

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
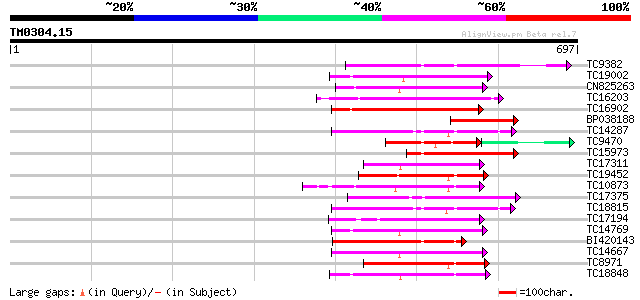
Query= TM0304.15
(697 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC9382 weakly similar to UP|ABL_DROME (P00522) Tyrosine-protein ... 174 3e-44
TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866)... 122 3e-28
CN825263 117 5e-27
TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodula... 115 2e-26
TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-... 114 4e-26
BP038188 114 4e-26
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 112 2e-25
TC9470 similar to UP|EPA5_HUMAN (P54756) Ephrin type-A receptor ... 104 3e-25
TC15973 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F... 112 3e-25
TC17311 similar to UP|Q8K2Y2 (Q8K2Y2) Receptor-interacting serin... 111 4e-25
TC19452 homologue to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, parti... 110 8e-25
TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partia... 109 2e-24
TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corn... 107 5e-24
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 107 6e-24
TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete 107 8e-24
TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete 106 1e-23
BI420143 106 1e-23
TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, com... 106 1e-23
TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P... 104 5e-23
TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, pa... 102 2e-22
>TC9382 weakly similar to UP|ABL_DROME (P00522) Tyrosine-protein kinase Abl
(D-ash) , partial (5%)
Length = 1197
Score = 174 bits (442), Expect = 3e-44
Identities = 94/278 (33%), Positives = 156/278 (55%), Gaps = 1/278 (0%)
Frame = +2
Query: 414 VAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLF 473
VAIKV + + L+++ +E+ IM+++RH NV+ F+GA L IVTE + RGSL+
Sbjct: 17 VAIKVLKPERISTDMLKEFAQEVYIMRKIRHKNVVQFIGACTRTPNLCIVTEFMSRGSLY 196
Query: 474 KTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFG 533
LHK + L++A+D+++GMNYLH N I+HRDLK+ NLL+D+N VKV DFG
Sbjct: 197 DFLHKQRGVFKLPSLLKVAIDVSKGMNYLHQNN--IIHRDLKTGNLLMDENELVKVADFG 370
Query: 534 LSR-LKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLN 592
++R + + ++T ++ GT +WMAPE++ ++P ++K+DV+S+G+ LWEL+T +P+ L
Sbjct: 371 VARVITQSGVMTAET--GTYRWMAPEVIEHKPYDQKADVFSFGIALWELLTGELPYSYLT 544
Query: 593 SLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLL 652
LQ V R +P+ P ++ ++ CW++
Sbjct: 545 PLQAAVGVVQKGLRPSIPKNTHPRLSELLQRCWKQ------------------------- 649
Query: 653 VLNMSSDPEQRPSFEELIQRMLFVLNRVTAESVKRSSE 690
