

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0302a.8
(578 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
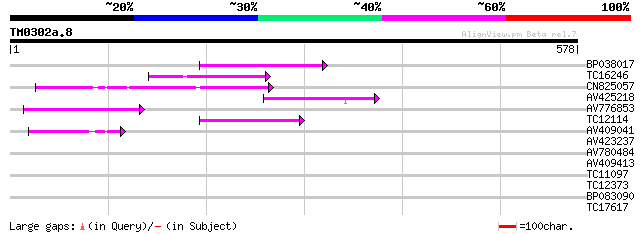
Score E
Sequences producing significant alignments: (bits) Value
BP038017 109 1e-24
TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired re... 91 5e-19
CN825057 88 3e-18
AV425218 85 3e-17
AV776853 83 1e-16
TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transp... 78 3e-15
AV409041 42 4e-04
AV423237 37 0.011
AV780484 30 1.1
AV409413 27 7.0
TC11097 similar to UP|P93520 (P93520) Calcium/calmodulin-depende... 27 7.0
TC12373 27 7.0
BP083090 27 7.0
TC17617 weakly similar to UP|Q84SA8 (Q84SA8) Laccase (Fragment),... 27 7.0
>BP038017
Length = 570
Score = 109 bits (273), Expect = 1e-24
Identities = 49/131 (37%), Positives = 78/131 (59%)
Frame = +2
Query: 194 DAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGK 253
DA+ ++Y + + K+P Y +D+ + +FW+D S+ Y FGD V FD Y
Sbjct: 167 DAQNLLNYFKKMQGKNPGFYYAIQLDDENRMINVFWADARSRSAYNYFGDAVIFDTMYRP 346
Query: 254 NRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDK 313
N+Y+ PF F GVNHH ++ +F AL+ +E ++ WL +L AM + P+ + TD D+
Sbjct: 347 NQYQVPFAPFTGVNHHGQNVLFGCALLLDESESSFTWLFRTWLSAMNDRPPVSITTDQDR 526
Query: 314 AMKAAIKQVFP 324
A++AA+ QVFP
Sbjct: 527 AIQAAVAQVFP 559
>TC16246 weakly similar to UP|Q8S2M2 (Q8S2M2) Far-red impaired response
protein-like, partial (4%)
Length = 660
Score = 90.9 bits (224), Expect = 5e-19
Identities = 44/125 (35%), Positives = 71/125 (56%)
Frame = +2
Query: 142 NNMSRSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISY 201
+ M+ G+ I GG++ + F+K ++ N + K++ + DA A+SY
Sbjct: 2 DGMNLYGVRACHIMALMLGPKGGHESLGFTKTDLSN---HIAKPKRERIQNGDAAAALSY 172
Query: 202 IRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFV 261
+ G DP+ + + D +L+ LFW DG+S+++Y VFGDV+AFD+TY KN+Y P V
Sbjct: 173 LEGKADNDPMFFYKFTKTGDESLENLFWCDGVSRMDYNVFGDVIAFDSTYKKNKYNKPLV 352
Query: 262 VFYGV 266
VF V
Sbjct: 353 VFLHV 367
>CN825057
Length = 721
Score = 88.2 bits (217), Expect = 3e-18
Identities = 66/244 (27%), Positives = 108/244 (44%), Gaps = 1/244 (0%)
Frame = +1
Query: 27 AYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKRQRESRTD 85
AY FY Y GF K + K I F C + G+ + ++R +TD
Sbjct: 1 AYSFYQEYAKSMGFTTSIKNSRRSKKTKEFIDAKFACSRYGVTPESDGGSNRRSSVKKTD 180
Query: 86 FRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMS 145
C A V K +G+W F EHNHEL + HR + ++ +M+ +
Sbjct: 181 ----CKACMHVK-RKPDGKWIIHEFIKEHNHELLP-ALAYHFRIHRNVKLAEKNNMDILH 342
Query: 146 RSGISTPQIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISYIRGL 205
T +++ + Q+GG + G++ + + + + E DA+ + Y + +
Sbjct: 343 AVSERTRKMYVEMSRQSGGCLNIESLVGDLNDQFKKGQYLAMDEG---DAQVMLEYFKHI 513
Query: 206 GSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYG 265
