

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0284.4
(247 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
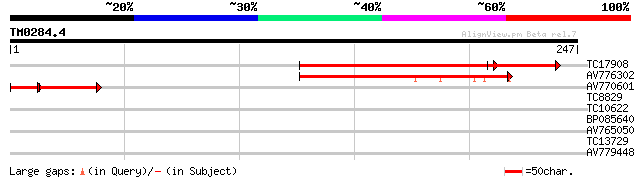
Sequences producing significant alignments: (bits) Value
TC17908 similar to UP|PMM_ARATH (O80840) Probable phosphomannomu... 165 5e-56
AV776302 92 6e-20
AV770601 54 1e-09
TC8829 30 0.50
TC10622 similar to UP|O24316 (O24316) Acetyl-CoA carboxylase (F... 27 3.3
BP085640 26 7.3
AV765050 26 7.3
TC13729 similar to UP|Q9FFC3 (Q9FFC3) Protease-like protein, par... 26 7.3
AV779448 25 9.5
>TC17908 similar to UP|PMM_ARATH (O80840) Probable phosphomannomutase (PMM)
, partial (34%)
Length = 574
Score = 165 bits (417), Expect(2) = 5e-56
Identities = 77/87 (88%), Positives = 82/87 (93%)
Frame = +1
Query: 127 MLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVFPQ 186
MLNVSPIGR+CS EERDEFEKYDKVQ +RP+MVSVLREKFAHLNLTFSIGGQISFDVFPQ
Sbjct: 1 MLNVSPIGRDCSLEERDEFEKYDKVQYLRPRMVSVLREKFAHLNLTFSIGGQISFDVFPQ 180
Query: 187 GWDKTYCLRYLDGFNEIHFFGDKTYKG 213
WD TYCLRYLDGFNEIHFFGD+T +G
Sbjct: 181 AWD*TYCLRYLDGFNEIHFFGDETLQG 261
Score = 68.9 bits (167), Expect(2) = 5e-56
Identities = 30/32 (93%), Positives = 31/32 (96%)
Frame = +2
Query: 209 KTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
K YKGGNDH+IYESERTIGHTVTSPEDTIKQC
Sbjct: 248 KLYKGGNDHDIYESERTIGHTVTSPEDTIKQC 343
>AV776302
Length = 514
Score = 92.4 bits (228), Expect = 6e-20
Identities = 60/100 (60%), Positives = 67/100 (67%), Gaps = 7/100 (7%)
Frame = -1
Query: 127 MLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSI-GGQISFDVFP 185
MLNVSP R+CSQ ERDEFEKYDKVQN+RP MVSVLREKFAHLNLTFS + FDVF
Sbjct: 514 MLNVSPTARDCSQTERDEFEKYDKVQNLRPHMVSVLREKFAHLNLTFSXWRTR*RFDVFL 335
Query: 186 Q--GWDKTYCLRYLDGFN-EIHF-FGDKTYKGGND--HEI 219
G + RYL + EIH FGD+ G + HEI
Sbjct: 334 SRLGTKHSLPCRYL*WVSIEIHIPFGDRNLTQGWELTHEI 215
>AV770601
Length = 163
Score = 54.3 bits (129), Expect(2) = 1e-09
Identities = 26/27 (96%), Positives = 27/27 (99%)
Frame = +2
Query: 14 VDGTLTAPRKVANSEMLGFMQELRKVV 40
+DGTLTAPRKVANSEMLGFMQELRKVV
Sbjct: 83 LDGTLTAPRKVANSEMLGFMQELRKVV 163
Score = 23.9 bits (50), Expect(2) = 1e-09
Identities = 11/14 (78%), Positives = 12/14 (85%)
Frame = +1
Query: 1 MAVAKPGVIALFDV 14
MAVAK GV+A FDV
Sbjct: 43 MAVAKAGVMAWFDV 84
>TC8829
Length = 511
Score = 29.6 bits (65), Expect = 0.50
Identities = 16/52 (30%), Positives = 28/52 (53%)
Frame = -2
Query: 91 FLGDEKLKEFINFTLHYIADLDIPIKRGTFIEFRSGMLNVSPIGRNCSQEER 142
+L + LK+ NF+ I+ ++ + R F RSG++N+ P NC +R
Sbjct: 372 YLAKQSLKDEENFSSLTISQINY-LSRKPFSVMRSGVINIWPCRNNCKWIDR 220
>TC10622 similar to UP|O24316 (O24316) Acetyl-CoA carboxylase (Fragment) ,
partial (42%)
Length = 522
Score = 26.9 bits (58), Expect = 3.3
Identities = 17/59 (28%), Positives = 30/59 (50%), Gaps = 3/59 (5%)
Frame = -3
Query: 69 FSENGLVAHKQGKLIGTESLKTFLGDEKLKEFI---NFTLHYIADLDIPIKRGTFIEFR 124
FS+ L +H QG ++ L ++LG + K F+ N L + + + G F++FR
Sbjct: 325 FSDYTLCSHSQGSIMKFSKLGSYLGIHRQKLFLTGFNLLLQGLNGTRVSVSLG-FLKFR 152
>BP085640
Length = 331
Score = 25.8 bits (55), Expect = 7.3
Identities = 8/17 (47%), Positives = 12/17 (70%)
Frame = +3
Query: 188 WDKTYCLRYLDGFNEIH 204
WD+TY + +L GF +H
Sbjct: 279 WDQTYHISHLAGFQHVH 329
>AV765050
Length = 234
Score = 25.8 bits (55), Expect = 7.3
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = -3
Query: 173 FSIGGQISFDVFPQGW 188
F GG I F VFP+GW
Sbjct: 160 FISGGIIPFSVFPKGW 113
>TC13729 similar to UP|Q9FFC3 (Q9FFC3) Protease-like protein, partial (15%)
Length = 508
Score = 25.8 bits (55), Expect = 7.3
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = +1
Query: 46 GGSDLIKISEQLGSTVTHDYDYVFSENGLVA 76
G S+ + E GST HD ++F + GL A
Sbjct: 214 GRSEFVNAGEFSGSTSLHDGPHMFIKTGLFA 306
>AV779448
Length = 509
Score = 25.4 bits (54), Expect = 9.5
Identities = 12/30 (40%), Positives = 17/30 (56%), Gaps = 2/30 (6%)
Frame = +3
Query: 209 KTYKGGNDHEIYESERTIGHTVTS--PEDT 236
+TYKGGN +R +GH + PE+T
Sbjct: 123 ETYKGGNGRATRRVQRLVGHVPSI**PENT 212
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.140 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,529,806
Number of Sequences: 28460
Number of extensions: 42103
Number of successful extensions: 163
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 162
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 163
length of query: 247
length of database: 4,897,600
effective HSP length: 88
effective length of query: 159
effective length of database: 2,393,120
effective search space: 380506080
effective search space used: 380506080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0284.4