DP +RP+F E+I+ + + V R +
Sbjct: 650 ------DPIERPAFSEIIEILQNIAKEVNDTKTDRHKD 745
>TC19002 weakly similar to UP|Q9XJ10 (Q9XJ10) ESTs C22458(C62866), partial
(48%)
Length = 780
Score = 122 bits (305), Expect = 3e-28
Identities = 71/206 (34%), Positives = 120/206 (57%), Gaps = 6/206 (2%)
Frame = +1
Query: 394 DEIGQGSYAVVYHG-IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMG 452
+++G+G + VY G +W+GS +A+K ++ + ++ E++I+ R+RH N+L G
Sbjct: 76 NKLGEGGFGSVYWGQLWDGSQIAVKRL--KVWSNKADMEFAVEVEILARVRHKNLLSLRG 249
Query: 453 AVYSLERLAIVTELLPRGSLFKTLHKSNQT---LDIRRRLRMALDIARGMNYLHHR-NPP 508
+ IV + +P SL LH + + LD RR+ +A+ A G+ YLHH+ P
Sbjct: 250 YCAEGQERLIVYDYMPNLSLLSHLHGQHSSECLLDWNRRMNIAIGSAEGIVYLHHQATPH 429
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRL-KDTTLLTTKSGRGTPQWMAPEILRNEPSNE 567
I+HRD+K+SN+L+D ++ +V DFG ++L D T +GT ++APE +NE
Sbjct: 430 IIHRDIKASNVLLDSDFQARVADFGFAKLIPDGATHVTTRVKGTLGYLAPEYAMLGKANE 609
Query: 568 KSDVYSYGVVLWELMTQSIPWENLNS 593
DV+S+G++L EL + P E L+S
Sbjct: 610 CCDVFSFGILLLELASGKKPLEKLSS 687
>CN825263
Length = 663
Score = 117 bits (294), Expect = 5e-27
Identities = 74/194 (38%), Positives = 113/194 (58%), Gaps = 7/194 (3%)
Frame = +1
Query: 401 YAVVYHGIWN-GSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLER 459
+ +VY GI N G DVA+K+ + + +++ E++++ RL H N++ +G +
Sbjct: 4 FGLVYKGILNDGRDVAVKILKRDD--QRGGREFLAEVEMLSRLHHRNLVKLIGICIEKQT 177
Query: 460 LAIVTELLPRGSLFKTLH---KSNQTLDIRRRLRMALDIARGMNYLHH-RNPPIVHRDLK 515
++ EL+P GS+ LH K LD R+++AL ARG+ YLH NP ++HRD K
Sbjct: 178 RCLIYELVPNGSVESHLHGADKETGPLDWNARMKIALGAARGLAYLHEDSNPCVIHRDFK 357
Query: 516 SSNLLVDKNWTVKVGDFGLSR--LKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYS 573
SSN+L++ ++T KV DFGL+R L + + GT ++APE KSDVYS
Sbjct: 358 SSNILLECDFTPKVSDFGLARTALDEGNKHISTHVMGTFGYLAPEYAMTGHLLVKSDVYS 537
Query: 574 YGVVLWELMTQSIP 587
YGVVL EL+T + P
Sbjct: 538 YGVVLLELLTGTKP 579
>TC16203 UP|Q8GRU6 (Q8GRU6) LRR receptor-like kinase (Hypernodulation aberrant
root formation protein), complete
Length = 3308
Score = 115 bits (289), Expect = 2e-26
Identities = 82/237 (34%), Positives = 134/237 (55%), Gaps = 7/237 (2%)
Frame = +2
Query: 378 DSVVDCEIHWEDLHLRDE--IGQGSYAVVYHGIW-NGSDVAIKVYFGNGYTEETLQDYKK 434
+ VV+C L++E IG+G +VY G NG+DVAIK G G ++
Sbjct: 2174 EDVVEC--------LKEENIIGKGGAGIVYRGSMPNGTDVAIKRLVGQGSGRNDY-GFRA 2326
Query: 435 EIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQT-LDIRRRLRMAL 493
EI+ + ++RH N++ +G V + + ++ E +P GSL + LH + L R ++A+
Sbjct: 2327 EIETLGKIRHRNIMRLLGYVSNKDTNLLLYEYMPNGSLGEWLHGAKGGHLRWEMRYKIAV 2506
