

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
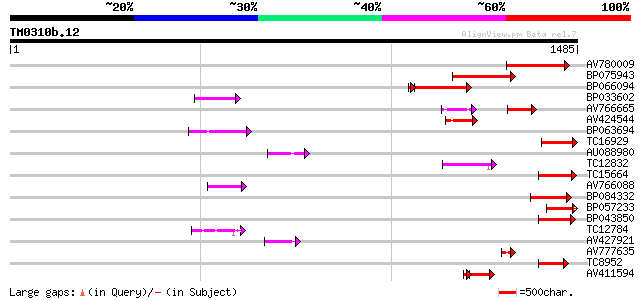
Query= TM0310b.12
(1485 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV780009 167 1e-41
BP075943 131 7e-31
BP066094 107 2e-26
BP033602 103 2e-22
AV766665 98 1e-20
AV424544 94 1e-19
BP063694 93 4e-19
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 92 5e-19
AU088980 88 1e-17
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 87 1e-17
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 85 1e-16
AV766088 82 6e-16
BP084332 80 3e-15
BP057233 76 4e-14
BP043850 74 2e-13
TC12784 similar to UP|Q850H6 (Q850H6) Gag-pol polyprotein (Fragm... 65 1e-10
AV427921 64 2e-10
AV777635 63 4e-10
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 62 5e-10
AV411594 54 8e-10
>AV780009
Length = 529
Score = 167 bits (423), Expect = 1e-41
Identities = 79/166 (47%), Positives = 113/166 (67%)
Frame = -1
Query: 1301 QAVDRILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTW 1360
QA R+L+Y+K +P +GL F L ++ Y+D+D+AG RRS TGY +FLG +L++W
Sbjct: 529 QAATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISW 350
Query: 1361 RSKKQNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHN 1420
R+KKQ V+RSS+EAE+RA+A VCE+ + + LK+ P+ LFCDN+SA+ IAHN
Sbjct: 349 RTKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKLNVPLPVPLFCDNQSALHIAHN 170
Query: 1421 PVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLP 1466
P H+RTKHIE+D H ++ KL + LI + + QLAD+ TK P
Sbjct: 169 PTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKPCP 32
>BP075943
Length = 547
Score = 131 bits (330), Expect = 7e-31
Identities = 69/165 (41%), Positives = 104/165 (62%), Gaps = 1/165 (0%)
Frame = -3
Query: 1160 NGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGKLKYFLGMEVAYSKQGIFI 1219
+G T LL+YV+D+++AGND E Q ++ L QF +KDLG+ KYFLG+E+A S GI +
Sbjct: 539 SGSFTALLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVL 360
Query: 1220 SQRKYVLDLLKETGKLGCKPMGVPIEQNHK-IGSSEEDLRVHKAQYQRLVGKLIYLSHTR 1278
+QRKY L L+ ++G G +P N + +G++ Y+R+VG+L+YL+ TR
Sbjct: 359 NQRKYALQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTR 180
Query: 1279 PDIAYPVSVVSQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKS 1323
PDI + V+ +SQF+ P + H Q + L+YL SPG GL + S
Sbjct: 179 PDITFAVNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPAS 45
>BP066094
Length = 532
Score = 107 bits (267), Expect(2) = 2e-26
Identities = 53/150 (35%), Positives = 94/150 (62%)
Frame = +2
Query: 1060 AKMNTVRIILSLAAHFDWELQQFDVKNAFLHGDLEEEVYMEIPPGYDPTSGRNKVCKLKK 1119
+K +R+++S + + + L Q +VK+AFL+G + EEVY+ PPG + + + KLKK
Sbjct: 86 SKTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKK 265
Query: 1120 ALYGLKQSPRAWFGRFTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGND 1179
+LYGLKQ+PRAW+ R ++ ++ + + D TLF K T + ++ +YV+D+I +
Sbjct: 266 SLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCK-TYKDDILIVQIYVDDIIFGSAN 442
Query: 1180 ELEKQNLRKELAVQFEMKDLGKLKYFLGME 1209
+ + + +FEM+ +G+LKYFLG++
Sbjct: 443 PSLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
Score = 30.4 bits (67), Expect(2) = 2e-26
Identities = 13/18 (72%), Positives = 15/18 (83%)
Frame = +3
Query: 1044 YTQTYGIDYDETFAPVAK 1061
Y+Q GIDY ETFAPVA+
Sbjct: 39 YSQQ*GIDYTETFAPVAR 92
>BP033602
Length = 533
Score = 103 bits (257), Expect = 2e-22