++P + N++ L +FW D S +Y F DVV+FD +Y K+ K PF F G
Sbjct: 514 QKENPNFFYSIDLNEEQRLRNIFWIDAKSINDYLSFNDVVSFDTSYIKSNEKLPFAPFVG 693
Query: 266 VNHH 269
VNHH
Sbjct: 694 VNHH 705
>AV425218
Length = 419
Score = 85.1 bits (209), Expect = 3e-17
Identities = 42/123 (34%), Positives = 64/123 (51%), Gaps = 4/123 (3%)
Frame = +2
Query: 259 PFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAA 318
P GVNHH +S +F AL+S+E E++VWL + L M G P +ITD +AMK A
Sbjct: 5 PLATLVGVNHHGQSVLFGCALLSSEDSESFVWLFQSLLHCMSGVPPQGIITDHSEAMKKA 184
Query: 319 IKQVFPNAHHRLCAWHILDNARK----HVHIRAFNTELRKCMFMELDVSEFEEHWAATVA 374
I+ V P+ HR C +I+ + + + L+ ++ + + EFE +W V
Sbjct: 185 IETVLPSTRHRWCLSYIMKKLPQKLLGYAQYESIRHHLQNVVYDAVVIDEFERNWKKIVE 364
Query: 375 KFG 377
FG
Sbjct: 365 DFG 373
>AV776853
Length = 625
Score = 83.2 bits (204), Expect = 1e-16
Identities = 45/123 (36%), Positives = 61/123 (49%)
Frame = +1
Query: 15 DLMKYDFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLVIQRNFVCHKEGLREDRGLT 74
D+ DF E AY FY Y GF VRK + G V R FVC+++GLR +
Sbjct: 226 DIRHLDFGSEEEAYQFYQAYAKYQGFIVRKDDIGRDYHGNVNMRQFVCNRQGLRSKKHYN 405
Query: 75 MDKRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMN 134
R+R+ + C AK RVH+D G+W F + HNHEL + I +R MN
Sbjct: 406 RTDRKRDHKPVTHTNCLAKLRVHLDYKIGKWKVVSFEECHNHELTPARFVHFIPPYRVMN 585
Query: 135 ESD 137
++D
Sbjct: 586 DAD 594
>TC12114 weakly similar to GB|BAC16510.1|23237937|AP005198 transposase-like
{Oryza sativa (japonica cultivar-group);}, partial (8%)
Length = 488
Score = 78.2 bits (191), Expect = 3e-15
Identities = 38/107 (35%), Positives = 59/107 (54%)
Frame = +1
Query: 194 DAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGK 253
+A + + Y + ++ Y + +D+ + +FW D ++Y FGD+V+ D+TY
Sbjct: 43 EAGYILQYFQRKLVENSPFYHAYQLDDEDQITNVFWVDARMLIDYGYFGDMVSLDSTYCT 222
Query: 254 NRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMK 300
+ P VF G NHH K+ IF AL+ +E E+Y WL E FLEA K
Sbjct: 223 HSSNRPLAVFSGFNHHRKAVIFGAALLYDETTESY*WLFESFLEAHK 363
>AV409041
Length = 349
Score = 41.6 bits (96), Expect = 4e-04
Identities = 29/100 (29%), Positives = 41/100 (41%), Gaps = 1/100 (1%)
Frame = +3
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGL-VIQRNFVCHKEGLREDRGLTMDKR 78
+F E AY FY Y GF K ++ I F C + G ++ ++ R
Sbjct: 48 EFESHEAAYAFYKEYAKSAGFGTAKLSSRRSRASKEFIDAKFSCIRYGNKQQSDDAINPR 227
Query: 79 QRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
+ GC A V + +G+WY F EHNHEL
Sbjct: 228 PSP-----KIGCKASMHVK-RRQDGKWYVYSFVKEHNHEL 329
>AV423237
Length = 310
Score = 36.6 bits (83), Expect = 0.011
Identities = 20/62 (32%), Positives = 32/62 (51%)
Frame = +2
Query: 230 SDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYV 289
+D S++NY FGD V FD T + + +G + I+ AL++NE ++V
Sbjct: 14 ADATSRMNYSYFGDAVIFDTTI--DNHIESICSSWG*SSWATCVIWLVALIANESESSFV 187
Query: 290 WL 291
