

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269a.3
(490 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
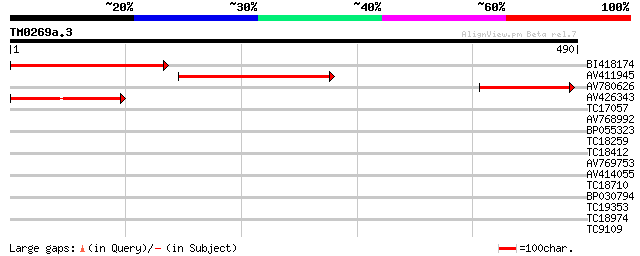
Score E
Sequences producing significant alignments: (bits) Value
BI418174 261 1e-70
AV411945 166 6e-42
AV780626 162 9e-41
AV426343 101 2e-22
TC17057 similar to UP|VATD_ARATH (Q9XGM1) Vacuolar ATP synthase ... 33 0.14
AV768992 28 4.4
BP055323 28 4.4
TC18259 homologue to UP|PSA4_SPIOL (P52427) Proteasome subunit a... 27 5.8
TC18412 27 7.5
AV769753 27 7.5
AV414055 27 7.5
TC18710 27 9.9
BP030794 27 9.9
TC19353 homologue to UP|AX2D_PHAAU (O24542) Auxin-induced protei... 27 9.9
TC18974 weakly similar to PIR|S16321|S16321 light-induced protei... 27 9.9
TC9109 homologue to UP|O23140 (O23140) AP47/50P, partial (18%) 27 9.9
>BI418174
Length = 602
Score = 261 bits (668), Expect = 1e-70
Identities = 136/137 (99%), Positives = 136/137 (99%)
Frame = +2
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MTKISPEIEIKMPM AVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE
Sbjct: 191 MTKISPEIEIKMPM*AVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 370
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 120
QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL
Sbjct: 371 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 550
Query: 121 RVRFAGRNKEDVEKAIS 137
RVRFAGRNKEDVEKAIS
Sbjct: 551 RVRFAGRNKEDVEKAIS 601
>AV411945
Length = 405
Score = 166 bits (421), Expect = 6e-42
Identities = 88/134 (65%), Positives = 110/134 (81%)
Frame = +3
Query: 147 TQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQ 206
TQ EGELIQEK EVKKL NFLKQASEDAKKLV++E++FA AEI++AR+ V R+ EAL+E
Sbjct: 3 TQREGELIQEKSEVKKLANFLKQASEDAKKLVDEERAFARAEIDNARAAVQRVEEALKEH 182
Query: 207 EQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQK 266
E+ SQAS QD++ L +EVQEARRIK+LHQPSKVM ME+EL+ALR Q+ EK+ ++LQK
Sbjct: 183 ERMSQASGKQDLEQLKKEVQEARRIKMLHQPSKVMDMEHELQALRAQLAEKTRHYLRLQK 362
Query: 267 ELTMSKRDEENKSH 280
EL +K+ EEN H
Sbjct: 363 ELARTKKGEENVPH 404
>AV780626
Length = 557
Score = 162 bits (411), Expect = 9e-41
Identities = 79/82 (96%), Positives = 81/82 (98%)
Frame = -2
Query: 407 GRMRIKLCRGWITKAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERN 466
GRMRIKLCRGWITKAREIYSPSMQLCGVRGDAS+AAKALFWQARKGLSFVLTF SEKERN
Sbjct: 556 GRMRIKLCRGWITKAREIYSPSMQLCGVRGDASHAAKALFWQARKGLSFVLTFASEKERN 377
Query: 467 VAIMIARKYALDCNVVLAGPDD 488
VAIMIAR+YALDCNVVLAGPDD
Sbjct: 376 VAIMIARQYALDCNVVLAGPDD 311
>AV426343
Length = 418
Score = 101 bits (252), Expect = 2e-22
Identities = 53/100 (53%), Positives = 74/100 (74%)
Frame = +3
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MT++S + M EAVP VS+DV+F + RFP YK+GA++QI+E T +D + S+K+V+
Sbjct: 120 MTRVSRDFGDTMQNEAVPAVSSDVTFATSRFPSYKIGANNQIME-TKDDPKVVSMKEVVA 296
