

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.14.1
(63 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
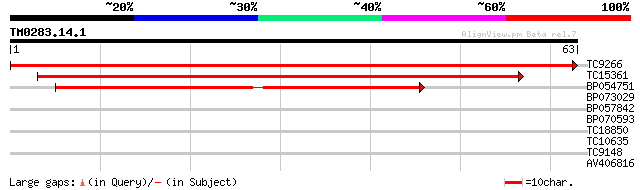
TC9266 similar to UP|Q9ASW5 (Q9ASW5) Epoxide hydrolase (Express... 114 4e-27
TC15361 weakly similar to UP|Q84VQ5 (Q84VQ5) Hydrolase, alpha/be... 48 3e-07
BP054751 38 3e-04
BP073029 33 0.011
BP057842 25 3.9
BP070593 25 3.9
TC18850 similar to UP|Q41414 (Q41414) Epoxide hydrolase , parti... 23 8.6
TC10635 similar to UP|Q946U8 (Q946U8) Long cell-linked locus pro... 23 8.6
TC9148 similar to UP|AAQ65137 (AAQ65137) At4g27270, partial (97%) 23 8.6
AV406816 23 8.6
>TC9266 similar to UP|Q9ASW5 (Q9ASW5) Epoxide hydrolase (Expressed
protein) (At2g18360/T30D6.13) , partial (34%)
Length = 506
Score = 114 bits (285), Expect = 4e-27
Identities = 54/63 (85%), Positives = 58/63 (91%)
Frame = +3
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
RIHLLWGE DQIFK ELA+NMKE+LGD A+FQGIKKAGHLVHLERPCVYNR LK+FIASF
Sbjct: 153 RIHLLWGETDQIFKVELAQNMKEELGDAASFQGIKKAGHLVHLERPCVYNRYLKQFIASF 332
Query: 61 FAS 63
AS
Sbjct: 333 LAS 341
>TC15361 weakly similar to UP|Q84VQ5 (Q84VQ5) Hydrolase, alpha/beta fold
family, partial (11%)
Length = 695
Score = 48.1 bits (113), Expect = 3e-07
Identities = 20/54 (37%), Positives = 33/54 (61%)
Frame = +1
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
++WGEND+IF ++A +KE + A + IK A H+ LE+P +N + F+
Sbjct: 301 IVWGENDRIFPVQMAHELKEAISQKARLELIKDASHVPQLEKPVEFNNIILNFL 462
>BP054751
Length = 553
Score = 38.1 bits (87), Expect = 3e-04
Identities = 19/41 (46%), Positives = 25/41 (60%)
Frame = -1
Query: 6 WGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
WGEND+I +LA + +L D A + I GHL HLE+P
Sbjct: 553 WGENDRIISNKLAVQLHCELPD-AILRQIPDCGHLPHLEKP 434
>BP073029
Length = 452
Score = 33.1 bits (74), Expect = 0.011
Identities = 23/54 (42%), Positives = 28/54 (51%), Gaps = 3/54 (5%)
Frame = -3
Query: 1 RIHLLWGENDQIFK-RELAKNMKEQLGDGATF-QGIKKAGHLVHLER-PCVYNR 51
RIH+LWGE D F+ K + GDGA+ +KK R PCVYNR
Sbjct: 375 RIHILWGETDHNFQGGACPKYERGN*GDGASLSMALKKDWSA*FTWREPCVYNR 214
>BP057842
Length = 516
Score = 24.6 bits (52), Expect = 3.9
Identities = 17/66 (25%), Positives = 28/66 (41%), Gaps = 11/66 (16%)
Frame = -3
Query: 4 LLWGENDQIFKR---ELAKNMKEQLGDGATFQG--------IKKAGHLVHLERPCVYNRC 52
++ GE D FK+ E+ + LG +G + GH HLE P
Sbjct: 295 IIHGEKDTKFKKIAQEMMHTICSGLGRNEHAEGNNIHQVVEVPNCGHAPHLENPLPVITA 116
Query: 53 LKRFIA 58
L++F++
Sbjct: 115 LRKFMS 98
>BP070593
Length = 376
Score = 24.6 bits (52), Expect = 3.9
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = -3
Query: 35 KKAGHLVHLERPCVYNRCLKRFIAS 59
K AGH +H E+P LK F+ S
Sbjct: 374 KNAGHAIHKEKPKDLCMALKSFLFS 300
>TC18850 similar to UP|Q41414 (Q41414) Epoxide hydrolase , partial (22%)
Length = 474
Score = 23.5 bits (49), Expect = 8.6
Identities = 12/36 (33%), Positives = 17/36 (46%)
Frame = +1
Query: 25 LGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
L D +G+ GH +H ERP N + F+ F
Sbjct: 115 LDDVVVLEGV---GHFLHQERPHEINNHIYDFLRKF 213
>TC10635 similar to UP|Q946U8 (Q946U8) Long cell-linked locus protein,
partial (16%)
Length = 1243
Score = 23.5 bits (49), Expect = 8.6
Identities = 8/22 (36%), Positives = 13/22 (58%)
Frame = +3
Query: 1 RIHLLWGENDQIFKRELAKNMK 22
R+ LLW +F+R LA ++
Sbjct: 465 RLSLLWARKGGVFRRRLATELR 530
>TC9148 similar to UP|AAQ65137 (AAQ65137) At4g27270, partial (97%)
Length = 980
Score = 23.5 bits (49), Expect = 8.6
Identities = 12/34 (35%), Positives = 17/34 (49%)
Frame = +1
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGI 34
++ LLW E Q + R L +K LG QG+
Sbjct: 139 KVQLLWKE*KQSYGRYLKHCLKRFLGRWEHLQGV 240
>AV406816
Length = 417
Score = 23.5 bits (49), Expect = 8.6
Identities = 7/29 (24%), Positives = 18/29 (61%)
Frame = -1
Query: 13 FKRELAKNMKEQLGDGATFQGIKKAGHLV 41
F+R + +++ ++G ++ G+ + HLV
Sbjct: 243 FRRRVGESVASEIGSDSSEAGVSEGNHLV 157
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.328 0.142 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,194,383
Number of Sequences: 28460
Number of extensions: 10350
Number of successful extensions: 71
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 70
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 70
length of query: 63
length of database: 4,897,600
effective HSP length: 39
effective length of query: 24
effective length of database: 3,787,660
effective search space: 90903840
effective search space used: 90903840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 49 (23.5 bits)
Lotus: description of TM0283.14.1