Identities = 56/126 (44%), Positives = 73/126 (57%), Gaps = 4/126 (3%)
Frame = +3
Query: 483 QIIGIAKEQDGLYCLQHE-ESKCHQTSSESWNTS---QIWLRHKRLGHPPFSILRTMFPH 538
++IG AK +DGLY + +K Q TS QI L H RLGHP F LR +FP
Sbjct: 156 KMIGSAKMKDGLYYFEDTFRNKAAQGLCGMSCTSVRDQIMLWHNRLGHPNFQYLRHLFPD 335
Query: 539 LFSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGPSSISNVSGAKWFVT 598
LF V+ S C+ C +K+ R+ + ++PF L+HSDVWGPS I+ G +WFVT
Sbjct: 336 LFKNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFYLIHSDVWGPSKITTQFGKRWFVT 515
Query: 599 FIDDCT 604
FIDD T
Sbjct: 516 FIDDHT 533
>AV766665
Length = 601
Score = 97.8 bits (242), Expect = 1e-20
Identities = 50/75 (66%), Positives = 58/75 (76%)
Frame = +3
Query: 1305 RILQYLKASPGRGLLFKKSEQLSMEVYTDADYAGSIVDRRSTTGYCMFLGGNLVTWRSKK 1364
RI YLKA+ RG LF+K + SM+ +T ADY GSIVDR ST GY MFL GNLVTWRSK+
Sbjct: 372 RIF*YLKANSRRGPLFQKEGKSSMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTWRSKQ 551
Query: 1365 QNVVARSSAEAEFRA 1379
QN++ARSS EAE RA
Sbjct: 552 QNIIARSSGEAELRA 596
Score = 53.5 bits (127), Expect = 2e-07
Identities = 38/94 (40%), Positives = 54/94 (57%), Gaps = 2/94 (2%)
Frame = +1
Query: 1130 AWFGRFTNAMVSLGYKQSQGDHTL--FIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLR 1187
AWF R T A + +S+ ++L + + L VYV+ M I G+DE+EKQ LR
Sbjct: 13 AWFVRLTEAT----HAESRWPYSLNKTCPQRQTHTSSHLCVYVDHMTIVGDDEVEKQILR 180
Query: 1188 KELAVQFEMKDLGKLKYFLGMEVAYSKQGIFISQ 1221
++LA QF++K KLKY +EV Y IFIS+
Sbjct: 181 EKLAPQFDLK---KLKYISVIEVVYP**NIFISK 273
>AV424544
Length = 276
Score = 94.4 bits (233), Expect = 1e-19
Identities = 48/83 (57%), Positives = 63/83 (75%)
Frame = +3
Query: 1142 LGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQFEMKDLGK 1201
LGY QS DH+LF K R+ T++LVYV+D+I+AGND E Q ++ +L +QF +KDLG
Sbjct: 30 LGYIQSAHDHSLFTKF-RDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQFRIKDLGT 206
Query: 1202 LKYFLGMEVAYSKQGIFISQRKY 1224
LKYFLG+EVA S G+F+SQRKY
Sbjct: 207 LKYFLGLEVARSSCGLFLSQRKY 275
>BP063694
Length = 511
Score = 92.8 bits (229), Expect = 4e-19
Identities = 52/169 (30%), Positives = 83/169 (48%), Gaps = 4/169 (2%)
Frame = -2
Query: 469 FFHSHGVFQDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTS-QIWLRHKRLGHP 527
F V Q+ TG ++G + GLY L + TSS S ++W H RLGH
Sbjct: 507 FLDDSFVIQNRKTGAVLGRXRCDQGLYVLNQDSQALLTTSSPLPRASFELW--HSRLGHV 334
Query: 528 PFSILRTMFPHL---FSKVSVESFHCDVCQFSKHHRSTFLPSNNKCAQPFDLVHSDVWGP 584
F I++ + H S + + C CQ +K R F +N + + DL+H D+ GP
Sbjct: 333 NFDIIKQLHKHGCLDVSSILPKPICCTSCQMAKSKRLVFHDNNKRASAVLDLIHCDLRGP 154
Query: 585 SSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVFQLFANFFHMIRTQF 633
S ++++ G +FV F+DD +R TW + +K KS+ + F + +F
Sbjct: 153 SPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVFMENRF 7
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 92.4 bits (228), Expect = 5e-19
Identities = 44/93 (47%), Positives = 61/93 (65%)
Frame = +1
Query: 1393 ILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVP 1452
+L DLK+ +E P ++CDN SA IA NPV H+RTKHIEID H ++E++ LI +
Sbjct: 16 LLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHLLPIS 195
Query: 1453 SGLQLADVLTKGLPLERFRELTCKLGMIDIHSP 1485
S LAD+ TK L + F ++ KLG+I+I SP
Sbjct: 196 SSEPLADIYTKALSPQNFHQICAKLGLINICSP 294
>AU088980
Length = 360
Score = 87.8 bits (216), Expect = 1e-17
Identities = 46/110 (41%), Positives = 64/110 (57%)
Frame = +2