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIRE 100
+E L +Q R+SVRDLA KF+K LAAAAKLS EA++RE
Sbjct: 297 RETAQLLEQQNRLSVRDLANKFEKGLAAAAKLSEEARLRE 416
>TC17057 similar to UP|VATD_ARATH (Q9XGM1) Vacuolar ATP synthase subunit D
(V-ATPase D subunit) (Vacuolar proton pump D subunit) ,
partial (59%)
Length = 561
Score = 32.7 bits (73), Expect = 0.14
Identities = 38/167 (22%), Positives = 71/167 (41%)
Frame = +3
Query: 85 NLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAV 144
N+ + K R V + GH LLKK DA L V+F ++ ++K +S E++
Sbjct: 99 NVVPTVTMLGVVKARLVGATRGHALLKKKSDA---LTVQF----RQILKKIVSTKESMGD 257
Query: 145 KLTQNEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALE 204
+ + L + K+ A ++ K +V + A + S V +
Sbjct: 258 IMKNSSFALTEAKY----------VAGDNIKHVVLENVKEASLRVRSRTENVAGVKLPKF 407
Query: 205 EQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALR 251
+ D ASK D+ GL Q+ ++ ++ + K + + EL +L+
Sbjct: 408 DYTADGDASK-NDLTGLARGGQQVQQCRVAY--IKAIEVLVELASLQ 539
>AV768992
Length = 473
Score = 27.7 bits (60), Expect = 4.4
Identities = 15/56 (26%), Positives = 31/56 (54%)
Frame = +1
Query: 185 ACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKV 240
A +++ AR+V+ IGE +E D+ +K D+D + ++ I L +P+++
Sbjct: 199 AISDVSQARTVLNLIGERPTHEEVDNARAKLADIDAQLS--RQLEEIVLQSRPTEI 360
>BP055323
Length = 581
Score = 27.7 bits (60), Expect = 4.4
Identities = 18/65 (27%), Positives = 32/65 (48%)
Frame = -2
Query: 70 HKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNK 129
HKR ++R + K ++++ E+ HV+LKKLR+ L+ RV++
Sbjct: 556 HKRNTIRTTSLK--------GRVASSEVFSELIHPLTHVILKKLRELLKIQRVKYPTAQP 401
Query: 130 EDVEK 134
+EK
Sbjct: 400 LRLEK 386
>TC18259 homologue to UP|PSA4_SPIOL (P52427) Proteasome subunit alpha type 4
(20S proteasome alpha subunit C) (20S proteasome
subunit alpha-3) (Proteasome 27 kDa subunit) , partial
(34%)
Length = 366
Score = 27.3 bits (59), Expect = 5.8
Identities = 15/35 (42%), Positives = 22/35 (62%)
Frame = -3
Query: 186 CAEIESARSVVLRIGEALEEQEQDSQASKPQDVDG 220
CA I S+R +VLRIGE EE+ D + + + +G
Sbjct: 139 CAAIISSRHIVLRIGEH-EERNGDWRTERAAEREG 38
>TC18412
Length = 394
Score = 26.9 bits (58), Expect = 7.5
Identities = 24/95 (25%), Positives = 46/95 (48%), Gaps = 2/95 (2%)
Frame = +1
Query: 9 EIKMPMEAVPPVSADVSFISDRFPKYKLG--ADSQILEETVEDNQGPSLKDVIEQEATNL 66
+++ P ++V V + F+ D + LG A S+ +E VED L D +E++ +
Sbjct: 121 DLQQPHQSVGDV-VEFEFLDDG-GEVMLGQSASSEEMELNVEDEPEKDLNDTVEEDTSFW 294
Query: 67 SDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREV 101
+QH+ + N+ + L E++IR+V
Sbjct: 295 ENQHQVLQA---------NICTTSSL--ESRIRQV 366
>AV769753
Length = 496
Score = 26.9 bits (58), Expect = 7.5
Identities = 10/22 (45%), Positives = 17/22 (76%)
Frame = +1
Query: 3 KISPEIEIKMPMEAVPPVSADV 24
K S ++ IK+P E +PP+S+D+
Sbjct: 181 KQSHQVRIKIPHELLPPLSSDI 246
>AV414055
Length = 431
Score = 26.9 bits (58), Expect = 7.5
