

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
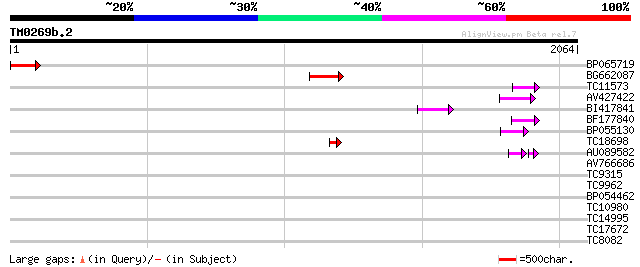
Query= TM0269b.2
(2064 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP065719 183 2e-46
BG662087 115 5e-26
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 73 5e-13
AV427422 65 1e-10
BI417841 64 2e-10
BF177840 59 8e-09
BP055130 53 6e-07
TC18698 50 5e-06
AU089582 38 2e-05
AV766686 33 0.35
TC9315 32 1.0
TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial... 32 1.3
BP054462 31 2.3
TC10980 similar to UP|Q8VZA1 (Q8VZA1) Laccase (Diphenol oxidase)... 30 5.0
TC14995 similar to UP|Q9SVJ8 (Q9SVJ8) Kinesin like protein, part... 30 5.0
TC17672 similar to UP|Q93X83 (Q93X83) Hsp70 interacting protein/... 29 6.6
TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-... 29 8.6
>BP065719
Length = 567
Score = 183 bits (465), Expect = 2e-46
Identities = 104/112 (92%), Positives = 104/112 (92%)
Frame = +1
Query: 1 T*PGTKSKQET*PSMKI*K*STSRVL*PKMHLPGSPPSLLGQSIHGPSWKEFSMNSSLGE 60
T*PGTKSKQET*PSMKI*K*STSRVL*PKM LPGSPPSLLGQSIHGPSWKEFSMNSSLGE
Sbjct: 232 T*PGTKSKQET*PSMKI*K*STSRVL*PKMPLPGSPPSLLGQSIHGPSWKEFSMNSSLGE 411
Query: 61 SAK*V*RT*LVLKESQQSLLMTT*TGLGCLNLDVSPMSLNTS*LS*PLVV*S 112
SAK*V RT LV KESQQSLLM T*TGLGCLNLD SPM LNTS*LS* LVV*S
Sbjct: 412 SAK*VXRTWLVSKESQQSLLMIT*TGLGCLNLDASPMCLNTS*LS*XLVV*S 567
>BG662087
Length = 373
Score = 115 bits (289), Expect = 5e-26
Identities = 58/122 (47%), Positives = 82/122 (66%)
Frame = +1
Query: 1093 NGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVS 1152
+GK R+ +D+ DLN A PKD Y +P + +VD A+ +E LSL+D YSGY+QI + D
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 1153 KTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRD 1212
KTAF A Y + +PFGLKNAGATYQ +M+ +F D + M+VY+D+++VKS R
Sbjct: 193 KTAFMT--ARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRA 366
Query: 1213 DH 1214
+H
Sbjct: 367 NH 372
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 72.8 bits (177), Expect = 5e-13
Identities = 35/96 (36%), Positives = 52/96 (53%)
Frame = +2
Query: 1832 DQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQ 1891
+ G F ++ F + GI++ S+ + Q NGQ EAANK ++ IK+ + W
Sbjct: 8 ENGIQFTSKQTQDFCDGMGIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWID 187
Query: 1892 TLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEV 1927
L VLW+Y P+ S TPF L YG + +LPVE+
Sbjct: 188 ELPIVLWSYNTMPQSSIKETPF*LTYGADTMLPVEI 295
>AV427422
Length = 417
Score = 65.1 bits (157), Expect = 1e-10
Identities = 36/133 (27%), Positives = 69/133 (51%), Gaps = 2/133 (1%)
Frame = +1
Query: 1782 SRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIE--FISSIVYRFGLPESLTTDQGTVFVG 1839
S+ ++ ++V +D +K+ +PL++ VI F+ +V G+P S+ +D+ +F+
Sbjct: 19 SKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDPLFMS 198
Query: 1840 QKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWA 1899
+ G KL ST Y+ +++GQ E N+ L + ++ + +PK+W + +
Sbjct: 199 NFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPWAEYW 378
Query: 1900 YRNSPKESTGATP 1912
Y S STG TP
Sbjct: 379 YNTSYHVSTGQTP 417
>BI417841
Length = 617
Score = 63.9 bits (154), Expect = 2e-10
Identities = 45/136 (33%), Positives = 69/136 (50%), Gaps = 6/136 (4%)
Frame = +1