Query: 494 DIARGMNYLHHR-NPPIVHRDLKSSNLLVDKNWTVKVGDFGLSR-LKDTTLLTTKSG-RG 550
+ ARG+ Y+HH +P I+HRD+KS+N+L+D ++ V DFGL++ L D + S G
Sbjct: 2507 EAARGLCYMHHDCSPLIIHRDVKSNNILLDADFEAHVADFGLAKFLYDPGASQSMSSIAG 2686
Query: 551 TPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRL 607
+ ++APE +EKSDVYS+GVVL EL+ P V +VG++++ +
Sbjct: 2687 SYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIGRKPVGEFG--DGVDIVGWVNKTM 2851
>TC16902 homologue to UP|Q8VZH5 (Q8VZH5) Receptor protein kinase-like
protein, partial (46%)
Length = 941
Score = 114 bits (286), Expect = 4e-26
Identities = 67/192 (34%), Positives = 119/192 (61%), Gaps = 5/192 (2%)
Frame = +3
Query: 396 IGQGSYAVVYHG-IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAV 454
+G G + VY+G + G+ VAIK GN +E+ + +++ EI+++ +LRH +++ +G
Sbjct: 15 LGVGGFGKVYYGEVDGGTKVAIKR--GNPLSEQGVHEFQTEIEMLSKLRHRHLVSLIGYC 188
Query: 455 YSLERLAIVTELLPRGSLFKTLHKSNQT-LDIRRRLRMALDIARGMNYLHH-RNPPIVHR 512
+ +V + + G+L + L+K+ + L ++RL + + ARG++YLH I+HR
Sbjct: 189 EENTEMILVYDHMAYGTLREHLYKTQKPPLPWKQRLEICIGAARGLHYLHTGAKYTIIHR 368
Query: 513 DLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSG--RGTPQWMAPEILRNEPSNEKSD 570
D+K++N+L+D+ W KV DFGLS+ T T S +G+ ++ PE R + +KSD
Sbjct: 369 DVKTTNILLDEKWVAKVSDFGLSKTGPTLDNTHVSTVVKGSFGYLDPEYFRRQQLTDKSD 548
Query: 571 VYSYGVVLWELM 582
VYS+GVVL+E++
Sbjct: 549 VYSFGVVLFEIL 584
>BP038188
Length = 539
Score = 114 bits (286), Expect = 4e-26
Identities = 49/83 (59%), Positives = 65/83 (78%)
Frame = +2
Query: 543 LTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGF 602
L++KS GTP+WMAPE+LR+EPSNEKSDVYS+G++LWEL T PW NLN QVV VGF
Sbjct: 5 LSSKSAAGTPEWMAPEVLRDEPSNEKSDVYSFGIILWELATLQQPWGNLNPAQVVAAVGF 184
Query: 603 MDRRLDLPEGLDPHVASIINDCW 625
+RL++P ++P +A++I CW
Sbjct: 185 KGKRLEIPCDVNPQLAAMIEACW 253
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 112 bits (281), Expect = 2e-25
Identities = 73/236 (30%), Positives = 131/236 (54%), Gaps = 8/236 (3%)
Frame = +3
Query: 396 IGQGSYAVVYHG--IWNGSDVAIKVYFGNGYTEETL-QDYKKEIDIMKRLRHPNVLLFMG 452
+GQG++A VYHG + +VAIKV ++ L + K+E+ +M+ +RHP+++
Sbjct: 450 LGQGNFAKVYHGRNLATNENVAIKVIKKEKLKKDRLVKQIKREVSVMRLVRHPHIVELKE 629
Query: 453 AVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHR 512
+ + ++ +V E + G LF ++K D R+ L A +++ H R + HR
Sbjct: 630 VMATKGKIFLVMEYVKGGELFTKVNKGKLNEDDARKYFQQLISA--VDFCHSRG--VTHR 797
Query: 513 DLKSSNLLVDKNWTVKVGDFGLSRL----KDTTLLTTKSGRGTPQWMAPEILRNEP-SNE 567
DLK NLL+D+N +KV DFGLS L +D +L T GTP ++APE+L+ +
Sbjct: 798 DLKPENLLLDENEDLKVSDFGLSALPEQRRDDGMLVTPC--GTPAYVAPEVLKKKGYDGS 971
