

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0306.6
(510 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
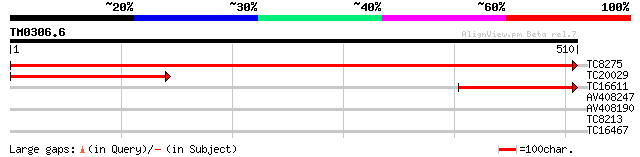
Score E
Sequences producing significant alignments: (bits) Value
TC8275 homologue to UP|Q94C02 (Q94C02) Myo-inositol-1-phosphate ... 1015 0.0
TC20029 homologue to UP|Q94G22 (Q94G22) 1L-myo-inositol-1-phosph... 284 3e-77
TC16611 homologue to GB|AAA85390.1|1161312|ATU04876 myo-inositol... 202 8e-53
AV408247 30 0.94
AV408190 29 1.6
TC8213 28 2.7
TC16467 UP|NUCC_LOTJA (Q9BBN8) NAD(P)H-quinone oxidoreductase ch... 27 6.1
>TC8275 homologue to UP|Q94C02 (Q94C02) Myo-inositol-1-phosphate synthase
, complete
Length = 1889
Score = 1015 bits (2624), Expect = 0.0
Identities = 510/510 (100%), Positives = 510/510 (100%)
Frame = +2
Query: 1 MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVP 60
MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVP
Sbjct: 113 MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVP 292
Query: 61 KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERANYFGSLTQASAIRVGSFQGEE 120
KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERANYFGSLTQASAIRVGSFQGEE
Sbjct: 293 KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERANYFGSLTQASAIRVGSFQGEE 472
Query: 121 IYAPFKSLLPMVNPDDVVFGGWDISDMNLADAMGRARVFDIDLQKQLRPYMESMVPLPGI 180
IYAPFKSLLPMVNPDDVVFGGWDISDMNLADAMGRARVFDIDLQKQLRPYMESMVPLPGI
Sbjct: 473 IYAPFKSLLPMVNPDDVVFGGWDISDMNLADAMGRARVFDIDLQKQLRPYMESMVPLPGI 652
Query: 181 YDPDFIAANQGDRANNVIKGTKKEQIQQIIKDIKEFKEASKVDRVVVLWTANTERYSNLV 240
YDPDFIAANQGDRANNVIKGTKKEQIQQIIKDIKEFKEASKVDRVVVLWTANTERYSNLV
Sbjct: 653 YDPDFIAANQGDRANNVIKGTKKEQIQQIIKDIKEFKEASKVDRVVVLWTANTERYSNLV 832
Query: 241 VGLNDTMENLLASVDRNEAEISPSTLYAIACVMENIPFINGSPQNTFVPGLIDLAIKKNS 300
VGLNDTMENLLASVDRNEAEISPSTLYAIACVMENIPFINGSPQNTFVPGLIDLAIKKNS
Sbjct: 833 VGLNDTMENLLASVDRNEAEISPSTLYAIACVMENIPFINGSPQNTFVPGLIDLAIKKNS 1012
Query: 301 LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS 360
LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS
Sbjct: 1013LIGGDDFKSGQTKMKSVLVDFLVGAGIKPTSIVSYNHLGNNDGMNLSAPQTFRSKEISKS 1192
Query: 361 NVVDDMVNSNAILYEPGEHPDHVVVIKYVPYVADSKRAMDEYTSEIFMGGKNTIVLHNTC 420
NVVDDMVNSNAILYEPGEHPDHVVVIKYVPYVADSKRAMDEYTSEIFMGGKNTIVLHNTC
Sbjct: 1193NVVDDMVNSNAILYEPGEHPDHVVVIKYVPYVADSKRAMDEYTSEIFMGGKNTIVLHNTC 1372
Query: 421 EDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLTKAPLVPPGTPVVN 480
EDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLTKAPLVPPGTPVVN
Sbjct: 1373EDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLTKAPLVPPGTPVVN 1552
Query: 481 ALSKQRAMLENILRACVGLAPENNMILEYK 510
ALSKQRAMLENILRACVGLAPENNMILEYK
Sbjct: 1553ALSKQRAMLENILRACVGLAPENNMILEYK 1642
>TC20029 homologue to UP|Q94G22 (Q94G22) 1L-myo-inositol-1-phosphate
synthase (Fragment), partial (31%)
Length = 506
Score = 284 bits (726), Expect = 3e-77
Identities = 133/144 (92%), Positives = 142/144 (98%)
Frame = +3
Query: 1 MFIENFKVESPNVKYTETEIQSVYNYETTELVHENRNGSYQWIVKPKSVKYEFKTSTNVP 60
MFIE+FKVESPNVKYTETEIQSVYNYETTELVHEN+N +YQW+VKPK+VKYEFKT TNVP
