

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
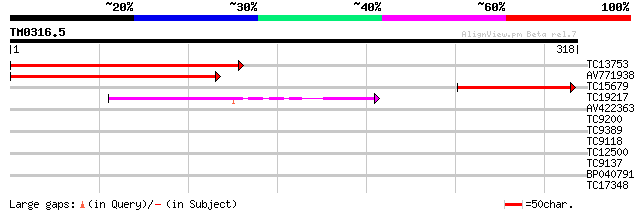
Query= TM0316.5
(318 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC13753 similar to UP|Q9LQQ6 (Q9LQQ6) F24B9.6, partial (38%) 258 7e-70
AV771938 222 6e-59
TC15679 137 3e-33
TC19217 67 5e-12
AV422363 32 0.19
TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, ... 31 0.24
TC9389 similar to UP|Q8GT82 (Q8GT82) Fiber protein Fb11, partial... 28 2.7
TC9118 similar to UP|Q84JE4 (Q84JE4) Squamosa promoter binding l... 27 3.5
TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein... 27 4.6
TC9137 UP|CAF02297 (CAF02297) Cysteine-rich polycomb-like protei... 27 4.6
BP040791 27 6.0
TC17348 similar to UP|CAE26126 (CAE26126) Possible ABC transport... 26 7.8
>TC13753 similar to UP|Q9LQQ6 (Q9LQQ6) F24B9.6, partial (38%)
Length = 614
Score = 258 bits (660), Expect = 7e-70
Identities = 131/131 (100%), Positives = 131/131 (100%)
Frame = +3
Query: 1 MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 60
MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL
Sbjct: 222 MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 401
Query: 61 EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKL 120
EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKL
Sbjct: 402 EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKL 581
Query: 121 MKVGENAATSD 131
MKVGENAATSD
Sbjct: 582 MKVGENAATSD 614
>AV771938
Length = 582
Score = 222 bits (566), Expect = 6e-59
Identities = 113/118 (95%), Positives = 114/118 (95%)
Frame = +2
Query: 1 MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 60
MAETVNHLNNHDSKEKEASPLAA LKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL
Sbjct: 227 MAETVNHLNNHDSKEKEASPLAAWLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 406
Query: 61 EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQ 118
EAKNLLLL+YCQSLVYY RKAKGFSIEEHPVVRSIVEIRLYLEKIR IDKKQQYQIQ
Sbjct: 407 EAKNLLLLSYCQSLVYYWWRKAKGFSIEEHPVVRSIVEIRLYLEKIRAIDKKQQYQIQ 580
>TC15679
Length = 609
Score = 137 bits (344), Expect = 3e-33
Identities = 66/66 (100%), Positives = 66/66 (100%)
Frame = +1
Query: 252 LFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNG 311
LFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNG
Sbjct: 1 LFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKGSNRNG 180
Query: 312 RLKKRK 317
RLKKRK
Sbjct: 181 RLKKRK 198
>TC19217
Length = 637
Score = 66.6 bits (161), Expect = 5e-12
Identities = 43/155 (27%), Positives = 78/155 (49%), Gaps = 3/155 (1%)
Frame = +2
Query: 56 GFSYLEAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQY 115
G Y E K LLLL+YCQ++ +YLL K++G + +HPVV + EIR L++I+ +D K
Sbjct: 14 GVRYFELKQLLLLSYCQAITFYLLLKSEGHPVHDHPVVARLAEIRELLDQIKQLDAKLPV 193
Query: 116 QIQKLMKVG---ENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAY 172
++ ++K E SD ++ P+ D +++ + P LVS E L
Sbjct: 194 GLEDILKESNGLETVVNSD--NENAPITID---SITRNQELP--LVSAESLE-------- 328
Query: 173 RPVKFAPTSMDQDKTSKQERNALRREKEILKESKH 207
+ P +D+ K ++ + + +++ + H
Sbjct: 329 ---EAMPIKVDEIKKLDSSKDGVPKARKVKHQKDH 424
>AV422363
Length = 485
Score = 31.6 bits (70), Expect = 0.19
Identities = 22/71 (30%), Positives = 37/71 (51%), Gaps = 4/71 (5%)
Frame = +2
Query: 185 DKTSKQERNA-LRREKEILKESKHSSFIRTLM---NDIEEKPEEIRDYEGSSKEVERHIA 240
+K KQ R +RR + +LKES+H + + M N++ E E + EG SK++ + A
Sbjct: 239 NKRRKQLREQFMRRTRLVLKESEHDPWCKRYMELYNELRENWERLYWDEGFSKKLAQDHA 418
Query: 241 KMEERAWQEEE 251
E +E+
Sbjct: 419 NYESAEEDDED 451
>TC9200 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(31%)