Query: 568 KSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIIND 623
K+D++S GV+L+ L++ +P++ N +++ + PE + P ++I++
Sbjct: 972 KADIWSCGVILYALLSGYLPFQGENVMRIYRKA--FKAEYEFPEWISPQAKNLISN 1133
>TC9470 similar to UP|EPA5_HUMAN (P54756) Ephrin type-A receptor 5
precursor (Tyrosine-protein kinase receptor EHK-1) (Eph
homology kinase-1) (Receptor protein-tyrosine kinase
HEK7) , partial (3%)
Length = 1204
Score = 104 bits (259), Expect(2) = 3e-25
Identities = 62/124 (50%), Positives = 81/124 (65%), Gaps = 6/124 (4%)
Frame = +2
Query: 463 VTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVD 522
VTE + GSL L K+ + LD R+RL +A+D+A GM YLH +N IVH DLKS NLLV+
Sbjct: 17 VTEYMVNGSLRNALQKNGRNLDKRKRLLIAMDVAFGMEYLHGKN--IVHFDLKSDNLLVN 190
Query: 523 ----KNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPS--NEKSDVYSYGV 576
KVGD GLS++K TL+ + RGT WMAPE+L S +EK DV+S+G+
Sbjct: 191 LRDPHRPICKVGDLGLSKVKCQTLI-SGGVRGTLPWMAPELLNGSSSLVSEKVDVFSFGI 367
Query: 577 VLWE 580
V+WE
Sbjct: 368 VMWE 379
Score = 28.5 bits (62), Expect(2) = 3e-25
Identities = 19/114 (16%), Positives = 44/114 (37%)
Frame = +3
Query: 581 LMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYG 640
L+T P+ +L+ ++G + R +PE DP ++ CW
Sbjct: 381 LLTGEEPYADLHYGAIIGGIVNNTLRPPVPESCDPDWKVLMEKCW--------------- 515
Query: 641 RNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVTAESVKRSSETCNA 694
SS+P +RP+F E+ + + ++++ + + + ++
Sbjct: 516 ----------------SSEPSERPTFTEIASELRSIGSKISPKGQNQQQQPASS 629
>TC15973 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F4P13_4
{Arabidopsis thaliana;}, partial (49%)
Length = 944
Score = 112 bits (279), Expect = 3e-25
Identities = 52/137 (37%), Positives = 87/137 (62%)
Frame = +3
Query: 489 LRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSG 548
+++ALD+ARG++YLH + IVHRD+K+ N+L+DK TVK+ DFG++R++ +
Sbjct: 66 IQLALDLARGLSYLHSQK--IVHRDVKTENMLLDKTRTVKIADFGVARVEASNPNDMTGE 239
Query: 549 RGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLD 608
GT +MAPE+L P N K DVYS+G+ LWE+ +P+ +L+ ++ V + R +
Sbjct: 240 TGTLGYMAPEVLNGNPYNRKCDVYSFGICLWEIYCCDMPYPDLSFSEITSAVVRQNLRPE 419
Query: 609 LPEGLDPHVASIINDCW 625
+P +A+++ CW
Sbjct: 420 IPRCCPSSLANVMKKCW 470
>TC17311 similar to UP|Q8K2Y2 (Q8K2Y2) Receptor-interacting serine-threonine
kinase 3, partial (7%)
Length = 606
Score = 111 bits (277), Expect = 4e-25
Identities = 65/163 (39%), Positives = 94/163 (56%), Gaps = 14/163 (8%)
Frame = +3
Query: 435 EIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKS-----------NQTL 483
E++I+ +RH N++ + S E L +V + L SL + LHK N +