Sbjct: 75 MFIESFKVESPNVKYTETEIQSVYNYETTELVHENKNNAYQWVVKPKTVKYEFKTKTNVP 254
Query: 61 KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERANYFGSLTQASAIRVGSFQGEE 120
KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKI+++NYFGSLTQASAIRVGSFQGEE
Sbjct: 255 KLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIQQSNYFGSLTQASAIRVGSFQGEE 434
Query: 121 IYAPFKSLLPMVNPDDVVFGGWDI 144
IYAPFKSLLPMVNPDD+VFGGWDI
Sbjct: 435 IYAPFKSLLPMVNPDDIVFGGWDI 506
>TC16611 homologue to GB|AAA85390.1|1161312|ATU04876
myo-inositol-1-phosphate synthase {Arabidopsis
thaliana;} , partial (21%)
Length = 527
Score = 202 bits (515), Expect = 8e-53
Identities = 100/107 (93%), Positives = 105/107 (97%)
Frame = +2
Query: 404 SEIFMGGKNTIVLHNTCEDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATIL 463
+EIFMGGKNTIV+HNTCEDSLLAAPIILDLVLLAELSTRIQFK+E E KFHSFHPVATIL
Sbjct: 8 AEIFMGGKNTIVMHNTCEDSLLAAPIILDLVLLAELSTRIQFKAENESKFHSFHPVATIL 187
Query: 464 SYLTKAPLVPPGTPVVNALSKQRAMLENILRACVGLAPENNMILEYK 510
SYLTKAPLVPPGTPVVNALSKQRAMLENI+RACVGLAPENNMI+EYK
Sbjct: 188 SYLTKAPLVPPGTPVVNALSKQRAMLENIMRACVGLAPENNMIMEYK 328
>AV408247
Length = 394
Score = 30.0 bits (66), Expect = 0.94
Identities = 18/44 (40%), Positives = 23/44 (51%), Gaps = 4/44 (9%)
Frame = +1
Query: 440 STRIQFKSEAEDKFHS----FHPVATILSYLTKAPLVPPGTPVV 479
+TR F ++ HS F P+A LS LT P+ PP PVV
Sbjct: 190 TTRSPFPIKSSPVLHSIPIIFDPIALFLS-LTPPPMAPPAAPVV 318
>AV408190
Length = 382
Score = 29.3 bits (64), Expect = 1.6
Identities = 18/54 (33%), Positives = 29/54 (53%)
Frame = -1
Query: 46 PKSVKYEFKTSTNVPKLGVMLVGWGGNNGSTLTGGVIANKEGISWATKDKIERA 99
P+ + E S +V + G+ +VG GG+ S + G V +EG+ A +D I A
Sbjct: 241 PEVLVGELAASGSVDEGGLAMVGAGGDESSDV*GVVWREREGLPLAVED*IGAA 80
>TC8213
Length = 935
Score = 28.5 bits (62), Expect = 2.7
Identities = 26/59 (44%), Positives = 28/59 (47%)
Frame = +1
Query: 416 LHNTCEDSLLAAPIILDLVLLAELSTRIQFKSEAEDKFHSFHPVATILSYLTKAPLVPP 474
LHN SLL IL L L L TR Q K A HSF+ V IL + K L PP
Sbjct: 1 LHNVTYCSLLTW--ILILAWLPHLFTRGQTKL-AHHAQHSFNWVFLILGWSYKTSLCPP 168
>TC16467 UP|NUCC_LOTJA (Q9BBN8) NAD(P)H-quinone oxidoreductase chain H,
chloroplast (NAD(P)H dehydrogenase, chain H)
(NADH-plastoquinone oxidoreductase 49 kDa subunit) ,
complete
Length = 2407
Score = 27.3 bits (59), Expect = 6.1
Identities = 10/34 (29%), Positives = 22/34 (64%)
Frame = +3
Query: 57 TNVPKLGVMLVGWGGNNGSTLTGGVIANKEGISW 90
+++ +G+++ G+G NN + GG+ A + IS+
Sbjct: 1668 SSIAPIGLLMSGYGSNNKYSFLGGLRAAAQSISY 1769
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,115,895
Number of Sequences: 28460
Number of extensions: 82776
Number of successful extensions: 408
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 407
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 408
length of query: 510
length of database: 4,897,600
effective HSP length: 94
effective length of query: 416
effective length of database: 2,222,360
effective search space: 924501760
effective search space used: 924501760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0306.6