WL
Sbjct: 188 WL 193
>AV780484
Length = 529
Score = 30.0 bits (66), Expect = 1.1
Identities = 14/38 (36%), Positives = 21/38 (54%)
Frame = -1
Query: 223 NLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPF 260
+L +FW D +L+Y+ +V Y KN+YK PF
Sbjct: 124 HLTSVFWVDIKGRLDYETSMMLVLIHTHYLKNKYKIPF 11
>AV409413
Length = 422
Score = 27.3 bits (59), Expect = 7.0
Identities = 20/77 (25%), Positives = 36/77 (45%), Gaps = 1/77 (1%)
Frame = +2
Query: 118 LEDDKMCGMIAAHRKMNESDIMHMNNMSRS-GISTPQIHNTFASQTGGYQKVRFSKGNMY 176
+ +D++ G + + K+ E +I+ + RS G N T + GNM
Sbjct: 23 VSEDELKGFFSKYGKVVEHEIIRDHTTKRSRGFGFIVFDNEKVVDT------ILTDGNMI 184
Query: 177 NVQGRQRRMKKKEPEKT 193
++ G Q +KK EP+K+
Sbjct: 185 DMAGTQVEIKKAEPKKS 235
>TC11097 similar to UP|P93520 (P93520) Calcium/calmodulin-dependent protein
kinase homolog|CaM kinase homolog|MCK1, partial (36%)
Length = 949
Score = 27.3 bits (59), Expect = 7.0
Identities = 19/74 (25%), Positives = 33/74 (43%)
Frame = -1
Query: 88 CGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRS 147
C CT F++H+ + G+W+ ++ D N L C H ++ + D+ S
Sbjct: 556 CSCTKFFKIHL--SIGKWFHHMY-DIKNP*LPHGISCMTCKGHFEIIKGDVPISIRFQ*S 386
Query: 148 GISTPQIHNTFASQ 161
+ P+IH F Q
Sbjct: 385 EL-RPKIHQLFLCQ 347
>TC12373
Length = 450
Score = 27.3 bits (59), Expect = 7.0
Identities = 15/44 (34%), Positives = 24/44 (54%)
Frame = +3
Query: 111 SDEHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRSGISTPQI 154
S+ N + + M G MN S IMH+++ SR+ ++TP I
Sbjct: 60 SERANRLIVERFMIGSSEYMHPMNFSTIMHLDSTSRT*LATPLI 191
>BP083090
Length = 465
Score = 27.3 bits (59), Expect = 7.0
Identities = 18/60 (30%), Positives = 25/60 (41%)
Frame = -3
Query: 336 LDNARKHVHIRAFNTELRKCMFMELDVSEFEEHWAATVAKFGWKKILGLRNYMKEE*CGL 395
LD +H H+ F + R C L + + + AA + F WK G EE GL
Sbjct: 304 LDLRGQHFHLIPFGSGRRGCPGTSLALQVVQTNLAAMIQCFEWKVSGGKERVDMEEKLGL 125
>TC17617 weakly similar to UP|Q84SA8 (Q84SA8) Laccase (Fragment), partial
(53%)
Length = 1139
Score = 27.3 bits (59), Expect = 7.0
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +2
Query: 145 SRSGISTPQIHNTFASQTGGYQKVRFSKGN 174
++ + PQI NT A GG+ +RF N
Sbjct: 737 AKFNLVNPQIRNTIAVPVGGWAVIRFQANN 826
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.149 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,190,319
Number of Sequences: 28460
Number of extensions: 138238
Number of successful extensions: 893
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 891
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 892
length of query: 578
length of database: 4,897,600
effective HSP length: 95
effective length of query: 483
effective length of database: 2,193,900
effective search space: 1059653700
effective search space used: 1059653700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0302a.8