Sbjct: 3 EVEILSNIRHNNIVKLQCCISSEESLLLVYDYLENLSLDRWLHKKSKPPTVSGSVQNNII 182
Query: 484 DIRRRLRMALDIARGMNYLHHR-NPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL--KDT 540
D RRL +A+ +A+G+ Y+HH +PP+VHRDLK SN+L+D + KV DFGL+ + K
Sbjct: 183 DWPRRLHIAIGVAQGLCYMHHDCSPPVVHRDLKPSNILLDSQFNAKVADFGLAMMSVKPE 362
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMT 583
L T + GT ++APE NEK DVYS+GV+L EL T
Sbjct: 363 ELATMSAVAGTFGYIAPEYALTIRVNEKIDVYSFGVILLELTT 491
>TC19452 homologue to UP|Q8S9J9 (Q8S9J9) At1g14000/F7A19_9, partial (41%)
Length = 562
Score = 110 bits (275), Expect = 8e-25
Identities = 61/168 (36%), Positives = 105/168 (62%), Gaps = 8/168 (4%)
Frame = +3
Query: 429 LQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRR 488
+QD++ E++++ +LRHPN++ F+GAV + L ++TE L G L + L K +L
Sbjct: 30 IQDFRHEVNLLVKLRHPNIVQFLGAVTERKPLMLITEYLRGGDLHQYL-KEKGSLSPSTA 206
Query: 489 LRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWT--VKVGDFGLSRL------KDT 540
+ ++DI RGM YLH+ I+HRDLK N+L+ + +KVGDFGLS+L D
Sbjct: 207 INFSMDIVRGMAYLHNEPNVIIHRDLKPRNVLLVNSSADHLKVGDFGLSKLITVQNSHDV 386
Query: 541 TLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPW 588
+T ++ G+ ++MAPE+ ++ ++K DVYS+ ++L+E++ P+
Sbjct: 387 YKMTGET--GSYRYMAPEVFKHRKYDKKVDVYSFAMILYEMLEGEPPF 524
>TC10873 similar to UP|Q9LDZ5 (Q9LDZ5) F2D10.13 (F5M15.3), partial (77%)
Length = 1478
Score = 109 bits (272), Expect = 2e-24
Identities = 77/233 (33%), Positives = 132/233 (56%), Gaps = 9/233 (3%)
Frame = +1
Query: 360 SSRESVGSHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHG-IWNGSDVAIKV 418
SS S +++S G + + D ++++ +L IG+G + VY G + G VA+K
Sbjct: 373 SSNGSSNGKTAAASFGFRE-LADATRNFKEANL---IGEGGFGKVYKGRLTTGEAVAVKQ 540
Query: 419 YFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSL----FK 474
+G + Q++ E+ ++ L H N++ +G ++ +V E +P GSL F+
Sbjct: 541 LSHDG--RQGFQEFVMEVLMLSLLHHTNLVRLIGYCTDGDQRLLVYEYMPMGSLEDHLFE 714
Query: 475 TLHKSNQTLDIRRRLRMALDIARGMNYLH-HRNPPIVHRDLKSSNLLVDKNWTVKVGDFG 533
H + L+ R+++A+ ARG+ YLH +PP+++RDLKS+N+L+D + K+ DFG
Sbjct: 715 LSH-DKEPLNWSTRMKVAVGAARGLEYLHCTADPPVIYRDLKSANILLDNEFNPKLSDFG 891
Query: 534 LSRL---KDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMT 583
L++L D T ++T+ GT + APE + KSD+YS+GVVL EL+T
Sbjct: 892 LAKLGPVGDNTHVSTRV-MGTYGYCAPEYAMSGKLTLKSDIYSFGVVLLELLT 1047
>TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corniculatus
var. japonicus]
Length = 1422
Score = 107 bits (268), Expect = 5e-24
Identities = 64/215 (29%), Positives = 110/215 (50%), Gaps = 3/215 (1%)
Frame = +3
