

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
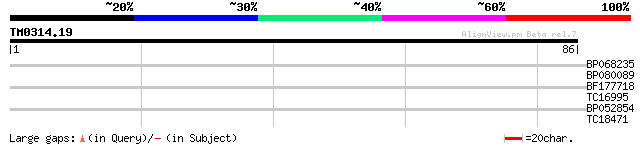
Query= TM0314.19
(86 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP068235 27 0.64
BP080089 24 5.5
BF177718 23 7.1
TC16995 similar to PIR|T05493|T05493 pathogenesis-related protei... 23 9.3
BP052854 23 9.3
TC18471 weakly similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein (At... 23 9.3
>BP068235
Length = 511
Score = 26.9 bits (58), Expect = 0.64
Identities = 17/47 (36%), Positives = 24/47 (50%), Gaps = 4/47 (8%)
Frame = -2
Query: 30 GIDKVRFCKPVIAGDTLVMRMTLTKL----QKSFELAKMEGKAYVGG 72
GID+V +P ++ MTL + Q SFE++ GKAY G
Sbjct: 384 GIDQVVMAQPWPEHAQTLLEMTLRRYKHGNQLSFEVSHFSGKAYQPG 244
>BP080089
Length = 392
Score = 23.9 bits (50), Expect = 5.5
Identities = 12/32 (37%), Positives = 19/32 (58%)
Frame = -1
Query: 35 RFCKPVIAGDTLVMRMTLTKLQKSFELAKMEG 66
R C P I+ +M+MTL L + FE+ ++G
Sbjct: 320 RMC-PGISFGLQLMKMTLATLLQGFEIVTLDG 228
>BF177718
Length = 484
Score = 23.5 bits (49), Expect = 7.1
Identities = 14/47 (29%), Positives = 23/47 (48%)
Frame = +3
Query: 2 LLQAMAQVGVLVMLQPEGGGSRENFFFVGIDKVRFCKPVIAGDTLVM 48
+L A++ L LQP+GG NF ++ K C + +T V+
Sbjct: 93 MLMISAELENLTNLQPQGGCDDPNFSYLFKVKCGRCGELSQRETCVV 233
>TC16995 similar to PIR|T05493|T05493 pathogenesis-related protein 19K4.140
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(22%)
Length = 614
Score = 23.1 bits (48), Expect = 9.3
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +2
Query: 3 LQAMAQVGVLVMLQPEGGGSRENFFFVG 30
LQ + G L+ LQP+G RE FVG
Sbjct: 284 LQ*LQLHGWLIWLQPQGVLQREAGIFVG 367
>BP052854
Length = 456
Score = 23.1 bits (48), Expect = 9.3
Identities = 11/35 (31%), Positives = 18/35 (51%)
Frame = +1
Query: 44 DTLVMRMTLTKLQKSFELAKMEGKAYVGGEVVCEG 78
DTL + + + S + A + G YV E++C G
Sbjct: 205 DTLHLHNSKPDKESSEDSAVLNGSCYVNLEMMCPG 309
>TC18471 weakly similar to UP|Q9SA38 (Q9SA38) F3O9.19 protein
(At1g16390/F3O9_19), partial (12%)
Length = 385
Score = 23.1 bits (48), Expect = 9.3
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = +2
Query: 71 GGEVVCEGEFLMAIG 85
G EVVC G F +AIG
Sbjct: 110 GSEVVCYGVFGLAIG 154
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.142 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,092,897
Number of Sequences: 28460
Number of extensions: 9480
Number of successful extensions: 45
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 45
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 45
length of query: 86
length of database: 4,897,600
effective HSP length: 62
effective length of query: 24
effective length of database: 3,133,080
effective search space: 75193920
effective search space used: 75193920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0314.19


