Query: 853 NVSKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVE 912
+ + +VKNTLIFN FVLCQ+FNEFN+R ++ N+F+G+ KN+LF+GIVG+T+VLQ+++VE
Sbjct: 638 DHAVKVKNTLIFNAFVLCQIFNEFNARKPDEFNIFKGVTKNYLFMGIVGLTVVLQIVIVE 817
Score = 130 bits (328), Expect = 8e-31
Identities = 70/107 (65%), Positives = 80/107 (74%), Gaps = 3/107 (2%)
Frame = +3
Query: 650 FRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIG 709
FR ++ ER E D I VM RSSP DKLL+VQ L++KGHVVAVTGDGTNDAPAL EADIG
Sbjct: 9 FRAMSDAERDEIADAISVMGRSSPNDKLLLVQALRRKGHVVAVTGDGTNDAPALHEADIG 188
Query: 710 LSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRC---VYNNIQKF 753
L+MGI GTEVAKESSDI+ILDDNF SV V G ++ N+ F
Sbjct: 189 LAMGIAGTEVAKESSDIIILDDNFASVVKVSDGGGLYMQIFRNLSSF 329
>BF177756
Length = 455
Score = 173 bits (438), Expect = 1e-43
Identities = 85/150 (56%), Positives = 109/150 (72%), Gaps = 4/150 (2%)
Frame = +1
Query: 755 QFQLTVNVAALVINFIAAVSSGDVPLTTVQLLWVNLIMDTLGALALATERPTKELMQKKP 814
QFQLTVNV ALVINF +A +G PLT VQLLWVNLIMDTLGALALATE P L+++ P
Sbjct: 1 QFQLTVNVVALVINFFSACITGSAPLTAVQLLWVNLIMDTLGALALATEPPNDGLLKRPP 180
Query: 815 IGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNVS----KEVKNTLIFNTFVLC 870
+ R ITK MWRN++ Q++YQ+ VL++ F GK + +S V NTLIFN+FV C
Sbjct: 181 VARGASFITKAMWRNIIGQSIYQLIVLVILTFDGKRLLRLSGSDATRVLNTLIFNSFVFC 360
Query: 871 QVFNEFNSRSMEKLNVFEGILKNHLFLGIV 900
QVFNE NSR +EK+N+F G+ + +F+ I+
Sbjct: 361 QVFNEINSRDIEKINIFRGMFDSWIFVAII 450
>BP066255
Length = 540
Score = 144 bits (362), Expect = 9e-35
Identities = 77/162 (47%), Positives = 110/162 (67%)
Frame = +2
Query: 243 AQMLVTAVGANTAWGQMMSSISGDNSERTPLQARLDKLTSSIGKIGLAVAFLVLLVLLIR 302
++ML+T VG T WG++M++++ + TPLQ +L+ + + IGKIGL A +V +L++
Sbjct: 32 SKMLITTVGMRTQWGKLMATLTEGGDDETPLQVKLNGVATIIGKIGLFFA-IVTFAVLVQ 208
Query: 303 YFTGNTEDENGNKEYKGSKTDINDVCNAVVSIVAAAVTIVVVAIPEGLPLAVTLTLAYSM 362
+ ++ + G D ++ A AVTIVVVA+PEGLPLAVTL+LA++M
Sbjct: 209 GLVSHKLQQDSFWSWTG------DDALEMLEYFAIAVTIVVVAVPEGLPLAVTLSLAFAM 370
Query: 363 KRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTK 404
K+MM D+A+VR L+ACETMGSAT IC+DKTGTLT N M V K
Sbjct: 371 KKMMNDKALVRHLAACETMGSATTICSDKTGTLTTNHMTVVK 496
>AV779952
Length = 557
Score = 125 bits (315), Expect = 2e-29
Identities = 73/169 (43%), Positives = 102/169 (60%), Gaps = 1/169 (0%)
Frame = +2
Query: 468 LGMDMDELKQKHKVLHVETFNSEKKRSGVAVRKETNNTVHVHWKGAAEMVLAMCSNYIDS 527