Query: 1484 LFFDGSSHKN--GTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGA 1541
L FDGSS N G G + + G + + N +NN+AEY L GL+ G
Sbjct: 142 LEFDGSSKGNPGSAGAGAVLRAEDGSKVYLREGV-GNQTNNQAEYRGLILGLKHAHEQGY 318
Query: 1542 KNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSRVDNQEANEL 1601
+++ VKGDS+LV KQ+ +K + N+A +A L +KF + HV R N EA+
Sbjct: 319 QHINVKGDSQLVCKQVEGSWKARNPNIASLCNEAKELKSKFQSFDINHVPRQYNSEADVQ 498
Query: 1602 AQIA----SGYMVDKC 1613
A + +G++ + C
Sbjct: 499 ANLGVNLPAGHVEEYC 546
>BF177840
Length = 410
Score = 58.9 bits (141), Expect = 8e-09
Identities = 31/100 (31%), Positives = 51/100 (51%)
Frame = +2
Query: 1828 SLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPK 1887
S+ +D+ T F+ + G KLL ST + Q +GQ E NKTL +L++ + R K
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 1888 NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEV 1927
W L + +AY +T +PF + YG + P+++
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYGYNPLTPLDL 313
>BP055130
Length = 567
Score = 52.8 bits (125), Expect = 6e-07
Identities = 34/104 (32%), Positives = 52/104 (49%), Gaps = 2/104 (1%)
Frame = +2
Query: 1788 IIVAIDYFTKWVEAIPLQ-NVTQDTVIE-FISSIVYRFGLPESLTTDQGTVFVGQKVASF 1845
IIV +D +K+ P + N T V E F+S+IV G+P ++ +D+ F F
Sbjct: 191 IIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAFWKHF 370
Query: 1846 AESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNW 1889
+ G L S+ Y+ Q +GQ EA NK L ++ V P+ W
Sbjct: 371 FKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLW 502
>TC18698
Length = 808
Score = 49.7 bits (117), Expect = 5e-06
Identities = 23/44 (52%), Positives = 32/44 (72%)
Frame = -2
Query: 1165 YEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKS 1208
Y + VMP GLKN TYQR+M+ IFH I ++VY++D++VKS
Sbjct: 771 YYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKS 640
>AU089582
Length = 383
Score = 37.7 bits (86), Expect(2) = 2e-05
Identities = 19/65 (29%), Positives = 34/65 (52%)
Frame = +2
Query: 1815 FISSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTL 1874
++ IV G+P S+ +D+G F SF + G +L ST ++ Q +GQ E + L
Sbjct: 32 YLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQSERTIQIL 211
Query: 1875 ISLIK 1879
+++
Sbjct: 212 EDMLR 226
Score = 28.9 bits (63), Expect(2) = 2e-05
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = +3
Query: 1888 NWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPV 1925
+W Q L + +AY NS + S PF YG+ P+
Sbjct: 252 SWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRRCRSPI 365
>AV766686
Length = 536
Score = 33.5 bits (75), Expect = 0.35
Identities = 27/87 (31%), Positives = 39/87 (44%), Gaps = 1/87 (1%)
Frame = -2
Query: 1872 KTLISL-IKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQ 1930
KTL S+ H R+P+ T G WA R E+ A F A G LP+ + Q
Sbjct: 283 KTLFSISYLGHFIRRPRRPPSTFG*TFWANRRFFGETDNAVEFLAAKG*RNNLPLCLRRQ 104
Query: 1931 SCRIQRQEEIPSEDYWNMMLDELVNLD 1957
S R R+ ++P + D+ + LD
Sbjct: 103 SLRQPREPQLP----LRLSTDKFLKLD 35
>TC9315
Length = 632
Score = 32.0 bits (71), Expect = 1.0
Identities = 14/28 (50%), Positives = 17/28 (60%)
Frame = +2
Query: 560 PSNREQPYHQIQLRKKGRMLSKMRMKTC 587
PSNRE +H Q+R R +S RMK C
Sbjct: 179 PSNRENKWHCRQIRDAARFISFCRMKNC 262
>TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial (21%)