Identities = 23/82 (28%), Positives = 36/82 (43%), Gaps = 2/82 (2%)
Frame = +2
Query: 133 EKAISMVEALAVKLTQNEGELIQEKFEVKKLLN--FLKQASEDAKKLVNQEKSFACAEIE 190
E +S E + K+ + I+++ E LL F+K K V Q + I+
Sbjct: 185 EDLLSKPEQIRSKIVEGR---IRKRLEELALLEQPFIKNDKVPVKDWVKQTIATIGENIK 355
Query: 191 SARSVVLRIGEALEEQEQDSQA 212
R V +GE LE++ QD A
Sbjct: 356 VKRFVRFNLGEGLEKKSQDFAA 421
>TC18710
Length = 843
Score = 26.6 bits (57), Expect = 9.9
Identities = 11/24 (45%), Positives = 15/24 (61%)
Frame = -3
Query: 371 SHVEALLRKSNTDFHVVISQMNGK 394
SHV+A+L K DFH ++ GK
Sbjct: 835 SHVKAVLNKKRKDFHSEEEEIRGK 764
>BP030794
Length = 513
Score = 26.6 bits (57), Expect = 9.9
Identities = 22/104 (21%), Positives = 48/104 (46%)
Frame = +1
Query: 161 KKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDG 220
K++ +++ E +K ++ + R++ LEE + ++
Sbjct: 94 KEMASYIVALEEFRRKAFEED*FLNIRHPDGTRTIHELTKALLEEDLRPELEKDLKEFLA 273
Query: 221 LVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKL 264
VEE+QE +++L+ K ++E ++ + D + EK+ SIKL
Sbjct: 274 FVEEIQELSQLELMLLEEK-ESVEGRIKTIEDSL-EKTELSIKL 399
>TC19353 homologue to UP|AX2D_PHAAU (O24542) Auxin-induced protein 22D
(Indole-3-acetic acid induced protein ARG13), partial
(41%)
Length = 560
Score = 26.6 bits (57), Expect = 9.9
Identities = 15/35 (42%), Positives = 22/35 (62%), Gaps = 3/35 (8%)
Frame = +1
Query: 415 RGWITKAREIYSP---SMQLCGVRGDASNAAKALF 446
RGW+ I+SP SMQ+CGV+ S+A + +F
Sbjct: 223 RGWV-----IFSPKKQSMQICGVKLVYSDAIQEIF 312
>TC18974 weakly similar to PIR|S16321|S16321 light-induced protein CPRF-2 -
parsley {Petroselinum crispum;} , partial (17%)
Length = 507
Score = 26.6 bits (57), Expect = 9.9
Identities = 12/48 (25%), Positives = 24/48 (50%)
Frame = +2
Query: 171 SEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDV 218
SED ++ + + ACA + R +++ + E S+AS P ++
Sbjct: 338 SEDYHAILKTKLNLACAAVAMTRGSLVKSQDLATLPENGSRASNPSEI 481
>TC9109 homologue to UP|O23140 (O23140) AP47/50P, partial (18%)
Length = 614
Score = 26.6 bits (57), Expect = 9.9
Identities = 13/31 (41%), Positives = 17/31 (53%), Gaps = 7/31 (22%)
Frame = -3
Query: 412 KLCRGWITK-------AREIYSPSMQLCGVR 435
KLCR WIT+ + SPSM LC ++
Sbjct: 357 KLCRTWITRWNCAFNYLGSLQSPSMNLCHLK 265
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.131 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,553,122
Number of Sequences: 28460
Number of extensions: 75637
Number of successful extensions: 385
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 384
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 384
length of query: 490
length of database: 4,897,600
effective HSP length: 94
effective length of query: 396
effective length of database: 2,222,360
effective search space: 880054560
effective search space used: 880054560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0269a.3