LG D + +Q K++ VE FNS KK GV ++ H KGA+E+VLA C +DS
Sbjct: 65 LGGDFIKGRQVVKLVKVEPFNSTKKSMGVVLQLPDGG-FRAHCKGASEIVLAACDKVVDS 241
Query: 528 NGTQKSLDEER-SKIEKIIQGMAASSLRCIAFAYMEISEGGDYIEKGKPRQVLREDGLTL 586
NG LDE+ ++++ I+ A +LR + YM+I D G P + G T
Sbjct: 242 NGEVVPLDEDSINQVKDTIEKFADDALRTLCLTYMDIQ---DEF*VGSP---IPTSGYTC 403
Query: 587 LGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGIL 635
+GIVG+KDP RP V+++V C+ AG+ ++M+TGDNI TAKAIA ECGIL
Sbjct: 404 IGIVGIKDPVRPGVRESVAICRSAGITVRMVTGDNIDTAKAIARECGIL 550
>AV417225
Length = 359
Score = 115 bits (287), Expect = 4e-26
Identities = 56/119 (47%), Positives = 78/119 (65%), Gaps = 4/119 (3%)
Frame = +3
Query: 796 GALALATERPTKELMQKKPIGRTEPLITKIMWRNLLAQALYQIAVLLVFQFYGKSIFNV- 854
GALALATE P LM++ P+GR ITK MWRN+ Q++YQ+ VL V F GK + +
Sbjct: 3 GALALATEPPNDGLMERLPVGRRASFITKPMWRNIFGQSIYQLIVLGVLNFDGKRLLGLT 182
Query: 855 ---SKEVKNTLIFNTFVLCQVFNEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLM 910
+ V NT+IFNTFV CQVFNE NSR +EK+N+F G+ + +F ++ T+ Q ++
Sbjct: 183 GSDATAVLNTVIFNTFVFCQVFNEINSREIEKINIFRGMFDSGIFFTVIFSTVAFQAII 359
>AV780873
Length = 487
Score = 112 bits (280), Expect = 3e-25
Identities = 56/146 (38%), Positives = 91/146 (61%)
Frame = +3
Query: 28 RLSNMVKDKNLEAYSEFGGVEGVADVLGTIPAKGILGSDDDTAARRELFGTNTYVRPPPK 87
+L+++ ++ + A ++GGV ++++L T KG++ D D R+ FG+N Y R +
Sbjct: 39 QLASISREHDTAALQQYGGVAVISNLLKTNLEKGVIDDDADLLKRKNAFGSNNYPRKKGR 218
Query: 88 IFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSIFLAVFLVVVVSALSN 147
F F+ +A D T++IL+ A SL GIK G EGWY+GGSI AV LV+ V+A+S+
Sbjct: 219 PFWMFLWDACKDLTLIILMVAAAASLALGIKSEGIEEGWYDGGSIAFAVLLVIFVTAVSD 398
Query: 148 FRQDRQFDKLSKISNDIKVEVVRNGR 173
++Q QF L++ +I +EV+R GR
Sbjct: 399 YKQSLQFRDLNEEKRNIHLEVIRGGR 476
>AV409404
Length = 421
Score = 106 bits (265), Expect = 2e-23
Identities = 63/142 (44%), Positives = 88/142 (61%), Gaps = 8/142 (5%)
Frame = +1
Query: 356 LTLAYSMKRMMADQAMVRKLSACETMGSATVICTDKTGTLTLNQMRVTKFWL-------- 407
L+LA++MK++M D+A+VR LSACETMGSA ICTDKTGTLT N M V K W+
Sbjct: 4 LSLAFAMKKLMNDRALVRHLSACETMGSANCICTDKTGTLTTNHMVVDKIWICEKTTEIK 183
Query: 408 GLENVVENFSNAMAPTVLELFHQGVGLNTTGSVYKPSAESEPEISGSPTEKAMLLWAVSD 467
G E+ VE + ++ V+ +F Q + NT+ V E + I G+PTE A+L + +
Sbjct: 184 GNES-VEKLKSEISEEVISIFLQAIFQNTSSEVVN-DKEGKKAILGTPTESALLEFGLLS 357
Query: 468 LGMDMDELKQKHKVLHVETFNS 489
G D D ++ +K+L VE FNS
Sbjct: 358 -GGDFDAQRRDYKILKVEPFNS 420
