

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296a.5
(754 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
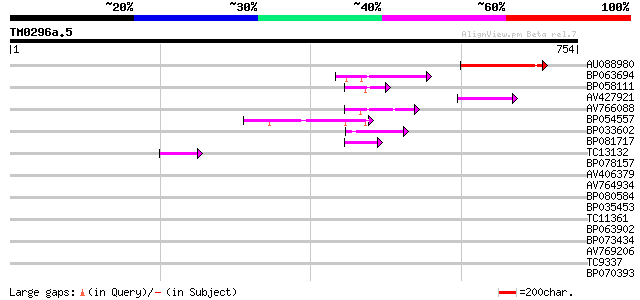
Score E
Sequences producing significant alignments: (bits) Value
AU088980 111 5e-25
BP063694 74 7e-14
BP058111 58 5e-09
AV427921 56 2e-08
AV766088 54 1e-07
BP054557 52 3e-07
BP033602 48 7e-06
BP081717 46 2e-05
TC13132 similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement... 43 2e-04
BP078157 40 0.001
AV406379 39 0.002
AV764934 39 0.002
BP080584 39 0.003
BP035453 32 0.009
TC11361 37 0.009
BP063902 36 0.020
BP073434 35 0.034
AV769206 34 0.098
TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B pre... 33 0.22
BP070393 31 0.83
>AU088980
Length = 360
Score = 111 bits (277), Expect = 5e-25
Identities = 56/116 (48%), Positives = 75/116 (64%)
Frame = +2
Query: 600 NGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFIILHNL 659
N +VERKH+HILN+ RALM S L W+FAV HA L+N LP+ +L P+ L+N
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 660 KPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTREV 715
P + LKVFG+LCY+++LT + K RARKC+ LG K G GY++ DL T +
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKT-GTXGYIVMDLPTTXI 346
>BP063694
Length = 511
Score = 74.3 bits (181), Expect = 7e-14
Identities = 48/136 (35%), Positives = 68/136 (49%), Gaps = 8/136 (5%)
Frame = -2
Query: 434 NMSTVSSSNKLPN----IWHHRLGHPSQPCYISLQQM----YPPITTSPCKNCDTCHFAK 485
+ + +++S+ LP +WH RLGH + L + I P C +C AK
Sbjct: 411 SQALLTTSSPLPRASFELWHSRLGHVNFDIIKQLHKHGCLDVSSILPKPIC-CTSCQMAK 235
Query: 486 QKRLSFSLRNTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSE 545
KRL F N VL+L+H D+ GP VASI G YF I V D SRFTW +K KS+
Sbjct: 234 SKRLVFHDNNKRASAVLDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSD 55
Query: 546 ARAALQTFILYSQRQY 561
L F ++ + ++
Sbjct: 54 FSDVLLRFKVFMENRF 7
>BP058111
Length = 570
Score = 58.2 bits (139), Expect = 5e-09
Identities = 31/65 (47%), Positives = 39/65 (59%), Gaps = 4/65 (6%)
Frame = -1
Query: 446 NIWHHRLGHPSQPCYISLQQMYPPIT----TSPCKNCDTCHFAKQKRLSFSLRNTIYGTV 501
N+WH RLGH S LQ+ +P IT TSPC DTCH AKQK+LSF T+ +
Sbjct: 189 NLWHLRLGHTSSKKLAILQKNFPFITCKKITSPC---DTCHMAKQKKLSFPNSVTLSSEI 19
Query: 502 LELLH 506
+L+H
Sbjct: 18 FDLIH 4
>AV427921
Length = 284
Score = 56.2 bits (134), Expect = 2e-08
Identities = 30/80 (37%), Positives = 46/80 (57%)
Frame = +1
Query: 596 TPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFII 655