Query: 675 NGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVSPVLVMTSF 734
N + ERK+RH+L +TRAL+F +PK W AV AAYLIN LPS +L P + +
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 735 FPSVPIKSGLQSRVFGCSAFVHVHGHHRGKLDPRAIKCIFIGYASDKKGY 784
P++ L +VFG + H+ KLDPRA KC+F+G+ + GY
Sbjct: 182 SPTL-----LDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGY 316
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 87.4 bits (215), Expect = 1e-17
Identities = 46/147 (31%), Positives = 81/147 (54%), Gaps = 8/147 (5%)
Frame = +2
Query: 1135 FTNAMVSLGYKQSQGDHTLFIKHTRNGKLTLLLVYVNDMIIAGNDELEKQNLRKELAVQF 1194
F + ++SLGY + DH + K + +LL+YV+DM++ G ++ Q L+ +LA +F
Sbjct: 2 FDSFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREF 181
Query: 1195 EMKDLGKLKYFLGMEVAYSKQG--IFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGS 1252
+MKDLG LGM++ ++ I++SQ+ Y+ +L+ C P+ P+ N+K+ S
Sbjct: 182 DMKDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSS 361
Query: 1253 S------EEDLRVHKAQYQRLVGKLIY 1273
S E + + + Y VG L+Y
Sbjct: 362 SMIPSSEAERMEMSRVPYASAVGSLMY 442
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 84.7 bits (208), Expect = 1e-16
Identities = 44/100 (44%), Positives = 64/100 (64%)
Frame = +1
Query: 1385 CELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSS 1444
CEL + IL DL++ + +P L+CD++SA IA N V H+RTKH++ID H ++EKL +
Sbjct: 1 CEL*WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAK 180
Query: 1445 LISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMIDIHS 1484
L + S Q AD+LTK L F L KLG+++I+S
Sbjct: 181 LFHLLPISSVDQTADILTKPLESGPFSHLVSKLGVLNIYS 300
>AV766088
Length = 501
Score = 82.0 bits (201), Expect = 6e-16
Identities = 41/103 (39%), Positives = 61/103 (58%), Gaps = 2/103 (1%)
Frame = -3
Query: 519 LRHKRLGHPPFSILRTMFPHLFSKVSVESFH--CDVCQFSKHHRSTFLPSNNKCAQPFDL 576
L H RLGHP S LR + + F + ++ C+ C F KH R F+ SNN PFD+
Sbjct: 307 LWHSRLGHPSSSALRYLRSNKFISYELLNYSPVCESCVFGKHVRLPFVSSNNVTVMPFDI 128
Query: 577 VHSDVWGPSSISNVSGAKWFVTFIDDCTRITWVFLMKDKSEVF 619
+HSD+W S + + +G +++V F+DD T W F + +KS+VF
Sbjct: 127 LHSDLW-TSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVF 2
>BP084332
Length = 368
Score = 79.7 bits (195), Expect = 3e-15
Identities = 41/106 (38%), Positives = 69/106 (64%)
Frame = +2
Query: 1365 QNVVARSSAEAEFRAMAQGVCELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQH 1424
Q+ +A S+AEAE+ + A ++L MK L+D +I E+ + ++CDN +AIS++ NP+ H
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILH 208
Query: 1425 DRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLPLERF 1470
R KHIE+ HFI++ + ++ ++V + Q AD+ TK L +RF
Sbjct: 209 SRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRF 346
>BP057233
Length = 473
Score = 75.9 bits (185), Expect = 4e-14
Identities = 40/79 (50%), Positives = 55/79 (68%)
Frame = -2
Query: 1407 LFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSLISTQYVPSGLQLADVLTKGLP 1466
L+CDN SA ++A NPV H R+KHIEID H+I++++ + + YVP+ Q+AD LTK L
Sbjct: 445 LWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIADCLTKPLS 266
Query: 1467 LERFRELTCKLGMIDIHSP 1485
RF +L KLG+ IHSP
Sbjct: 265 HTRFSQLRDKLGV--IHSP 215
>BP043850
Length = 515
Score = 73.6 bits (179), Expect = 2e-13
Identities = 36/96 (37%), Positives = 59/96 (60%)
Frame = -1
Query: 1386 ELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSSL 1445
E++ ++ +L +L P L DN SAI IA NPV H+ T+HIE+D H ++E + +