Query: 416 IKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKT 475
+KV+ + ++E L+ +EI+++ + HPN++ + G+ E L++ E + GS+ K
Sbjct: 24 VKVFSDDKTSKECLKQLNQEINLLNQFSHPNIVQYYGSELGEESLSVYLEYVSGGSIHKL 203
Query: 476 LHKSNQTLD--IRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFG 533
L + + I+ R I G+ YLH RN VHRD+K +N+LVD N +K+ DFG
Sbjct: 204 LQEYGAFKEPVIQNYTRQ---IVSGLAYLHSRNT--VHRDIKGANILVDPNGEIKLADFG 368
Query: 534 LSRLKDTTLLTTKSGRGTPQWMAPEILRNEPS-NEKSDVYSYGVVLWELMTQSIPWENLN 592
+S+ + + S +G+P WMAPE++ N D+ S G + E+ T PW
Sbjct: 369 MSK-HINSAASMLSFKGSPYWMAPEVVMNTNGYGLPVDISSLGCTILEMATSKPPWSQFE 545
Query: 593 SLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
+ + +G ++PE L + I C +R
Sbjct: 546 GVAAIFKIGNSKDMPEIPEHLSDDAKNFIKQCLQR 650
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 107 bits (267), Expect = 6e-24
Identities = 78/235 (33%), Positives = 124/235 (52%), Gaps = 8/235 (3%)
Frame = +2
Query: 396 IGQGSYAVVYHG--IWNGSDVAIKVYFGNGYTEETLQDY-KKEIDIMKRLRHPNVLLFMG 452
+G+G+ A VY I +G VAIKV +E + D K+EI IM+ +RHPN++
Sbjct: 56 LGKGTLAKVYFAKEITSGEGVAIKVMSKARIKKEGMMDQIKREISIMRLVRHPNIVNLKE 235
Query: 453 AVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHR 512
+ + ++ + E + G LF + K D+ RR L A ++Y H R + HR
Sbjct: 236 VMATKTKIFFIMEYIRGGELFAKVAKGKLKDDLARRYFQQLISA--VDYCHSRG--VSHR 403
Query: 513 DLKSSNLLVDKNWTVKVGDFGLS----RLKDTTLLTTKSGRGTPQWMAPEILRNEP-SNE 567
DLK NLL+D+N +KV DFGLS +L+ LL T+ GTP ++APE+LR +
Sbjct: 404 DLKPENLLLDENENLKVSDFGLSGLPEQLRQDGLLHTQC--GTPAYVAPEVLRKKGYDGF 577
Query: 568 KSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIIN 622
K+D +S GV+L+ L+ +P+++ N + + V + P P +I+
Sbjct: 578 KTDTWSCGVILYALLAGCLPFQHENLMTMYNKV--LRAEFQFPPWFSPESKKLIS 736
>TC17194 UP|CAE02590 (CAE02590) Nod-factor receptor 1b, complete
Length = 2193
Score = 107 bits (266), Expect = 8e-24
Identities = 71/196 (36%), Positives = 113/196 (57%), Gaps = 4/196 (2%)
Frame = +1
Query: 392 LRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFM 451
L ++IGQG + VY+ G AIK TE + E+ ++ + H N++ +
Sbjct: 1093 LDNKIGQGGFGAVYYAELRGKKTAIKKMDVQASTE-----FLCELKVLTHVHHLNLVRLI 1257
Query: 452 GAVYSLE-RLAIVTELLPRGSLFKTLHKSN-QTLDIRRRLRMALDIARGMNYLHHRNPPI 509
G Y +E L +V E + G+L + LH S + L R+++ALD ARG+ Y+H P+
Sbjct: 1258 G--YCVEGSLFLVYEHIDNGNLGQYLHGSGKEPLPWSSRVQIALDAARGLEYIHEHTVPV 1431
Query: 510 -VHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGR-GTPQWMAPEILRNEPSNE 567
+HRD+KS+N+L+DKN KV DFGL++L + T ++ GT +M PE + +
Sbjct: 1432 YIHRDVKSANILIDKNLRGKVADFGLTKLIEVGNSTLQTRLVGTFGYMPPEYAQYGDISP 1611
Query: 568 KSDVYSYGVVLWELMT 583
