

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
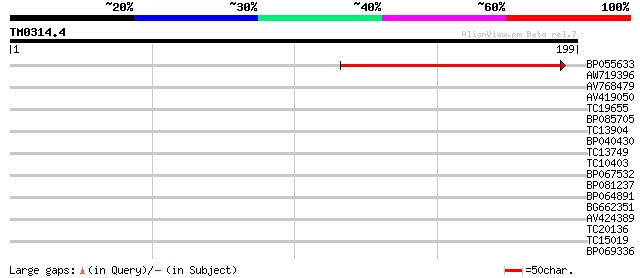
Query= TM0314.4
(199 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP055633 71 1e-13
AW719396 29 0.62
AV768479 29 0.62
AV419050 29 0.62
TC19655 27 2.4
BP085705 27 3.1
TC13904 weakly similar to UP|Q9ZU51 (Q9ZU51) RING-H2 finger prot... 26 4.0
BP040430 25 6.9
TC13749 similar to UP|Q9FKW9 (Q9FKW9) Alpha-mannosidase, partial... 25 6.9
TC10403 similar to UP|O80815 (O80815) T8F5.22 protein, partial (... 25 6.9
BP067532 25 9.0
BP081237 25 9.0
BP064891 25 9.0
BG662351 25 9.0
AV424389 25 9.0
TC20136 25 9.0
TC15019 UP|Q9FY16 (Q9FY16) Ferredoxin-nitrite reductase precurso... 25 9.0
BP069336 25 9.0
>BP055633
Length = 528
Score = 71.2 bits (173), Expect = 1e-13
Identities = 35/79 (44%), Positives = 49/79 (61%)
Frame = -2
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFL 176
W FK A++ +FQ T +PF LL++KQ+GSV + FE A ++G D E +F+
Sbjct: 518 WTKFKEAMLEQFQLTSNSSPFAALLALKQEGSVEEFVGQFERFAGMLKGIDEEHYMDIFV 339
Query: 177 NGLQEEIRAEMKLYPAADL 195
NGL+EEI AE+KLY L
Sbjct: 338 NGLKEEIAAEIKLYEPKSL 282
>AW719396
Length = 492
Score = 28.9 bits (63), Expect = 0.62
Identities = 32/119 (26%), Positives = 49/119 (40%)
Frame = +2
Query: 72 INRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPT 131
I +E+ L R G + MV + A +Q WE + +R+WE +L + +
Sbjct: 26 IRELEKALDLGR-GANPKGYMVEEIWQELAKAKYQEWERSSTKRSWE--LQSLKQACESA 196
Query: 132 LVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLNGLQEEIRAEMKLY 190
L + F GS M E F + A E L+GVF + +I AE+ Y
Sbjct: 197 LKEKHF-------LDGSDM---EGFVDDATTTHLKQLEALEGVFNKAAEADIPAEVPDY 343
>AV768479
Length = 462
Score = 28.9 bits (63), Expect = 0.62
Identities = 27/109 (24%), Positives = 43/109 (38%), Gaps = 16/109 (14%)
Frame = -2
Query: 55 GRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPE 114
G R I FDG Y +++V R + + +ALG +WWE++ P
Sbjct: 455 GFRAPIGFFDGMAEY-----------ITKVAGFPRKGLPECGQKEEALGGLRWWEDKEPA 309
Query: 115 R-------AWEPFKHALIRRFQPTL--------VQNPFG-PLLSIKQKG 147
+ +K IRR T+ V+N G PL +++ G
Sbjct: 308 HVARLLHYCEDYYKKGFIRRLNETMRDTWHDRVVENVLGVPLTTMRPCG 162
>AV419050
Length = 409
Score = 28.9 bits (63), Expect = 0.62
Identities = 17/41 (41%), Positives = 21/41 (50%)
Frame = +1
Query: 1 RTRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRER 41
R R R R R R+RDR+ + SR+ SR DH R R
Sbjct: 31 RERRREKEERDR-GDRDRDRDRTRSKRSRSRSRSIDHGRSR 150
>TC19655
Length = 473
Score = 26.9 bits (58), Expect = 2.4
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = +1
Query: 104 WFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPF 137
W W +PE+AW P H + R F P + + F
Sbjct: 178 WSLLWSSHSPEKAW-PSLHLMPR*FHPRRLLHHF 276
>BP085705
Length = 358
Score = 26.6 bits (57), Expect = 3.1
Identities = 12/24 (50%), Positives = 14/24 (58%), Gaps = 1/24 (4%)
Frame = -2
Query: 91 EMVMIAMEGKALG-WFQWWEEQAP 113
EM M M G LG W++WW E P
Sbjct: 309 EMQMTTMSGGGLGLWWRWWCEY*P 238
>TC13904 weakly similar to UP|Q9ZU51 (Q9ZU51) RING-H2 finger protein RHA2b,
partial (32%)
Length = 535
Score = 26.2 bits (56), Expect = 4.0
Identities = 7/26 (26%), Positives = 14/26 (52%)
Frame = -3
Query: 107 WWEEQAPERAWEPFKHALIRRFQPTL 132
WW + +R W ++ +R + P+L
Sbjct: 125 WWSHPSKQRRWNVWRQGSLRTWSPSL 48
>BP040430
Length = 516
Score = 25.4 bits (54), Expect = 6.9
Identities = 14/56 (25%), Positives = 24/56 (42%)
Frame = -3
Query: 98 EGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYR 153
E +AL WE+ + W PFK +R ++ L +++ V AY+