Sbjct: 512 EIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRRV 333
Query: 1446 ISTQYVPSGLQLADVLTKGLPLERFRELTCKLGMID 1481
I+ +V + +Q+AD+LTK L +R L KL +++
Sbjct: 332 ITLPHVSTSVQIADILTKSLTRQRHNFLVSKLMLVE 225
>TC12784 similar to UP|Q850H6 (Q850H6) Gag-pol polyprotein (Fragment),
partial (15%)
Length = 460
Score = 64.7 bits (156), Expect = 1e-10
Identities = 53/156 (33%), Positives = 75/156 (47%), Gaps = 14/156 (8%)
Frame = +2
Query: 477 QDLATGQIIGIAKEQDGLYCLQHEESKCHQTSSESWNTSQI--WLRHKRLGHPPFSILRT 534
QDL+T +IIG +E DGLY L SK +S ++ TS + + H R GHP +LR
Sbjct: 35 QDLSTPRIIGRGRESDGLYLLDEGVSK----ASTAYRTSSVSPFQLHCRFGHPSLGVLRK 202
Query: 535 MFPHLFSKVSVESFHCDVCQFS-KHHRSTFLPSNNKCAQP-FDLVHSDVWGPS------- 585
+ P L SV +F C+ C RS +LP NK A F +VH G
Sbjct: 203 LCPPL---ESVSNFSCESCHVC*TSXRSVYLPRLNKRAMSLFHIVHL*CMGSMPCNF*IR 373
Query: 586 ---SISNVSGAKWFVTFIDDCTRITWVFLMKDKSEV 618
S+S++ W + R W++LMK +SE+
Sbjct: 374 LRVSLSHL----WMIIL-----RAPWLYLMKSRSEL 454
>AV427921
Length = 284
Score = 63.5 bits (153), Expect = 2e-10
Identities = 35/96 (36%), Positives = 53/96 (54%)
Frame = +1
Query: 667 TCVDAPQQNGVAERKNRHLLEITRALLFQMNVPKFYWGEAVLTAAYLINRLPSRVLSNVS 726
T PQQNGV+ERKNR +L + R+LL +PK + EAV+ + +++NR P+ V+ N+
Sbjct: 10 TAAYTPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMM 189
Query: 727 PVLVMTSFFPSVPIKSGLQSRVFGCSAFVHVHGHHR 762
P + P+V R+ C A+ HV R
Sbjct: 190 PEEAWSGRQPAVD-----HFRISRCLAYAHVPDQKR 282
>AV777635
Length = 382
Score = 62.8 bits (151), Expect = 4e-10
Identities = 31/36 (86%), Positives = 32/36 (88%)
Frame = -2
Query: 1289 SQFMHDPQERHLQAVDRILQYLKASPGRGLLFKKSE 1324
SQFMHDP ERHL DRILQYLKASPGRGLLF+KSE
Sbjct: 381 SQFMHDPHERHL---DRILQYLKASPGRGLLFRKSE 283
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 62.4 bits (150), Expect = 5e-10
Identities = 33/79 (41%), Positives = 49/79 (61%)
Frame = +3
Query: 1385 CELLRMKIILDDLKIKYEAPMRLFCDNKSAISIAHNPVQHDRTKHIEIDRHFIKEKLNSS 1444
CE L + L DL+I + ++CDN+SA+ +A N V H RT++IEID H + K+
Sbjct: 18 CEALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFG 197
Query: 1445 LISTQYVPSGLQLADVLTK 1463
++ +VPS Q+ADV TK
Sbjct: 198 ILHLLHVPSSDQVADVFTK 254
>AV411594
Length = 244
Score = 53.5 bits (127), Expect(2) = 8e-10
Identities = 26/65 (40%), Positives = 40/65 (61%)
Frame = +2
Query: 1205 FLGMEVAYSKQGIFISQRKYVLDLLKETGKLGCKPMGVPIEQNHKIGSSEEDLRVHKAQY 1264
FL +A SK+GI +S+RKY L LL +T +GCKP P++ + K+ S++ +L Y
Sbjct: 50 FLRT*IAKSKKGILLSRRKYALQLLDDTRNMGCKPPPFPLDPSIKLNSTDGELLSDVTLY 229
Query: 1265 QRLVG 1269
R +G
Sbjct: 230 IRFIG 244
Score = 28.1 bits (61), Expect(2) = 8e-10
Identities = 12/21 (57%), Positives = 16/21 (76%)
Frame = +3
Query: 1189 ELAVQFEMKDLGKLKYFLGME 1209
ELA F++K L +KYFLG+E
Sbjct: 3 ELAKSFKLKVLDDMKYFLGLE 65
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.135 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,227,572
Number of Sequences: 28460
Number of extensions: 389939
Number of successful extensions: 2230
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 2150
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2194
length of query: 1485
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1383
effective length of database: 1,994,680
effective search space: 2758642440
effective search space used: 2758642440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0310b.12