TPQQNGV ERK++ ILN+ R+L+ S L F AV+ ++ ++N PT V+ P
Sbjct: 22 TPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEEA 201
Query: 656 LHNLKPDIEHLKVFGSLCYA 675
+P ++H ++ L YA
Sbjct: 202 WSGRQPAVDHFRISRCLAYA 261
>AV766088
Length = 501
Score = 53.5 bits (127), Expect = 1e-07
Identities = 32/104 (30%), Positives = 50/104 (47%), Gaps = 4/104 (3%)
Frame = -3
Query: 446 NIWHHRLGHPSQPCYISLQQ----MYPPITTSPCKNCDTCHFAKQKRLSFSLRNTIYGTV 501
++WH RLGHPS L+ Y + SP C++C F K RL F N +
Sbjct: 310 SLWHSRLGHPSSSALRYLRSNKFISYELLNYSPV--CESCVFGKHVRLPFVSSNNVTVMP 137
Query: 502 LELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSE 545
++LH D+W V S G +++ + + D + F W + KS+
Sbjct: 136 FDILHSDLW-TSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQ 8
>BP054557
Length = 538
Score = 52.0 bits (123), Expect = 3e-07
Identities = 56/187 (29%), Positives = 80/187 (41%), Gaps = 13/187 (6%)
Frame = -2
Query: 311 IMDSGATDHIASSLSFFSSYYSIKHVPVSLPNR----AHAMKTI*GQYFFHLP*LYIMFS 366
++D+GAT H+ +L F + + V +S+PN AH ++ F L + + S
Sbjct: 537 LLDTGATHHVCHNLDVFQALRQVHPVLLSMPNGQKEIAHMADSVYFSEKFFLAXVLYVPS 358
Query: 367 MYLNFLSILFLYTSLP-KY*ITD*FSLIPCV*FWTRPL*R*LGELRSRMGYTSWSFHLSL 425
N +S+ L SL + I D C RP + +G R G +
Sbjct: 357 FNFNLISVSALVKSLNCRLVIFD-----ECCLLQERPTLKTIGAAEERGGLYAVIDPAVK 193
Query: 426 VSNHFHSVNMST-VSSSNKLP---NIWHHRLGHPSQPCYISLQQMYPPIT----TSPCKN 477
+ N+ T VS N N+WH RLGH S LQ+ +P IT TSP
Sbjct: 192 AKHPPAEGNIKTQVSHFNFCQFDCNLWHLRLGHTSSKKLAILQKNFPFITCKQITSP--- 22
Query: 478 CDTCHFA 484
CDTCH A
Sbjct: 21 CDTCHMA 1
>BP033602
Length = 533
Score = 47.8 bits (112), Expect = 7e-06
Identities = 24/87 (27%), Positives = 46/87 (52%), Gaps = 3/87 (3%)
Frame = +3
Query: 447 IWHHRLGHPSQPCYISLQQMYPPITTSP-CKN--CDTCHFAKQKRLSFSLRNTIYGTVLE 503
+WH+RLGHP+ + L+ ++P + + C + C++C AK +R + +
Sbjct: 276 LWHNRLGHPN---FQYLRHLFPDLFKNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFY 446
Query: 504 LLHVDIWGPYSVASITGAKYFHIIVYD 530
L+H D+WGP + + G ++F + D
Sbjct: 447 LIHSDVWGPSKITTQFGKRWFVTFIDD 527
>BP081717
Length = 291
Score = 46.2 bits (108), Expect = 2e-05
Identities = 19/51 (37%), Positives = 29/51 (56%)
Frame = -1
Query: 446 NIWHHRLGHPSQPCYISLQQMYPPITTSPCKNCDTCHFAKQKRLSFSLRNT 496
++WH RLGHPS + +++P + CD CH AKQ +L F + +T
Sbjct: 171 DMWHFRLGHPSSKVLQHISKVFPYVKLHSNIICDFCHMAKQTKLPFPISDT 19
>TC13132 similar to UP|Q9FJA1 (Q9FJA1) Similarity to retroelement pol
polyprotein, partial (3%)
Length = 591
Score = 42.7 bits (99), Expect = 2e-04