K DVY++GVVL+EL++
Sbjct: 1612 KIDVYAFGVVLFELIS 1659
>TC14769 UP|Q8LKX1 (Q8LKX1) Receptor-like kinase SYMRK, complete
Length = 3495
Score = 106 bits (265), Expect = 1e-23
Identities = 65/199 (32%), Positives = 112/199 (55%), Gaps = 7/199 (3%)
Frame = +3
Query: 396 IGQGSYAVVYHGIWN-GSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAV 454
IG+G + VY G N G +VA+KV + + + +++ E++++ ++H N++ +G
Sbjct: 2181 IGEGGFGSVYRGTLNDGQEVAVKVR--SSTSTQGTREFDNELNLLSAIQHENLVPLLGYC 2354
Query: 455 YSLERLAIVTELLPRGSLFKTLH---KSNQTLDIRRRLRMALDIARGMNYLH-HRNPPIV 510
++ +V + GSL L+ + LD RL +AL ARG+ YLH ++
Sbjct: 2355 NESDQQILVYPFMSNGSLQDRLYGEPAKRKILDWPTRLSIALGAARGLAYLHTFPGRSVI 2534
Query: 511 HRDLKSSNLLVDKNWTVKVGDFGLSRL--KDTTLLTTKSGRGTPQWMAPEILRNEPSNEK 568
HRD+KSSN+L+D + KV DFG S+ ++ + RGT ++ PE + + +EK
Sbjct: 2535 HRDIKSSNILLDHSMCAKVADFGFSKYAPQEGDSYVSLEVRGTAGYLDPEYYKTQQLSEK 2714
Query: 569 SDVYSYGVVLWELMTQSIP 587
SDV+S+GVVL E+++ P
Sbjct: 2715 SDVFSFGVVLLEIVSGREP 2771
>BI420143
Length = 509
Score = 106 bits (265), Expect = 1e-23
Identities = 63/169 (37%), Positives = 105/169 (61%), Gaps = 5/169 (2%)
Frame = +2
Query: 398 QGSYAVVYHGIWNGSDVAIKVYFG--NGYTEETL--QDYKKEIDIMKRLRHPNVLLFMGA 453
QG++ +Y G +NG DVAIK+ N + L Q +++E+ ++ L+HPN++ F+GA
Sbjct: 11 QGAFGKLYRGTYNGEDVAIKILERPENDPAKAQLMEQQFQQEVMMLATLKHPNIVRFIGA 190
Query: 454 VYSLERLAIVTELLPRGSLFKTLHK-SNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHR 512
IVTE GS+ + L K N+++ ++ ++ ALD+ARGM Y+H ++HR
Sbjct: 191 CRKPMVWCIVTEYAKGGSVRQFLTKRQNRSVPLKLAVKQALDVARGMAYVHGLG--LIHR 364
Query: 513 DLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILR 561
DLKS NLL+ + ++K+ DFG++R++ T T GT +WMAPE+++
Sbjct: 365 DLKSDNLLIFGDKSIKIADFGVARIEVQTEGMTPE-TGTYRWMAPEMIQ 508
>TC14667 homologue to UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1591
Score = 106 bits (264), Expect = 1e-23
Identities = 72/204 (35%), Positives = 107/204 (52%), Gaps = 12/204 (5%)
Frame = +2
Query: 396 IGQGSYAVVYHGIWN-GSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAV 454
IG+GSY VY+ N G+ VA+K + ET ++ ++ ++ RL++ N + G
Sbjct: 377 IGEGSYGRVYYATLNDGNAVAVKKLDVSS-EPETNNEFLTQVSMVSRLKNDNFVELHGYC 553
Query: 455 YSLERLAIVTELLPRGSLFKTLH--------KSNQTLDIRRRLRMALDIARGMNYLHHR- 505
+ E GSL LH + TLD +R+R+A+D ARG+ YLH +
Sbjct: 554 VEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDWIQRVRIAVDAARGLEYLHEKV 733
Query: 506 NPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNE 563
P I+HRD++SSN+L+ +++ K+ DF LS S R GT + APE
Sbjct: 734 QPAIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAARLHSTRVLGTFGYHAPEYAMTG 913