>TC14980 similar to UP|ACA9_ARATH (Q9LU41) Potential calcium-transporting
ATPase 9, plasma membrane-type (Ca(2+)-ATPase isoform
9) , partial (8%)
Length = 594
Score = 90.5 bits (223), Expect = 1e-18
Identities = 40/78 (51%), Positives = 54/78 (68%)
Frame = +1
Query: 874 NEFNSRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIG 933
NEFN+R E++NVF G+ KN LF+GIV +T +LQ++++E L KF DT RLNW W +
Sbjct: 1 NEFNARKPEEMNVFRGVTKNRLFMGIVVMTFILQIIIIEFLGKFTDTVRLNWTLWLASLL 180
Query: 934 IAAVSWPIAWLTKLTPVP 951
I +SWP+A K PVP
Sbjct: 181 IGLISWPLAIAGKFIPVP 234
>TC11343 similar to PIR|B96660|B96660 protein F2K11.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(21%)
Length = 909
Score = 85.9 bits (211), Expect = 3e-17
Identities = 53/182 (29%), Positives = 89/182 (48%)
Frame = +2
Query: 586 LLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGILDLNDAGGVVV 645
++G++ + DP +PN ++ V + M+TGDN TA +IA + GI
Sbjct: 161 VIGVLAVSDPLKPNAREVVSILNSMNIRSIMVTGDNWGTANSIARQAGI----------- 307
Query: 646 EGVEFRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKE 705
V+A + P K V+ L+ G+ VA+ GDG ND+PAL
Sbjct: 308 -------------------ETVIAEAQPQTKATKVKELQTSGYTVAMVGDGINDSPALVA 430
Query: 706 ADIGLSMGIQGTEVAKESSDIVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAAL 765
AD+G+++G GT++A E+SDIV++ N + L + ++ I+ + L N+ A+
Sbjct: 431 ADVGIAIG-AGTDIAIEASDIVLMRSNLEDIIIALDLAKKTFSRIRLNYLWALGYNLLAI 607
Query: 766 VI 767
I
Sbjct: 608 PI 613
>AV781319
Length = 524
Score = 64.3 bits (155), Expect = 9e-11
Identities = 32/73 (43%), Positives = 46/73 (62%)
Frame = +1
Query: 73 RELFGTNTYVRPPPKIFLHFVLEALNDTTILILLGCAGLSLGFGIKEHGPGEGWYEGGSI 132
++ G+N Y R + FL F+ +A D T++IL A SL GIK G EGWY+GGSI
Sbjct: 106 KKCIGSNNYPRKKGRSFLMFIWDAGKDLTLIILTVAAIASLALGIKSEGIKEGWYDGGSI 285
Query: 133 FLAVFLVVVVSAL 145
+AV LV++V+ +
Sbjct: 286 AVAVILVILVTGV 324
>TC17282 similar to UP|Q8L8A0 (Q8L8A0) Type IIB calcium ATPase MCA5, partial
(7%)
Length = 498
Score = 63.2 bits (152), Expect = 2e-10
Identities = 29/73 (39%), Positives = 46/73 (62%)
Frame = +1
Query: 878 SRSMEKLNVFEGILKNHLFLGIVGITIVLQVLMVELLRKFADTERLNWEQWGICIGIAAV 937
SR MEK+NV GIL+N++F+ ++ T + Q+++VE + FA+T L QW C+ + +
Sbjct: 1 SREMEKINVLNGILENYVFVAVLSATALFQIIIVEYMGTFANTTPLTLVQWFFCLVVGFL 180
Query: 938 SWPIAWLTKLTPV 950
PIA K+ PV
Sbjct: 181 GMPIAAGIKMIPV 219
>TC10275 similar to UP|Q7Y051 (Q7Y051) Paa2 P-type ATPase, partial (16%)
