

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.4
(147 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
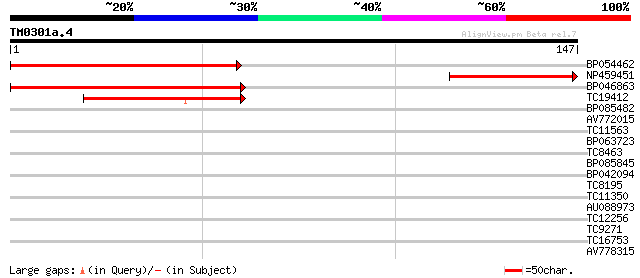
Score E
Sequences producing significant alignments: (bits) Value
BP054462 75 6e-15
NP459451 NDX3 protein [Lotus japonicus] 59 4e-10
BP046863 54 1e-08
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 46 2e-06
BP085482 37 0.002
AV772015 34 0.009
TC11563 33 0.020
BP063723 33 0.020
TC8463 similar to UP|CFI_VITVI (P51117) Chalcone--flavonone isom... 28 0.64
BP085845 27 1.4
BP042094 27 1.9
TC8195 similar to UP|Q7SEW0 (Q7SEW0) Predicted protein, partial ... 25 5.4
TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3,... 25 5.4
AU088973 25 7.1
TC12256 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_12... 25 7.1
TC9271 weakly similar to UP|Q9SFG3 (Q9SFG3) F2O10.5 protein (At3... 24 9.3
TC16753 similar to UP|Q9SIA7 (Q9SIA7) Legumin-like protein, part... 24 9.3
AV778315 24 9.3
>BP054462
Length = 422
Score = 74.7 bits (182), Expect = 6e-15
Identities = 35/60 (58%), Positives = 43/60 (71%)
Frame = +1
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL R S+ + +L+ YYGPYPI+ +IGAVAYRL+LP RVHPVFH SLLK
Sbjct: 241 VFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAKIGAVAYRLELPAHSRVHPVFHVSLLK 420
>NP459451 NDX3 protein [Lotus japonicus]
Length = 665
Score = 58.5 bits (140), Expect = 4e-10
Identities = 26/33 (78%), Positives = 29/33 (87%)
Frame = +3
Query: 115 DEPTWKDTLNIRSQFPVFNLEDKVDLSAGGIVR 147
DE TW+D + I+SQFP FNLEDKVDLSAGGIVR
Sbjct: 492 DEATWEDNITIKSQFPSFNLEDKVDLSAGGIVR 590
>BP046863
Length = 580
Score = 53.9 bits (128), Expect = 1e-08
Identities = 25/61 (40%), Positives = 39/61 (62%)
Frame = +1
Query: 1 VFIKLMALRENSVVTRDCPQLTAPYYGPYPIIQRIGAVAYRLQLPEGGRVHPVFHASLLK 60
VF+KL + ++ R +L+ +YGP+ +++R+ VAY L L RVHPVFH SLL+
Sbjct: 397 VFLKLQPYKLQNLAQRKNQKLSPRFYGPFKVLERVVQVAY*LDLXSESRVHPVFHLSLLE 576
Query: 61 E 61
+
Sbjct: 577 K 579
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 46.2 bits (108), Expect = 2e-06
Identities = 19/43 (44%), Positives = 33/43 (76%), Gaps = 1/43 (2%)
Frame = +3
Query: 20 QLTAPYYGPYPIIQRIGAVAYRLQL-PEGGRVHPVFHASLLKE 61
+L+ + GP+ +++R+G+V+YRL L P+ VHPVFH S+L++
Sbjct: 48 KLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRK 176
>BP085482
Length = 365
Score = 36.6 bits (83), Expect = 0.002
Identities = 17/17 (100%), Positives = 17/17 (100%)
Frame = -1
Query: 131 VFNLEDKVDLSAGGIVR 147
VFNLEDKVDLSAGGIVR
Sbjct: 365 VFNLEDKVDLSAGGIVR 315
>AV772015
Length = 456
Score = 34.3 bits (77), Expect = 0.009
Identities = 17/25 (68%), Positives = 19/25 (76%)
Frame = -1
Query: 123 LNIRSQFPVFNLEDKVDLSAGGIVR 147
L I+ QFP FNLEDKV L+ GGI R
Sbjct: 423 LVIQMQFPDFNLEDKVGLARGGIDR 349
>TC11563
Length = 470
Score = 33.1 bits (74), Expect = 0.020
Identities = 19/45 (42%), Positives = 24/45 (53%)
Frame = +2
Query: 102 LPQVWIQWQGKPADEPTWKDTLNIRSQFPVFNLEDKVDLSAGGIV 146
+PQ+ IQW+G A TW+ I+ FP F L DKV G V
Sbjct: 8 VPQLLIQWEG--AANCTWELLSYIQDSFPQFALADKVTFYGEGNV 136
>BP063723
Length = 479
Score = 33.1 bits (74), Expect = 0.020
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = -3
Query: 107 IQWQGKPADEPTWKDTLNIRSQFPVFNLEDKVDLSAG 143
I+W+ P E +W+D + FP LED+++L G
Sbjct: 477 IRWKDLPTFEDSWEDFCKLLDPFPNHQLEDQLNLQGG 367
>TC8463 similar to UP|CFI_VITVI (P51117) Chalcone--flavonone isomerase
(Chalcone isomerase) , partial (89%)
Length = 1042
Score = 28.1 bits (61), Expect = 0.64
Identities = 9/21 (42%), Positives = 16/21 (75%)
Frame = +2
Query: 98 QESTLPQVWIQWQGKPADEPT 118