Query: 564 PSNEKSDVYSYGVVLWELMTQSIP 587
+KSDVYS+GVVL EL+T P
Sbjct: 914 QLTQKSDVYSFGVVLLELLTGRKP 985
>TC8971 similar to GB|AAM16251.1|20334780|AY093990 At2g11520/F14P14.15
{Arabidopsis thaliana;}, partial (37%)
Length = 1079
Score = 104 bits (259), Expect = 5e-23
Identities = 57/161 (35%), Positives = 98/161 (60%), Gaps = 6/161 (3%)
Frame = +3
Query: 435 EIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHK-SNQTLDIRRRLRMAL 493
E++++ ++ H N++ +G + ++TE +P G+L + L + LD +RL +A+
Sbjct: 6 EVELLAKIDHRNLVKLLGFIDKGNERILITEYVPNGTLREHLDGLRGKILDFNQRLEIAI 185
Query: 494 DIARGMNYLH-HRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRL----KDTTLLTTKSG 548
D+A G+ YLH + I+HRD+KSSN+L+ ++ KV DFG +RL D T ++TK
Sbjct: 186 DVAHGLTYLHLYAEKQIIHRDVKSSNILLTESMRAKVADFGFARLGPVNGDQTHISTKV- 362
Query: 549 RGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWE 589
+GT ++ PE ++ KSDVYS+G++L E++T P E
Sbjct: 363 KGTVGYLDPEYMKTHQLTPKSDVYSFGILLLEILTGRRPVE 485
>TC18848 weakly similar to UP|Q9SSQ6 (Q9SSQ6) F6D8.24 protein, partial (64%)
Length = 969
Score = 102 bits (254), Expect = 2e-22
Identities = 65/205 (31%), Positives = 116/205 (55%), Gaps = 7/205 (3%)
Frame = +2
Query: 394 DEIGQGSYAVVYHG-IWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMG 452
+++G+G + VY G +G +A+K + ++ E++++ R+RH N+L G
Sbjct: 314 NKLGEGGFGSVYWGRTSDGLQIAVKKL--KAMNSKAEMEFAVEVEVLGRVRHKNLLGLRG 487
Query: 453 AVYSLERLAIVTELLPRGSLFKTLHKS---NQTLDIRRRLRMALDIARGMNYLHHR-NPP 508
++ IV + +P SL LH L+ ++R+++A+ A G+ YLHH P
Sbjct: 488 YCVGDDQRLIVYDYMPNLSLLSHLHGQFAVEVQLNWQKRMKIAIGSAEGILYLHHEVTPH 667
Query: 509 IVHRDLKSSNLLVDKNWTVKVGDFGLSRL--KDTTLLTTKSGRGTPQWMAPEILRNEPSN 566
I+HRD+K+SN+L++ ++ V DFG ++L + + +TT+ +GT ++APE +
Sbjct: 668 IIHRDIKASNVLLNSDFEPLVADFGFAKLIPEGVSHMTTRV-KGTLGYLAPEYAMWGKVS 844
Query: 567 EKSDVYSYGVVLWELMTQSIPWENL 591
E DVYS+G++L EL+T P E L
Sbjct: 845 ESCDVYSFGILLLELVTGRKPIEKL 919
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.132 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,071,353
Number of Sequences: 28460
Number of extensions: 165088
Number of successful extensions: 1283
Number of sequences better than 10.0: 233
Number of HSP's better than 10.0 without gapping: 1129
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1147
length of query: 697
length of database: 4,897,600
effective HSP length: 97
effective length of query: 600
effective length of database: 2,136,980
effective search space: 1282188000
effective search space used: 1282188000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0304.15