Length = 724
Score = 57.8 bits (138), Expect = 8e-09
Identities = 31/102 (30%), Positives = 54/102 (52%), Gaps = 1/102 (0%)
Frame = +1
Query: 667 VMARSSPMDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTE-VAKESSD 725
V A +P K + LK GH VA+ GDG NDAPAL AD+G+++ + E A +++
Sbjct: 7 VKASLAPQQKSEFISSLKAAGHHVAMVGDGINDAPALAVADVGIALQNEAQENAASDAAS 186
Query: 726 IVILDDNFNSVATVLRWGRCVYNNIQKFIQFQLTVNVAALVI 767
I++L + + V + + + + + + + NV A+ I
Sbjct: 187 IILLGNKISQVVDAIDLAQSTMAKVYQNLSWAVAYNVIAIPI 312
>TC19919 homologue to UP|Q7Y068 (Q7Y068) Plasma membrane H+-ATPase, partial
(21%)
Length = 586
Score = 48.5 bits (114), Expect = 5e-06
Identities = 56/241 (23%), Positives = 96/241 (39%)
Frame = +1
Query: 377 ACETMGSATVICTDKTGTLTLNQMRVTKFWLGLENVVENFSNAMAPTVLELFHQGVGLNT 436
A E M V+C+DKTGTLTLN++ V + N++E F+ + + L
Sbjct: 1 AIEEMAGMDVLCSDKTGTLTLNKLSVDR------NLIEVFTKGIEKEHVILL-------- 138
Query: 437 TGSVYKPSAESEPEISGSPTEKAMLLWAVSDLGMDMDELKQKHKVLHVETFNSEKKRSGV 496
+ E++ I A+ + D E + + +H FN KR+ +
Sbjct: 139 --AARASRTENQDAIDA----------AIVGMLADPKEARAGVREVHFLPFNPVDKRTAL 282
Query: 497 AVRKETNNTVHVHWKGAAEMVLAMCSNYIDSNGTQKSLDEERSKIEKIIQGMAASSLRCI 556
+++ + H KGA E +L +C+ ++ R + I A LR +
Sbjct: 283 TY-IDSDGSWHRASKGAPEQILNLCN----------CKEDVRKRAHSTIDKFAERGLRSL 429
Query: 557 AFAYMEISEGGDYIEKGKPRQVLREDGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKM 616
A + E G P Q +G++ L DP R + + + GV++KM
Sbjct: 430 GVARQVVPEKSK-DAPGAPWQ--------FVGLLPLFDPPRHDSAETITRALNLGVNVKM 582
Query: 617 I 617
I
Sbjct: 583 I 585
>TC19263 similar to UP|Q8L8A0 (Q8L8A0) Type IIB calcium ATPase MCA5, partial
(17%)
Length = 536
Score = 48.1 bits (113), Expect = 6e-06
Identities = 36/111 (32%), Positives = 54/111 (48%), Gaps = 8/111 (7%)
Frame = +3
Query: 1 IISKRNSHYVSKGTNHCSLVPHDVDKA-------RLSNMVKDKNLEAYSEFGGVEGVADV 53
++SK ++ VP DV A L ++V++ +++ + GGV GVA
Sbjct: 207 LVSKAALQFIQGSQPSEYKVPEDVKAAGFQICGDELGSIVENHDVKKFKFHGGVNGVAKK 386
Query: 54 LGTIPAKGILGSDDDTAARREL-FGTNTYVRPPPKIFLHFVLEALNDTTIL 103
L T +GI SD D RR+L +G N + K F FV EAL D T++
Sbjct: 387 LSTSVTEGI-SSDADILNRRQLIYGINKFTEIEAKSFWVFVWEALQDMTLM 536
>AU088926
Length = 487
Score = 47.8 bits (112), Expect = 8e-06
Identities = 38/137 (27%), Positives = 62/137 (44%), Gaps = 3/137 (2%)
Frame = +2
Query: 582 DGLTLLGIVGLKDPCRPNVKKAVETCKLAGVDIKMITGDNIFTAKAIATECGI-LDLNDA 640
D LG++ L DP R + + GV++KMITGD + K G+ ++ +