Length = 727
Score = 31.2 bits (69), Expect = 0.24
Identities = 22/79 (27%), Positives = 38/79 (47%), Gaps = 4/79 (5%)
Frame = +1
Query: 177 FAPTSMDQDKTSKQE----RNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSS 232
F P S+ + S E + ++ K + K S+ + L N +E E+++YE
Sbjct: 40 FNPHSLASLQESPNEAMVNESITKKRKLVPKSSQSAELPFKLQNKLEA---EVQEYEEVE 210
Query: 233 KEVERHIAKMEERAWQEEE 251
+EVE + + EE +EEE
Sbjct: 211 EEVEEEVEEEEEEEEEEEE 267
>TC9389 similar to UP|Q8GT82 (Q8GT82) Fiber protein Fb11, partial (97%)
Length = 572
Score = 27.7 bits (60), Expect = 2.7
Identities = 15/32 (46%), Positives = 19/32 (58%), Gaps = 7/32 (21%)
Frame = +1
Query: 254 NRVPLTKHERKREKYL-------KKSRNGLDG 278
N++P T ER+RE+ KKSRNG DG
Sbjct: 16 NKIPDTTQERRREQRRRRNKNLEKKSRNGSDG 111
>TC9118 similar to UP|Q84JE4 (Q84JE4) Squamosa promoter binding
like-protein, partial (16%)
Length = 779
Score = 27.3 bits (59), Expect = 3.5
Identities = 16/66 (24%), Positives = 30/66 (45%)
Frame = +2
Query: 203 KESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHE 262
+ S+H + EEK E++RD++ S K+ H + R + + R +H
Sbjct: 2 RSSRHEEEEEEEEEEEEEKREDVRDHKRSRKKHRSHRSSHTSR---DRDRDGRKHKRRHS 172
Query: 263 RKREKY 268
++KY
Sbjct: 173 SSKDKY 190
>TC12500 similar to GB|AAH45354.1|28279536|BC045354 FNBP4 protein {Danio
rerio;} , partial (5%)
Length = 678
Score = 26.9 bits (58), Expect = 4.6
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = +1
Query: 124 GENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAP 165
GE + LPS +P V+ P+P +L+S D AP
Sbjct: 277 GEASFPCSLPSPPQPPPPPPKYGVTTIPPSPPLLMSATDRAP 402
>TC9137 UP|CAF02297 (CAF02297) Cysteine-rich polycomb-like protein, complete
Length = 3086
Score = 26.9 bits (58), Expect = 4.6
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 2/64 (3%)
Frame = +1
Query: 125 ENAATSDLPSKKEPVAS--DKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSM 182
E+ D KE V+S +K D S P+M+ SREDL +E +D + P+
Sbjct: 1966 EDYVAPDHALSKERVSSNVEKGSD-STMLNKPEMVASREDLFQKEFYDQHHLSPITPSLQ 2142
Query: 183 DQDK 186
D+
Sbjct: 2143 CSDQ 2154
>BP040791
Length = 450
Score = 26.6 bits (57), Expect = 6.0
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = -1
Query: 43 TAKVKEGQYPTADGFSYLEAKNLLLLN 69
TA V G++ + G +LEA NLL LN
Sbjct: 441 TAAVFSGEFCSEKGGQFLEALNLLFLN 361
>TC17348 similar to UP|CAE26126 (CAE26126) Possible ABC transporter
ATP-binding component, membrane bound, partial (3%)
Length = 650
Score = 26.2 bits (56), Expect = 7.8
Identities = 27/102 (26%), Positives = 45/102 (43%), Gaps = 1/102 (0%)
Frame = +2
Query: 149 KYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNALRREKEILKESKHS 208
K RP+ R L P Y V++ P + +K +++R E + +H
Sbjct: 161 KRRPSDKPRRKRASLRPP---GPYAWVQYTPG--EPILPNKPNEGSVKRRNEKKRMRQHR 325
Query: 209 SFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKM-EERAWQE 249
+FI + + +E + + S K VER +A + ERAW E
Sbjct: 326 AFILAERKKRKAQMQEA-NRKKSIKRVERKMAAVARERAWAE 448
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.311 0.130 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,069,605
Number of Sequences: 28460
Number of extensions: 44395
Number of successful extensions: 210
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 205
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 208
length of query: 318
length of database: 4,897,600
effective HSP length: 90
effective length of query: 228
effective length of database: 2,336,200
effective search space: 532653600
effective search space used: 532653600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0316.5


