

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0321.6
(497 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
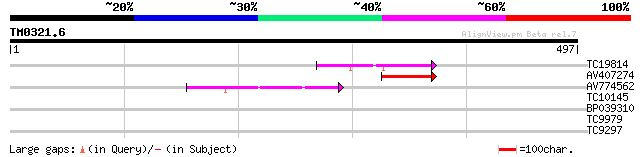
TC19814 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, pa... 67 9e-12
AV407274 55 2e-08
AV774562 45 3e-05
TC10145 similar to UP|Q8LJS8 (Q8LJS8) Homeodomain protein GhHOX1... 35 0.037
BP039310 30 1.2
TC9979 similar to UP|HIR5_MOUSE (Q9QZ23) HIRA-interacting protei... 29 1.5
TC9297 similar to UP|O65281 (O65281) ARABIDOPSIS THALIANA homeod... 27 7.7
>TC19814 homologue to UP|Q94C37 (Q94C37) At1g05230/YUP8H12_16, partial (17%)
Length = 389
Score = 66.6 bits (161), Expect = 9e-12
Identities = 40/112 (35%), Positives = 65/112 (57%), Gaps = 7/112 (6%)
Frame = +3
Query: 270 GEVKLLDGRKSVINLSNRMVGSFFKVLN--TQGIQNLLDPSNQLTTSIKLIAHRNTILD- 326
G + DGRKS++ L+ RMV SF ++ T + + T ++++ R ++ D
Sbjct: 54 GVITNQDGRKSMLKLAERMVISFCAGVSASTAHTWTTISGTGAETDDVRVMT-RKSVDDP 230
Query: 327 ----GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
GIV+ A +S WLPV PK +F FL+DE ++EWD L+ G +++ AHI+
Sbjct: 231 GRPPGIVLSAATSFWLPVPPKRVFEFLRDENSRSEWDILSNGGVVQEMAHIA 386
>AV407274
Length = 285
Score = 55.5 bits (132), Expect = 2e-08
Identities = 23/48 (47%), Positives = 35/48 (72%)
Frame = +2
Query: 327 GIVVIAVSSIWLPVTPKNLFLFLKDEFRKAEWDFLAGGRPMRQKAHIS 374
GIV+ A +S+W+PV+ LF FL+DE ++EWD L+ G PM++ HI+
Sbjct: 128 GIVLSAATSVWMPVSRHRLFDFLRDEQLRSEWDILSNGGPMQEMVHIA 271
>AV774562
Length = 460
Score = 45.1 bits (105), Expect = 3e-05
Identities = 38/139 (27%), Positives = 60/139 (42%), Gaps = 2/139 (1%)
Frame = +3
Query: 156 REYFFLRYCQQVEAGEWLIVDVSLDSFGNCAS--RSVAKKFPSGCRIRELPNGCCMVTWV 213
RE + +RYC+Q+ +W IVDVS+D ++ S A+ +R +
Sbjct: 3 REVYLVRYCKQLSGEQWAIVDVSIDKVKTTSTLPSSNAENAHPVALLRTSQMAIAK*SG* 182
Query: 214 EQVEVGQSFQLDMLVRDIVRNNIAYGAKRWLLELQRTFEKKACSEFSYTASTEGLEGEVK 273
E Q + + R IV + +A+GA+ W+ LQ E+ F T V
Sbjct: 183 STCEC-QKGAVHTMYRAIVNSGLAFGARHWIATLQLQCERLVF--FMATNVPMKDSTGVA 353
Query: 274 LLDGRKSVINLSNRMVGSF 292
RKS++ L+ RM F
Sbjct: 354 TWAERKSILKLAQRMTWGF 410
>TC10145 similar to UP|Q8LJS8 (Q8LJS8) Homeodomain protein GhHOX1, partial
(9%)
Length = 536
Score = 34.7 bits (78), Expect = 0.037
Identities = 22/66 (33%), Positives = 35/66 (52%), Gaps = 3/66 (4%)
Frame = +1
Query: 435 PSGFTISRDGREGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLM---LQRIKD 491
P T ++ + GSL T+ FQ+L ++S K T+ES+DS L+ L+ I+
Sbjct: 22 PLVITTRKEEKNTEGGSLFTIAFQILT----NASPTAKLTVESVDSVNTLVSCTLRNIRT 189
Query: 492 ALHCSD 497
+L C D
Sbjct: 190 SLQCED 207
>BP039310
Length = 584
Score = 29.6 bits (65), Expect = 1.2
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 304 LLDPSNQLTTSIKLIAHRNTILDGIVVIAVSSI 336
+L + TTS+ L+ HR T +DG V IA+ ++
Sbjct: 234 ILSLDHHTTTSLHLLQHRETTIDGSVQIALRNL 332
>TC9979 similar to UP|HIR5_MOUSE (Q9QZ23) HIRA-interacting protein 5
(mHIRIP5), partial (36%)
Length = 1304
Score = 29.3 bits (64), Expect = 1.5
Identities = 34/123 (27%), Positives = 52/123 (41%), Gaps = 10/123 (8%)
Frame = +2
Query: 325 LDGIVVIAVSSIWLPVTPKN--LFLFLKDEFRKAEWDFLAGGRPMRQKAHISINESNHV- 381
+DGI I S ++ VT + FLK E A DF + G+P+ + S +
Sbjct: 341 IDGITRIFFGSDFVTVTKSEDASWEFLKPEIFAAIMDFYSSGQPLFLDSQTSAAMDTAIK 520
Query: 382 -SILQTSSIIGEHMVIFQESSIDDLGGYVVYAPID------KLDAYMIVNGCDSSKKLIL 434
+ ++I E + ++ D GG +VY D KL +GC SS + L
Sbjct: 521 DDDSEIVAMIKELLETRIRPAVQDDGGDIVYRGSDPDTGIVKLQMQGACSGCPSS-SVTL 697
Query: 435 PSG 437
SG
Sbjct: 698 KSG 706
>TC9297 similar to UP|O65281 (O65281) ARABIDOPSIS THALIANA homeodomain
protein AHDP (SP:P93041), partial (7%)
Length = 598
Score = 26.9 bits (58), Expect = 7.7
Identities = 18/64 (28%), Positives = 36/64 (56%)
Frame = +3
Query: 427 DSSKKLILPSGFTISRDGREGNEGSLLTMCFQVLVDEEGSSSQVRKKTLESIDSSCMLML 486
DSS K + DG G+ G LLT+ FQ+LV+ ++ + ++++++++ +
Sbjct: 24 DSSNKSSTSQKGVAAADG--GSGGCLLTVGFQILVNNM-PTANLSVESVDTVNNLISCTI 194
Query: 487 QRIK 490
Q+IK
Sbjct: 195 QKIK 206
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,176,150
Number of Sequences: 28460
Number of extensions: 107850
Number of successful extensions: 435
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 429
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 432
length of query: 497
length of database: 4,897,600
effective HSP length: 94
effective length of query: 403
effective length of database: 2,222,360
effective search space: 895611080
effective search space used: 895611080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0321.6