Q++ +P + ++W+GKP DE T
Sbjct: 203 QDTAVPSLAVKWKGKPVDELT 265
>BP085845
Length = 464
Score = 26.9 bits (58), Expect = 1.4
Identities = 23/104 (22%), Positives = 44/104 (42%), Gaps = 5/104 (4%)
Frame = -2
Query: 27 GPYPIIQRIGAVAY----RLQLPEGGRVHPVFHASLLKEAVGNNSVELQLLDHLTGEEVA 82
GP+ I++ + +AY L L H V H +LL + + S + GE +
Sbjct: 391 GPFKILEMVCLIAY*LTHSLYLLAA---HIVLHVTLLWRNLYDQSQNICHEGVXLGEHWS 221
Query: 83 SV-HPFSVITSRFTTRQESTLPQVWIQWQGKPADEPTWKDTLNI 125
+ HP ++ + + + V + W+G E +W+ +I
Sbjct: 220 HMEHPIVMVDMKVRCMRPKNIDDVKVIWRGLSG*EKSWEYVTSI 89
>BP042094
Length = 409
Score = 26.6 bits (57), Expect = 1.9
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = -1
Query: 118 TWKDTLNIRSQFPVFNLEDKVDLSAG 143
+W D ++ QFP F+LEDK G
Sbjct: 406 SWVDEPALKCQFPSFSLEDKAAAIGG 329
>TC8195 similar to UP|Q7SEW0 (Q7SEW0) Predicted protein, partial (3%)
Length = 933
Score = 25.0 bits (53), Expect = 5.4
Identities = 16/48 (33%), Positives = 23/48 (47%)
Frame = -1
Query: 70 LQLLDHLTGEEVASVHPFSVITSRFTTRQESTLPQVWIQWQGKPADEP 117
LQ + T E+ SV P TS F + S + +G+PAD+P
Sbjct: 672 LQFAIYPTLPEMQSVRPSRKHTSGFNGTKRSQAGGTSTKERGEPADQP 529
>TC11350 similar to UP|Q9LJP7 (Q9LJP7) Genomic DNA, chromosome 3, P1 clone:
MIG10, partial (18%)
Length = 1118
Score = 25.0 bits (53), Expect = 5.4
Identities = 11/26 (42%), Positives = 16/26 (61%)
Frame = +3
Query: 47 GGRVHPVFHASLLKEAVGNNSVELQL 72
G +HP+ H + LK +GN +LQL
Sbjct: 474 GRSIHPISHKTSLKFKLGN*IYKLQL 551
>AU088973
Length = 173
Score = 24.6 bits (52), Expect = 7.1
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +2
Query: 81 VASVHPFSVITSRFTTRQESTLPQVWIQWQ 110
+A+ + FSV TSR T Q+ +P + QW+
Sbjct: 11 IAAKNKFSVSTSR-NTXQQQKIPLLGYQWR 97
>TC12256 weakly similar to UP|Q9ASZ6 (Q9ASZ6) AT4g35320/F23E12_120, partial
(33%)
Length = 640
Score = 24.6 bits (52), Expect = 7.1
Identities = 11/26 (42%), Positives = 13/26 (49%), Gaps = 1/26 (3%)
Frame = -1
Query: 94 FTTRQESTLPQVWIQWQG-KPADEPT 118
F R L Q+W W+G KP PT
Sbjct: 241 FPDRSIQDLWQIWASWRGSKPRSSPT 164
>TC9271 weakly similar to UP|Q9SFG3 (Q9SFG3) F2O10.5 protein (At3g05990),
partial (32%)
Length = 1035
Score = 24.3 bits (51), Expect = 9.3
Identities = 12/38 (31%), Positives = 21/38 (54%)
Frame = -2
Query: 94 FTTRQESTLPQVWIQWQGKPADEPTWKDTLNIRSQFPV 131
F T+ +S+ PQ+W QG D+ + + L + FP+
Sbjct: 581 FLTKGKSS-PQIWKIPQGAGGDKKLFPEVLRLSPGFPM 471
>TC16753 similar to UP|Q9SIA7 (Q9SIA7) Legumin-like protein, partial (42%)
Length = 534
Score = 24.3 bits (51), Expect = 9.3
Identities = 15/46 (32%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Frame = -2
Query: 66 NSVELQLLDHLTGEE-VASVHPFSVITSRFTTRQESTLPQVWIQWQ 110
NSV L H+T + A PF + T+ F++ ST+P + W+
Sbjct: 401 NSVSSLLYHHVTTPKGSAKASPFLIATT-FSSDSGSTIPTTPVPWR 267
>AV778315
Length = 559
Score = 24.3 bits (51), Expect = 9.3
Identities = 11/32 (34%), Positives = 16/32 (49%)
Frame = -3
Query: 107 IQWQGKPADEPTWKDTLNIRSQFPVFNLEDKV 138
+QW G +P W +T S+F F + D V
Sbjct: 548 LQWHGHFPGKPDWSET----SRFVAFTMLDSV 465
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,559,999
Number of Sequences: 28460
Number of extensions: 32683
Number of successful extensions: 151
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 150
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 150
length of query: 147
length of database: 4,897,600
effective HSP length: 82
effective length of query: 65
effective length of database: 2,563,880
effective search space: 166652200
effective search space used: 166652200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0301a.4