Identities = 20/59 (33%), Positives = 28/59 (46%), Gaps = 2/59 (3%)
Frame = +1
Query: 200 LCTHCGRQNHTVETCFLKHGYPSNFPNGYKPEKAYI--PTSGGESHSSSDQDEPPSSSL 256
+C+HCG HT CF HGYP YKP + P+SG ++ + P + L
Sbjct: 37 ICSHCGIPGHTAAKCFKLHGYPPGMKPKYKPSTHAVADPSSGTFQNAVTTLSPAPCNQL 213
>BP078157
Length = 395
Score = 40.4 bits (93), Expect = 0.001
Identities = 21/52 (40%), Positives = 27/52 (51%), Gaps = 3/52 (5%)
Frame = -2
Query: 443 KLPNIWHHRLGHPSQPCYISLQQMYPPITTSP---CKNCDTCHFAKQKRLSF 491
KLP +WH RLGH S + + P T P NC+ C +K +RLSF
Sbjct: 160 KLPRLWHSRLGHLSDK-VLKVVSNLVPFTVLPDFHSHNCNVCPLSKMRRLSF 8
>AV406379
Length = 226
Score = 39.3 bits (90), Expect = 0.002
Identities = 20/63 (31%), Positives = 34/63 (53%)
Frame = -3
Query: 429 HFHSVNMSTVSSSNKLPNIWHHRLGHPSQPCYISLQQMYPPITTSPCKNCDTCHFAKQKR 488
++ ++ST+ + + P++ H RLGHPS L+QM P ++ C++C K R
Sbjct: 221 YYMQPHLSTLCAIFESPDLLHCRLGHPSLE---RLKQMVPKLSHLKTLECESCQLGKHVR 51
Query: 489 LSF 491
SF
Sbjct: 50 ASF 42
>AV764934
Length = 406
Score = 39.3 bits (90), Expect = 0.002
Identities = 34/109 (31%), Positives = 48/109 (43%), Gaps = 4/109 (3%)
Frame = -2
Query: 413 RMGYTSWSFHLSLVSNHFHSVNMSTVSSSNKLPNIWHHRLGHPS----QPCYISLQQMYP 468
++GYT S+ S+ + S+VS+ L + WH RLGHP + S Y
Sbjct: 300 QLGYTR-----SMSSSPVSGRHPSSVSAFVTL-STWHSRLGHPHLDNLKRVLSSCNVSYS 139
Query: 469 PITTSPCKNCDTCHFAKQKRLSFSLRNTIYGTVLELLHVDIWGPYSVAS 517
P T + C K RL + T Y EL++ D+WGP V S
Sbjct: 138 PKDTVEFRT--ACCLGKAHRLPSQMSTTTY-LPFELIYSDLWGPAPVCS 1
>BP080584
Length = 507
Score = 38.9 bits (89), Expect = 0.003
Identities = 17/40 (42%), Positives = 24/40 (59%)
Frame = -3
Query: 446 NIWHHRLGHPSQPCYISLQQMYPPITTSPCKNCDTCHFAK 485
N+WH RLGHPS S++ ++P I + + CD C AK
Sbjct: 121 NLWHFRLGHPSFVKEQSIRDIFPYIVCNKEQVCDICPLAK 2
>BP035453
Length = 550
Score = 31.6 bits (70), Expect(2) = 0.009
Identities = 15/37 (40%), Positives = 25/37 (67%)
Frame = -1
Query: 432 SVNMSTVSSSNKLPNIWHHRLGHPSQPCYISLQQMYP 468
SVN+S+ + + N+WH+RLGHPS S++ ++P
Sbjct: 181 SVNVSSHFDPS-VHNLWHYRLGHPSFVKGQSIRDLFP 74
Score = 24.6 bits (52), Expect(2) = 0.009
Identities = 9/14 (64%), Positives = 10/14 (71%)
Frame = -3
Query: 478 CDTCHFAKQKRLSF 491
CD C KQK+LSF
Sbjct: 47 CDVCPLLKQKKLSF 6
>TC11361
Length = 549
Score = 37.4 bits (85), Expect = 0.009
Identities = 23/78 (29%), Positives = 35/78 (44%), Gaps = 3/78 (3%)
Frame = +3
Query: 201 CTHCGRQNHTVETCFLKHGYPSNFPNGYKPEKAYIPTSGGESHSSSDQDEP-PSSSLELT 259
CTHCG HT + C+ G+P ++ T+G E S++ P + ++
Sbjct: 42 CTHCGILGHTKDKCYRLIGFPPHYKKSKSVASVNAVTTGTEDSPSANSFVPFTAQQCQVL 221
