

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296b.2
(1291 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
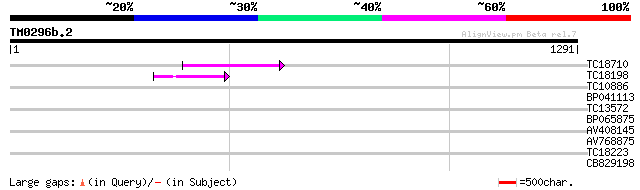
TC18710 155 3e-38
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 105 6e-23
TC10886 similar to UP|Q8GSM8 (Q8GSM8) Squalene monooxygenase 1 ... 37 0.020
BP041113 35 0.099
TC13572 33 0.38
BP065875 30 3.2
AV408145 30 3.2
AV768875 28 9.3
TC18223 28 9.3
CB829198 28 9.3
>TC18710
Length = 843
Score = 155 bits (393), Expect = 3e-38
Identities = 85/233 (36%), Positives = 127/233 (54%), Gaps = 1/233 (0%)
Frame = +2
Query: 394 KEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILV 453
++++R +++ G DG F+++ + V D+V + E ++ + E LV
Sbjct: 137 QKKLRLLFFLLETSQPRGMDGMNGLFYQKNGDVVKDSVT*AILAFLERAEIPNEINETLV 316
Query: 454 VLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLPGRGTMDN 513
LIPKV HP SI +FRPIS C YK+I+K+ V R++ + D++ PMQ+ F+ GR DN
Sbjct: 317 TLIPKVPHPESINQFRPISGCTFLYKVISKIFVARLKDAMPDLISPMQSGFIQGRQIQDN 496
Query: 514 AFLA*EIIHQMHRSMA-RKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCY 572
+ E H ++R A + + K+D+ KAYD + W FL +L + F + +IM
Sbjct: 497 LLIVQEAFHAINRPGALGRNHSIIKLDMNKAYDRVEWKFLESSLLAFGFSTNWVKMIMIL 676
Query: 573 VSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
VS + NG P P RGLRQGDP SPY F+ ME LS+ IQN G
Sbjct: 677 VSGVSYNYKINGVVGPKLLPQRGLRQGDPFSPYLFLFTMEVLSLLIQNSFNMG 835
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 105 bits (261), Expect = 6e-23
Identities = 59/177 (33%), Positives = 101/177 (56%), Gaps = 3/177 (1%)
Frame = -3
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYFQALFAGN--QEASQTPFPGDNVPKL-KEV 383
R N + ++K + G W K+ E A+ YF +++ + Q + P +PKL +
Sbjct: 524 RDFNRILKIKDAHGIWVEGQKKVNEAAVNYFSDIYSSSPLQYLRECLAP---IPKLVTDR 354
Query: 384 VRFLLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGK 443
+ L VS +E+++A+ SM APG DG F++Q E V + V + + F+ G+
Sbjct: 353 LNHKLCAQVSTDEIKEAVFSMGDLKAPGMDGINGLFYQQNREIVKNDVNNAILDFFDHGE 174
Query: 444 VDERLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPM 500
+ L E LV LIPK+ H +I++FRPIS C+ YK+I+K++V R++P + +++ PM
Sbjct: 173 LPVELNETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKIIVARLKPDMVNLISPM 3
>TC10886 similar to UP|Q8GSM8 (Q8GSM8) Squalene monooxygenase 1 , partial
(27%)
Length = 1135
Score = 37.0 bits (84), Expect = 0.020
Identities = 17/57 (29%), Positives = 32/57 (55%)
Frame = -3
Query: 816 WSLNGLYTVCDFDIASRLNRVTPTLSTAQEDQWIWGPGSRGTYLVKEAYSWLHERNR 872
WSL+ L+T+ ++ + + + + T +D W S+G Y+V+E YSW + +R
Sbjct: 281 WSLDWLWTMMP*NMLNLFSHLYLRVVTKLKDLWA*DSRSKGCYMVREFYSWFNSLDR 111
>BP041113
Length = 503
Score = 34.7 bits (78), Expect = 0.099
Identities = 16/28 (57%), Positives = 19/28 (67%)
Frame = +1
Query: 840 LSTAQEDQWIWGPGSRGTYLVKEAYSWL 867
LS ED+W+W PG GTY V AYS+L
Sbjct: 211 LSEGVEDKWLWLPG--GTYTVNSAYSFL 288
>TC13572
Length = 534
Score = 32.7 bits (73), Expect = 0.38
Identities = 21/76 (27%), Positives = 34/76 (44%), Gaps = 12/76 (15%)
Frame = -3
Query: 804 RLEWSDILQHGAWSLNGLYT------------VCDFDIASRLNRVTPTLSTAQEDQWIWG 851
+L+W+ I + G WS G++ D+ A L+ V L D+W+W
Sbjct: 259 KLKWASIDECGGWS-QGVWNWDFRWRRPLAGRELDWLQALSLDIVRVPLLAGVPDKWVWK 83
Query: 852 PGSRGTYLVKEAYSWL 867
P G+Y V A+ +L
Sbjct: 82 PSEDGSYSVNSAFIFL 35
>BP065875
Length = 495
Score = 29.6 bits (65), Expect = 3.2
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = -2
Query: 21 LGFSFAFVMEAVGFSGGIWV 40
+GF + V+EA GF GGIW+
Sbjct: 263 IGFDHSVVVEAEGFVGGIWI 204
>AV408145
Length = 422
Score = 29.6 bits (65), Expect = 3.2
Identities = 15/43 (34%), Positives = 21/43 (47%), Gaps = 2/43 (4%)
Frame = -2
Query: 845 EDQWIWGPGSRGT--YLVKEAYSWLHERNRVSAGAGNWRWLWR 885
ED ++G G + V A +W+ + V G G WRW WR
Sbjct: 175 EDVGVFGDEELGEVHHGVHVAAAWVWYGHHVGGGGGWWRWEWR 47
>AV768875
Length = 232
Score = 28.1 bits (61), Expect = 9.3
Identities = 23/58 (39%), Positives = 26/58 (44%)
Frame = +2
Query: 102 LGSCWGTLTKSYNLMRSEEELSYQAEQQNLQRFLMLVA*WILEWWIAGLLGKGK*TIG 159
LG GT K L + E S E LQ M+V * L W +A LLG K T G
Sbjct: 56 LGF*LGT*MKFCFLHKLGEVSSTLIEPNALQLSWMIVI*LTLAW*VASLLGSAKETTG 229
>TC18223
Length = 601
Score = 28.1 bits (61), Expect = 9.3
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +2
Query: 534 LAFKIDLEKAYDSISWSFLR 553
L+ +DL+ +YD ISW FLR
Sbjct: 308 LSHGVDLQGSYDKISWIFLR 367
>CB829198
Length = 562
Score = 28.1 bits (61), Expect = 9.3
Identities = 9/15 (60%), Positives = 9/15 (60%)
Frame = -2
Query: 870 RNRVSAGAGNWRWLW 884
RNR G G WRW W
Sbjct: 153 RNRTLIGGGRWRWWW 109
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.340 0.149 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,414,896
Number of Sequences: 28460
Number of extensions: 346123
Number of successful extensions: 2582
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 2521
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2571
length of query: 1291
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1190
effective length of database: 2,023,140
effective search space: 2407536600
effective search space used: 2407536600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0296b.2