Length = 585
Score = 31.6 bits (70), Expect = 1.3
Identities = 18/55 (32%), Positives = 25/55 (44%)
Frame = -1
Query: 359 TWKRFTPRLEKPLSSFYPAAKMARKKLCCALDAALFSTNPQPRPIKIQK*KGIIR 413
TW PR + L S + A+ A + C A +AA P P P +Q+ G R
Sbjct: 579 TWVARFPRTDPGLPSAHTRAEGAAARTCSASEAAAERATPYPSPGSVQQVMGRFR 415
>BP054462
Length = 422
Score = 30.8 bits (68), Expect = 2.3
Identities = 32/129 (24%), Positives = 56/129 (42%), Gaps = 3/129 (2%)
Frame = +1
Query: 1907 STGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLVLDT 1966
S +PF+ YG+E V LQ I +I + + + DEL++ LL
Sbjct: 19 SAKMSPFQALYGREP----PVLLQGTTIP--SKIAAVNDLQVGRDELLSDLRANLL---- 168
Query: 1967 LTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKD---KRYRKWVPNWEGPFTVEKV 2023
+ +D + NKK R + +GD V+ + P ++ K K P + GP+ +
Sbjct: 169 --KSQDMMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAK 342
Query: 2024 LVNNAYSIK 2032
+ AY ++
Sbjct: 343 IGAVAYRLE 369
>TC10980 similar to UP|Q8VZA1 (Q8VZA1) Laccase (Diphenol oxidase)-like
protein, partial (22%)
Length = 572
Score = 29.6 bits (65), Expect = 5.0
Identities = 16/82 (19%), Positives = 41/82 (49%)
Frame = -2
Query: 856 VMKRKMPLLGGKPMIKMINSSMARISAYAAENKIRTALEAERMAKEMAIDEASVSILPVE 915
V+K+ + LL + ++ + I + ++K+ T M ++ +++ + ++PV
Sbjct: 484 VIKKSLQLLQEELTHNKSSNCIVNIDC-SIQSKLLTTSSIASMLEDPSVEAEQIDLVPVH 308
Query: 916 LTPQEEATTNKDDTCIEENEQS 937
L PQ++ +T +NE++
Sbjct: 307 LPPQKQFSTPNQCAAPNDNERT 242
>TC14995 similar to UP|Q9SVJ8 (Q9SVJ8) Kinesin like protein, partial (4%)
Length = 783
Score = 29.6 bits (65), Expect = 5.0
Identities = 13/41 (31%), Positives = 24/41 (57%)
Frame = +1
Query: 1786 KYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVYRFGLP 1826
K +I+A+ + +K + IP+QN+ + ++ Y FGLP
Sbjct: 430 KPLIIALSFSSKVIMRIPIQNILKAIYRARLTLFFYGFGLP 552
>TC17672 similar to UP|Q93X83 (Q93X83) Hsp70 interacting protein/thioredoxin
chimera, partial (33%)
Length = 709
Score = 29.3 bits (64), Expect = 6.6
Identities = 20/50 (40%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = -2
Query: 221 L*KLILLMWPNAMQSMIYLCLMVCLWSLKTKN-FLLLNKGKGRNIANITT 269
L* +I L + + ++ + C VC S +T N F + +K KGRN ANI T
Sbjct: 438 L*GIIDLKRASMLYNLPFKCAFVC--SHRTVNLFAIFHKEKGRNTANIPT 295
>TC8082 similar to UP|Q8H9E8 (Q8H9E8) Resistant specific protein-3, partial
(54%)
Length = 1913
Score = 28.9 bits (63), Expect = 8.6
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 9/46 (19%)
Frame = +1
Query: 1940 IPSEDYWNMMLD---------ELVNLDEERLLVLDTLTRQKDRIAK 1976
+P E YW MML EL+ D +DT T+ KD+I+K
Sbjct: 97 LPPEIYWKMMLPSTPMPKAVIELLRGDVSIETCVDTYTQSKDKISK 234
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.144 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,325,844
Number of Sequences: 28460
Number of extensions: 475136
Number of successful extensions: 3516
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 3446
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3514
length of query: 2064
length of database: 4,897,600
effective HSP length: 104
effective length of query: 1960
effective length of database: 1,937,760
effective search space: 3798009600
effective search space used: 3798009600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0269b.2