Query: 260 REYLQGLL--ALLPQSKP 275
+ LQ L A LP S P
Sbjct: 222 LQMLQTQLSHAQLPDSAP 275
>BP063902
Length = 494
Score = 36.2 bits (82), Expect = 0.020
Identities = 19/50 (38%), Positives = 26/50 (52%), Gaps = 3/50 (6%)
Frame = -3
Query: 446 NIWHHRLGHPSQPCYISLQQMYPPITTSPCKN---CDTCHFAKQKRLSFS 492
++WHHRLGHP+ L+ I P ++ CD+C K RL FS
Sbjct: 186 SVWHHRLGHPASSALNHLRN-NKLIFCEPSRSSSVCDSCVLGKHVRLPFS 40
>BP073434
Length = 490
Score = 35.4 bits (80), Expect = 0.034
Identities = 26/78 (33%), Positives = 35/78 (44%), Gaps = 2/78 (2%)
Frame = -1
Query: 427 SNHFHSVNMSTVSSSNKLPNIWHHRLGHPSQPCYISLQQMYPPITTSPCKN--CDTCHFA 484
S F VN T +++NK IWH RLGHP Q ++ Q +S C C
Sbjct: 232 SPSFDIVNTITTTTTNKFL-IWHSRLGHPHQDALQAVLQSCNIHLSSRDMTDFCYACCLG 56
Query: 485 KQKRLSFSLRNTIYGTVL 502
K L SL ++Y +L
Sbjct: 55 KSHILPSSLSTSVYSFLL 2
>AV769206
Length = 536
Score = 33.9 bits (76), Expect = 0.098
Identities = 19/69 (27%), Positives = 38/69 (54%), Gaps = 5/69 (7%)
Frame = -3
Query: 683 TKFPLRARKCIVLGQKEGGVKGYLMFDLKTREVFLSRNVIFYETIFPFQA-----TDKVS 737
TK L +++C+ +G KGY D R +++S++V+F+E FP+ + T +
Sbjct: 513 TKLSLHSKECVFIGYSINH-KGYKCLDQSGR-IYISKDVLFHEHRFPYPSLFPDETTSST 340
Query: 738 LSSLFPINS 746
+ FP+++
Sbjct: 339 SAEFFPLST 313
>TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B precursor
(Pectin methylesterase) (PE) , partial (54%)
Length = 1145
Score = 32.7 bits (73), Expect = 0.22
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +3
Query: 478 CDTCHFAKQKRLSFSLRNTIYGTVLELLHVDIW 510
CD+C K RL FS TI ++LH D+W
Sbjct: 1029 CDSCVLGKHVRLPFSSSETITLRPFDILHSDLW 1127
>BP070393
Length = 359
Score = 30.8 bits (68), Expect = 0.83
Identities = 15/25 (60%), Positives = 18/25 (72%), Gaps = 1/25 (4%)
Frame = -3
Query: 432 SVNMSTVSS-SNKLPNIWHHRLGHP 455
S NMS+VSS N ++WH RLGHP
Sbjct: 129 SQNMSSVSSLGNSSYDLWHCRLGHP 55
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.337 0.145 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,182,504
Number of Sequences: 28460
Number of extensions: 245709
Number of successful extensions: 2415
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 2377
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2408
length of query: 754
length of database: 4,897,600
effective HSP length: 97
effective length of query: 657
effective length of database: 2,136,980
effective search space: 1403995860
effective search space used: 1403995860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0296a.5


