

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
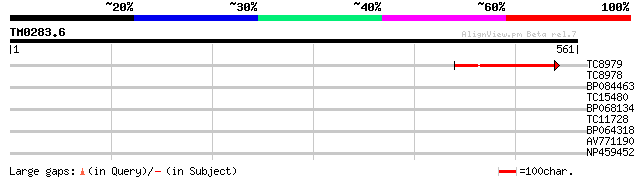
Query= TM0283.6
(561 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC8979 weakly similar to UP|Q8RY59 (Q8RY59) At1g32230/F3C3_1, pa... 103 6e-23
TC8978 30 0.79
BP084463 29 1.8
TC15480 similar to UP|V712_ARATH (Q9SIQ9) Vesicle-associated mem... 28 5.1
BP068134 27 6.7
TC11728 27 6.7
BP064318 27 8.8
AV771190 27 8.8
NP459452 NDX2 homeobox protein [Lotus japonicus] 27 8.8
>TC8979 weakly similar to UP|Q8RY59 (Q8RY59) At1g32230/F3C3_1, partial
(12%)
Length = 833
Score = 103 bits (258), Expect = 6e-23
Identities = 55/104 (52%), Positives = 73/104 (69%)
Frame = +1
Query: 441 RDNSSGANSDSHCSGDPVQSASSAYNEIAGNGVACTQKIPKSPWMPLHMLIAAINDKVPP 500
R N S NS+S S +QS S + VA + K P+SPWMP ML AAI +KVP
Sbjct: 1 RKNVSAVNSNSQGSKSLLQSQSPT-EDTGITTVASSHKAPRSPWMPFPMLFAAITEKVPS 177
Query: 501 NDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTITTLQ 544
NDM+LI +++ +++K+I+RDDFVK++RLIVGDTLLR TIT+LQ
Sbjct: 178 NDMNLINMHYQQFRSKKITRDDFVKKLRLIVGDTLLRATITSLQ 309
>TC8978
Length = 506
Score = 30.4 bits (67), Expect = 0.79
Identities = 14/19 (73%), Positives = 17/19 (88%)
Frame = +1
Query: 524 VKEMRLIVGDTLLRDTITT 542
VK++RLI GDTLLR TIT+
Sbjct: 1 VKKLRLIFGDTLLRATITS 57
>BP084463
Length = 380
Score = 29.3 bits (64), Expect = 1.8
Identities = 14/31 (45%), Positives = 17/31 (54%), Gaps = 1/31 (3%)
Frame = +2
Query: 319 CELFTTKCTYGVGVHLAAISCPYASARY-CD 348
C L T+ C Y +G + I C YA RY CD
Sbjct: 77 CILITSTCEYKLGAKII*ILCTYAYKRYTCD 169
>TC15480 similar to UP|V712_ARATH (Q9SIQ9) Vesicle-associated membrane
protein 712 (AtVAMP712), partial (53%)
Length = 482
Score = 27.7 bits (60), Expect = 5.1
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 539 TITTLQFMVHNIFFFFF 555
T TT ++H+IFFFFF
Sbjct: 6 TTTTFNSLIHHIFFFFF 56
>BP068134
Length = 462
Score = 27.3 bits (59), Expect = 6.7
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = -3
Query: 352 NGVRHLILCRIIMGNMELLCPSIGTSNVQ 380
NG L+L + GN+ LLCP+ TS ++
Sbjct: 295 NGRPGLVLAMYLHGNLLLLCPTAQTSRLE 209
>TC11728
Length = 832
Score = 27.3 bits (59), Expect = 6.7
Identities = 15/56 (26%), Positives = 24/56 (42%)
Frame = -3
Query: 362 IIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICPAFV 417
I + + CPS+ VQ QP + + IW++N+ IC AF+
Sbjct: 788 ITYNHCNVWCPSMLAKVVQTQPKPQTIHKSTSFSSKKKLFIIWDLNV---ICNAFL 630
>BP064318
Length = 331
Score = 26.9 bits (58), Expect = 8.8
Identities = 10/25 (40%), Positives = 17/25 (68%)
Frame = -3
Query: 391 GVDDIQCPRYYTIWNMNISTYICPA 415
GVD ++ P+ +T WN+ I + +C A
Sbjct: 119 GVDIMRKPKRHTRWNLRIVSLLCAA 45
>AV771190
Length = 497
Score = 26.9 bits (58), Expect = 8.8
Identities = 9/13 (69%), Positives = 12/13 (92%)
Frame = -2
Query: 429 YFCGTVDKNNVSR 441
YFCG+VD+NNV +
Sbjct: 253 YFCGSVDENNVKK 215
>NP459452 NDX2 homeobox protein [Lotus japonicus]
Length = 2223
Score = 26.9 bits (58), Expect = 8.8
Identities = 18/42 (42%), Positives = 22/42 (51%), Gaps = 1/42 (2%)
Frame = +1
Query: 115 NGRDLMLDFLHACQMDLKTGFIQPIAWIDKVGGYF-FPEVDV 155
N DLM L AC + L TGF+ P W + V P+VDV
Sbjct: 100 NHMDLMSSTLVACNLYLLTGFLSP-QWQEMVPVLLEHPKVDV 222
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,858,044
Number of Sequences: 28460
Number of extensions: 135583
Number of successful extensions: 694
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 690
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 694
length of query: 561
length of database: 4,897,600
effective HSP length: 95
effective length of query: 466
effective length of database: 2,193,900
effective search space: 1022357400
effective search space used: 1022357400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0283.6