Sbjct: 5 DEWEFLGLLPLFDPPRHDSAXTIRRALDLGVNVKMITGDQLAIGKETGRRLGMGTNMXPS 184
Query: 641 GGVVVEGVE--FRNYTEEERMEKVDKIRVMARSSPMDKLLMVQCLKKKGHVVAVTGDGTN 698
++ + + + +E +EK D A P K +V L H+ +TG G N
Sbjct: 185 SSLLGQSKDAAIASIPIDELIEKADGF---AGVFPEHKXEIVXRLXDTKHICGMTGXGVN 355
Query: 699 DAPALKEADIGLSMGIQ 715
DAPAL + + L +Q
Sbjct: 356 DAPALXKXTLVLQWLMQ 406
>TC19888 weakly similar to PIR|T08551|T08551 Ca2+-transporting ATPase
homolog F27B13.140 - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (3%)
Length = 464
Score = 40.4 bits (93), Expect = 0.001
Identities = 15/29 (51%), Positives = 20/29 (68%)
Frame = +1
Query: 923 LNWEQWGICIGIAAVSWPIAWLTKLTPVP 951
LNW+QW IC+ I + WP+A + K PVP
Sbjct: 1 LNWKQWLICVIIGFIGWPLAVVGKFIPVP 87
>AV409298
Length = 332
Score = 40.0 bits (92), Expect = 0.002
Identities = 27/87 (31%), Positives = 43/87 (49%)
Frame = +3
Query: 194 DQIPADGLFLGGHSLQVDESSMTGESDHVEIEPLKAPFLLSGAKVVDGYAQMLVTAVGAN 253
D+IPADG+ G S VDESS TGE + + + + +G+ ++G + V G
Sbjct: 3 DRIPADGIVRAGRS-TVDESSFTGEP--LPVTKVAGCEVAAGSINLNGTLTLEVRRPGGE 173
Query: 254 TAWGQMMSSISGDNSERTPLQARLDKL 280
TA ++ + S P+Q DK+
Sbjct: 174 TAMADIVRLVEEAQSREAPVQRLADKV 254
>BP075842
Length = 434
Score = 38.9 bits (89), Expect = 0.004
Identities = 19/57 (33%), Positives = 29/57 (50%)
Frame = -1
Query: 910 MVELLRKFADTERLNWEQWGICIGIAAVSWPIAWLTKLTPVPSKLFFTNAKWVKSSL 966
++E + AD LN QW ICI + A+SW I W + P + + T+A S+
Sbjct: 434 VIEYAKALADAMGLNATQWAICILVGALSWVIQWPLRNLPDFLRTYCTSASNTPESI 264
>BP042017
Length = 470
Score = 35.8 bits (81), Expect = 0.033
Identities = 19/52 (36%), Positives = 28/52 (53%)
Frame = +3
Query: 168 VVRNGRPQQISIFDVLVGDVIYLKIGDQIPADGLFLGGHSLQVDESSMTGES 219
VV+ Q+ DV+ GD++ + GD P D L L V ++S+TGES
Sbjct: 78 VVQTELKVQVDHRDVVPGDIVIFEPGDLFPGDIRLLSSTHLVVSQASLTGES 233
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,339,679
Number of Sequences: 28460
Number of extensions: 172545
Number of successful extensions: 843
Number of sequences better than 10.0: 48
Number of HSP's better than 10.0 without gapping: 829
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 833
length of query: 967
length of database: 4,897,600
effective HSP length: 99
effective length of query: 868
effective length of database: 2,080,060
effective search space: 1805492080
effective search space used: 1805492080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0322.1


