

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.2
(80 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
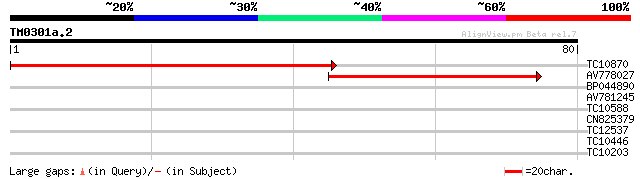
TC10870 similar to GB|CAA77438.1|2924285|CHNTXX Ycf2 protein {Ni... 91 3e-20
AV778027 55 3e-09
BP044890 25 2.0
AV781245 24 5.8
TC10588 similar to UP|Q9LPU4 (Q9LPU4) T22I11.8, partial (36%) 24 5.8
CN825379 23 7.5
TC12537 weakly similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-r... 23 7.5
TC10446 similar to UP|Q8GTR1 (Q8GTR1) Geranylgeranyl diphosphate... 23 9.8
TC10203 similar to UP|Q7X833 (Q7X833) Cytosolic phosphoglucose i... 23 9.8
>TC10870 similar to GB|CAA77438.1|2924285|CHNTXX Ycf2 protein {Nicotiana
tabacum;}, complete
Length = 6897
Score = 91.3 bits (225), Expect = 3e-20
Identities = 44/46 (95%), Positives = 44/46 (95%)
Frame = +1
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLIHRQRWLR NNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD
Sbjct: 6670 FELLIHRQRWLRTNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 6807
>AV778027
Length = 505
Score = 54.7 bits (130), Expect = 3e-09
Identities = 29/30 (96%), Positives = 30/30 (99%)
Frame = +2
Query: 46 DNTKMK*IIENSRGR*GSR*RQMSRLDILA 75
+NTKMK*IIENSRGR*GSR*RQMSRLDILA
Sbjct: 194 NNTKMK*IIENSRGR*GSR*RQMSRLDILA 283
>BP044890
Length = 558
Score = 25.4 bits (54), Expect = 2.0
Identities = 13/45 (28%), Positives = 27/45 (59%), Gaps = 4/45 (8%)
Frame = +2
Query: 10 WLRPNNSLSNGFFRSNT----LSESYQYLSNMFLSNGTLLDNTKM 50
W RP+ + FFR++T S S+ +L + F+S+ ++L ++ +
Sbjct: 215 WERPSLIIFKPFFRNSTPLLIPSRSFLFLFDGFMSSTSMLSHSSL 349
>AV781245
Length = 523
Score = 23.9 bits (50), Expect = 5.8
Identities = 8/24 (33%), Positives = 16/24 (66%)
Frame = +2
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRS 24
F+ L+H++ W+ PN +++ RS
Sbjct: 389 FISLLHQE*WVSPNKPVASPLSRS 460
>TC10588 similar to UP|Q9LPU4 (Q9LPU4) T22I11.8, partial (36%)
Length = 976
Score = 23.9 bits (50), Expect = 5.8
Identities = 13/45 (28%), Positives = 23/45 (50%), Gaps = 5/45 (11%)
Frame = +3
Query: 1 FLLLIHRQRWLRPN-----NSLSNGFFRSNTLSESYQYLSNMFLS 40
+ LL+H Q+WL N + GFF S++ + ++ S + S
Sbjct: 282 WFLLLHLQQWLAQNMPKFSEASYKGFFSSSSTTTRFRSDSRTYSS 416
>CN825379
Length = 646
Score = 23.5 bits (49), Expect = 7.5
Identities = 9/27 (33%), Positives = 17/27 (62%)
Frame = -2
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTL 27
++L + R+RWL+ ++ + FFR L
Sbjct: 174 WVLRLRRRRWLQSEPTVESPFFRRRWL 94
>TC12537 weakly similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich
protein, partial (8%)
Length = 444
Score = 23.5 bits (49), Expect = 7.5
Identities = 14/37 (37%), Positives = 21/37 (55%), Gaps = 1/37 (2%)
Frame = +2
Query: 14 NNSLSNGF-FRSNTLSESYQYLSNMFLSNGTLLDNTK 49
+N S GF FRS+ + SY+ + SNG +D +K
Sbjct: 305 SNCSSKGFDFRSDDVLCSYEDFTEQDSSNGNNIDPSK 415
>TC10446 similar to UP|Q8GTR1 (Q8GTR1) Geranylgeranyl diphosphate synthase,
partial (54%)
Length = 999
Score = 23.1 bits (48), Expect = 9.8
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = -2
Query: 4 LIHRQRWLRPNNSLSNGFFRSNTLSE 29
L+HR+++ NNS+SN R TLS+
Sbjct: 473 LLHRKQYWSSNNSMSN--VR*RTLSK 402
>TC10203 similar to UP|Q7X833 (Q7X833) Cytosolic phosphoglucose isomerase,
partial (43%)
Length = 851
Score = 23.1 bits (48), Expect = 9.8
Identities = 10/17 (58%), Positives = 13/17 (75%), Gaps = 2/17 (11%)
Frame = +3
Query: 6 HRQRWLRPNNS--LSNG 20
H+Q+WL P +S LSNG
Sbjct: 102 HKQKWLHPLSSLTLSNG 152
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.350 0.154 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,121,289
Number of Sequences: 28460
Number of extensions: 10223
Number of successful extensions: 132
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 132
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 132
length of query: 80
length of database: 4,897,600
effective HSP length: 56
effective length of query: 24
effective length of database: 3,303,840
effective search space: 79292160
effective search space used: 79292160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0301a.2