Sbjct: 505 EQQALEVCSKWEKNLKDPQWNPFKTITVRGETQEIIDEDDEKLERLRKDVGVGAYK 338
>TC13749 similar to UP|Q9FKW9 (Q9FKW9) Alpha-mannosidase, partial (17%)
Length = 618
Score = 25.4 bits (54), Expect = 6.9
Identities = 14/62 (22%), Positives = 28/62 (44%), Gaps = 14/62 (22%)
Frame = +1
Query: 67 DAYGWINRVERFF-----QLSRVGEEERLEMVMIAME---------GKALGWFQWWEEQA 112
D GW+ V+++F + E L+ V+ +++ + + +WW EQ+
Sbjct: 157 DDVGWLKTVDQYFVGSNNSIQGTSIENVLDSVVASLQKDPNRKFVFAEMAFFHRWWVEQS 336
Query: 113 PE 114
PE
Sbjct: 337 PE 342
>TC10403 similar to UP|O80815 (O80815) T8F5.22 protein, partial (15%)
Length = 1392
Score = 25.4 bits (54), Expect = 6.9
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = +3
Query: 19 DREASSHSGSRTGSRFSDHSRERFDHPHRV 48
DR+ SS GSRTG ++ R +HP V
Sbjct: 723 DRDRSSTPGSRTGRADYRNNGNRDEHPSGV 812
>BP067532
Length = 369
Score = 25.0 bits (53), Expect = 9.0
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = -1
Query: 102 LGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGP 139
+G ++ E PE P +LIRRF + + NP P
Sbjct: 357 MGGIMYYLEYEPE-TMAPVPRSLIRRFLASRISNPSPP 247
>BP081237
Length = 371
Score = 25.0 bits (53), Expect = 9.0
Identities = 8/23 (34%), Positives = 13/23 (55%)
Frame = -2
Query: 104 WFQWWEEQAPERAWEPFKHALIR 126
W +W EE+ ++ W+P H R
Sbjct: 334 WKRWKEEEL*QKMWDPKLHCRFR 266
>BP064891
Length = 480
Score = 25.0 bits (53), Expect = 9.0
Identities = 10/38 (26%), Positives = 20/38 (52%)
Frame = +2
Query: 71 WINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWW 108
W +VER L+R+G + +++++ K +G W
Sbjct: 293 WYLKVERDHTLTRIGIVAQFTQILMSLIDKKIGTILTW 406
>BG662351
Length = 407
Score = 25.0 bits (53), Expect = 9.0
Identities = 9/23 (39%), Positives = 13/23 (56%)
Frame = +2
Query: 107 WWEEQAPERAWEPFKHALIRRFQ 129
WW + +AW +K L+RR Q
Sbjct: 335 WWCRKPRVKAWNNYKQPLMRRKQ 403
>AV424389
Length = 309
Score = 25.0 bits (53), Expect = 9.0
Identities = 14/38 (36%), Positives = 20/38 (51%)
Frame = +3
Query: 115 RAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAY 152
R W PF + ++ PTL +P P LS K+ SV +
Sbjct: 108 RLWWPFHNPVMEAHGPTLHPSPV-PTLSCKKVPSVTLF 218
>TC20136
Length = 551
Score = 25.0 bits (53), Expect = 9.0
Identities = 12/39 (30%), Positives = 20/39 (50%)
Frame = -1
Query: 7 NTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHP 45
N P F + R+ + A+S + R RFS+ R + +P
Sbjct: 416 NIPHFPLSQRQERKHANSGASKR*NLRFSELMRTKQGYP 300
>TC15019 UP|Q9FY16 (Q9FY16) Ferredoxin-nitrite reductase precursor ,
complete
Length = 2098
Score = 25.0 bits (53), Expect = 9.0
Identities = 14/38 (36%), Positives = 21/38 (54%), Gaps = 1/38 (2%)
Frame = +2
Query: 84 VGEEERLEMVMIAMEGKALGWF-QWWEEQAPERAWEPF 120
VG + E ++ EG+A GW + +A ERA+ PF
Sbjct: 131 VGAQSGGEKWVLGFEGRAQGWH*SAGKGEAGERAYGPF 244
>BP069336
Length = 428
Score = 25.0 bits (53), Expect = 9.0
Identities = 9/30 (30%), Positives = 13/30 (43%)
Frame = -3
Query: 102 LGWFQWWEEQAPERAWEPFKHALIRRFQPT 131
L W + W+E WEP L+ + T
Sbjct: 309 LPWTRGWDESLDSFGWEPLHSELVLQLSTT 220
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,103,141
Number of Sequences: 28460
Number of extensions: 40773
Number of successful extensions: 298
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 298
length of query: 199
length of database: 4,897,600
effective HSP length: 86
effective length of query: 113
effective length of database: 2,450,040
effective search space: 276854520
effective search space used: 276854520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0314.4


