

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
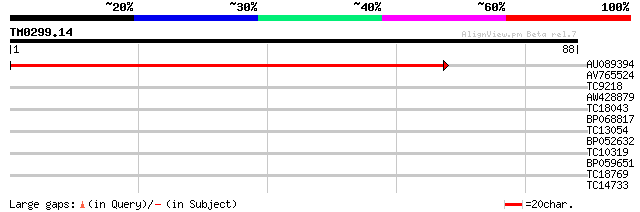
Query= TM0299.14
(88 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AU089394 125 1e-30
AV765524 26 1.1
TC9218 weakly similar to UP|Q9FPS1 (Q9FPS1) Ubiquitin-specific p... 25 1.8
AW428879 25 2.4
TC18043 25 3.1
BP068817 24 4.1
TC13054 similar to UP|NO93_SOYBN (Q02921) Early nodulin 93 (N-93... 24 5.4
BP052632 23 7.0
TC10319 similar to UP|Q8LE60 (Q8LE60) Cytochrome b561, partial (... 23 7.0
BP059651 23 9.1
TC18769 similar to GB|CAB81029.1|7269936|ATCHRIV72 cyclic nucleo... 23 9.1
TC14733 similar to UP|Q41058 (Q41058) Starch branching enzyme I ... 23 9.1
>AU089394
Length = 277
Score = 125 bits (314), Expect = 1e-30
Identities = 60/68 (88%), Positives = 62/68 (90%)
Frame = +3
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
MAEAKTQVE+IRKWV DHKLRTVGCLWL+GITGSIAYNWSRPNMKT V IIHA LHAQAL
Sbjct: 72 MAEAKTQVETIRKWVXDHKLRTVGCLWLTGITGSIAYNWSRPNMKTXVXIIHAXLHAQAL 251
Query: 61 TLAALAGA 68
TLA L GA
Sbjct: 252 TLAXLPGA 275
>AV765524
Length = 600
Score = 26.2 bits (56), Expect = 1.1
Identities = 13/32 (40%), Positives = 16/32 (49%)
Frame = -3
Query: 8 VESIRKWVVDHKLRTVGCLWLSGITGSIAYNW 39
V S+ WV +H L +VGCL L Y W
Sbjct: 247 VHSV*HWVGEHSLGSVGCLILKFCFLC*CYPW 152
>TC9218 weakly similar to UP|Q9FPS1 (Q9FPS1) Ubiquitin-specific protease
26, partial (6%)
Length = 553
Score = 25.4 bits (54), Expect = 1.8
Identities = 10/25 (40%), Positives = 15/25 (60%)
Frame = -1
Query: 29 SGITGSIAYNWSRPNMKTSVKIIHA 53
S I+ S+ WS NM TS ++H+
Sbjct: 88 SWISESLTITWSPANMFTSTSVVHS 14
>AW428879
Length = 384
Score = 25.0 bits (53), Expect = 2.4
Identities = 16/41 (39%), Positives = 21/41 (51%)
Frame = +2
Query: 18 HKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQ 58
H+ VGCLW GS A N + P ++ +KI RL Q
Sbjct: 47 HRESWVGCLWSIWKGGSTAANTAAPILR-YMKISSPRLPLQ 166
>TC18043
Length = 546
Score = 24.6 bits (52), Expect = 3.1
Identities = 15/54 (27%), Positives = 26/54 (47%), Gaps = 3/54 (5%)
Frame = +1
Query: 2 AEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSR---PNMKTSVKIIH 52
A+A+ + + R +D +L VGC + G+ +Y W R N+ S+ H
Sbjct: 178 AKARLEWKEFRTRCLDIELFDVGCYSMCRFHGNPSYRW*RALLSNIFRSISTTH 339
>BP068817
Length = 110
Score = 24.3 bits (51), Expect = 4.1
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = +2
Query: 27 WLSGITGSIAYNWSRPNMKTSVKIIHARL 55
W I+++W+R MK+S IIH L
Sbjct: 2 WFGN*KLDISFDWTRK*MKSSPFIIHKTL 88
>TC13054 similar to UP|NO93_SOYBN (Q02921) Early nodulin 93 (N-93), partial
(93%)
Length = 558
Score = 23.9 bits (50), Expect = 5.4
Identities = 16/48 (33%), Positives = 22/48 (45%)
Frame = +1
Query: 39 WSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRAEAKRAKNS 86
W+R N+ + AQAL ++ +AGAA D A KNS
Sbjct: 229 WARANLNHT---------AQALLISTVAGAAYFIVADKTVLATARKNS 345
>BP052632
Length = 489
Score = 23.5 bits (49), Expect = 7.0
Identities = 8/17 (47%), Positives = 12/17 (70%)
Frame = +2
Query: 14 WVVDHKLRTVGCLWLSG 30
+++ L +GCLWLSG
Sbjct: 92 YLLGKTLPRIGCLWLSG 142
>TC10319 similar to UP|Q8LE60 (Q8LE60) Cytochrome b561, partial (81%)
Length = 1039
Score = 23.5 bits (49), Expect = 7.0
Identities = 17/56 (30%), Positives = 27/56 (47%)
Frame = +2
Query: 20 LRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYD 75
L +G + + G ++Y S P K K+IH LHA AL L + A + ++
Sbjct: 155 LMLIGLIVIGG-EAIVSYK-SLPLKKEVKKLIHLTLHAIALVLGIVGICAAFKNHN 316
>BP059651
Length = 376
Score = 23.1 bits (48), Expect = 9.1
Identities = 10/28 (35%), Positives = 14/28 (49%)
Frame = +2
Query: 3 EAKTQVESIRKWVVDHKLRTVGCLWLSG 30
E T +E+IR W T +WL+G
Sbjct: 170 ENTTGLETIRHWTYAKSCFTPNTVWLTG 253
>TC18769 similar to GB|CAB81029.1|7269936|ATCHRIV72 cyclic nucleotide and
calmodulin-regulated ion channel-like protein
{Arabidopsis thaliana;} , partial (29%)
Length = 646
Score = 23.1 bits (48), Expect = 9.1
Identities = 14/51 (27%), Positives = 25/51 (48%)
Frame = -2
Query: 38 NWSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRAEAKRAKNSEQ 88
NW N+++S + A +ALT+ G V D ++A+ + +S Q
Sbjct: 543 NWLATNLRSSARRAKASTSLRALTVLVEEGKFTV---DFGSKAQESSSSPQ 400
>TC14733 similar to UP|Q41058 (Q41058) Starch branching enzyme I precursor ,
partial (8%)
Length = 1188
Score = 23.1 bits (48), Expect = 9.1
Identities = 9/30 (30%), Positives = 16/30 (53%)
Frame = -2
Query: 21 RTVGCLWLSGITGSIAYNWSRPNMKTSVKI 50
+T+G W S I + + WS + K +K+
Sbjct: 311 KTLGLQWASWILLHLCHGWSHLSDKLLIKV 222
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,503,862
Number of Sequences: 28460
Number of extensions: 17070
Number of successful extensions: 96
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 96
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 96
length of query: 88
length of database: 4,897,600
effective HSP length: 64
effective length of query: 24
effective length of database: 3,076,160
effective search space: 73827840
effective search space used: 73827840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 48 (23.1 bits)
Lotus: description of TM0299.14


