

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
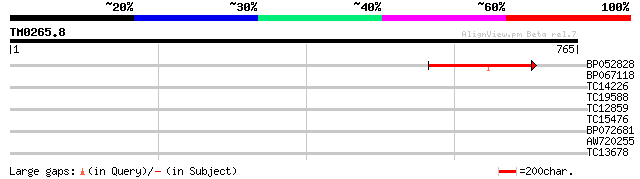
Query= TM0265.8
(765 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP052828 119 1e-27
BP067118 31 0.85
TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete 30 1.1
TC19588 30 1.4
TC12859 weakly similar to UP|CAE55354 (CAE55354) PE-PGRS FAMILY ... 30 1.9
TC15476 similar to UP|THRC_SOLTU (Q9MT28) Threonine synthase, ch... 30 1.9
BP072681 28 5.5
AW720255 28 7.2
TC13678 27 9.4
>BP052828
Length = 523
Score = 119 bits (299), Expect = 1e-27
Identities = 63/148 (42%), Positives = 94/148 (62%), Gaps = 3/148 (2%)
Frame = +2
Query: 566 PVNFGASKQVRALCDLGSSVNLMPLSMFERLNVGELKPTMMMLQLADRSIVTPWGVVEDV 625
P G +K + DLG+ +N+MP S++ L+ G L+ T +++QLA+RS P GVVEDV
Sbjct: 71 PCTIGDNKYENCMLDLGAGINVMPTSIYNILHPGPLQHTGLIVQLANRSNARPKGVVEDV 250
Query: 626 LVRVGEFEFPVDFVIIDMD---EDSKIPLILGRPFLATSQAKINVGKGTISLRVADEKIG 682
LV+V E FP DF I+DM+ + S+ P+ILGRPF+ T++ KI+V GT+S+ D
Sbjct: 251 LVQVNELIFPADFYILDMEGETKSSRAPIILGRPFMKTAKTKIDVDNGTMSMEFGDIIAK 430
Query: 683 FNIFDLKPKPVEKNDVFLVKMMEEWSDE 710
FNI D P+E + VF + ++ E D+
Sbjct: 431 FNIDDAMKHPLEDHSVFHIGVVSELVDD 514
>BP067118
Length = 554
Score = 30.8 bits (68), Expect = 0.85
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = +3
Query: 452 KKKQKEKEEEKRKVELEKKFTKVPFPPFPT 481
K K+KEK E+++K + E++ VP PP T
Sbjct: 108 KPKRKEKSEKEKKKQEEEQEQSVPIPPIQT 197
>TC14226 UP|Q9MBE4 (Q9MBE4) Cytochrome P450, complete
Length = 1878
Score = 30.4 bits (67), Expect = 1.1
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -1
Query: 268 WNMRWSSCQ*RLRCRF*RGSQGFEKFPK 295
WN+RW SC+ RC F RGS + P+
Sbjct: 393 WNLRWCSCR---RCTFQRGSVAGSQAPR 319
>TC19588
Length = 489
Score = 30.0 bits (66), Expect = 1.4
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 5/50 (10%)
Frame = -1
Query: 450 ENKKKQKEKEEEKRKVELE-----KKFTKVPFPPFPTNIAKRRLEKQFSK 494
ENK K KE+ K E ++ TKV FP +PT +A+ L + S+
Sbjct: 459 ENKSKLNCKEQAKIGQSQEGNAHSQRLTKVEFPKYPTTLARFPLPYELSR 310
>TC12859 weakly similar to UP|CAE55354 (CAE55354) PE-PGRS FAMILY PROTEIN,
partial (5%)
Length = 528
Score = 29.6 bits (65), Expect = 1.9
Identities = 18/38 (47%), Positives = 21/38 (54%), Gaps = 4/38 (10%)
Frame = +2
Query: 327 SEAIFKSKIP---VSEFRT-STESGSRQWQWEEELRRI 360
SE FKS IP E RT TES R W+W+ +R I
Sbjct: 407 SELKFKSSIPR*ASDE*RTRETESDRRSWRWQRRVRTI 520
>TC15476 similar to UP|THRC_SOLTU (Q9MT28) Threonine synthase, chloroplast
precursor (TS) , partial (19%)
Length = 658
Score = 29.6 bits (65), Expect = 1.9
Identities = 24/67 (35%), Positives = 35/67 (51%)
Frame = -3
Query: 524 EILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGS 583
+I+ K +L+ N VE EC I+ ++L KD + T+P N SKQ + LG+
Sbjct: 488 KIIYNKIQLACSN--VEQHTECQ*IIPKRLKANTKD--NTTVPKN---SKQAEQIAKLGT 330
Query: 584 SVNLMPL 590
NLM L
Sbjct: 329 LANLMIL 309
>BP072681
Length = 461
Score = 28.1 bits (61), Expect = 5.5
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +2
Query: 53 SIWRICH*RCTCSSRAVHQKLQYYMSSQPRACSFATFSFF 92
SI+ H C C+ H + + +P++CS +FSFF
Sbjct: 11 SIFIFIHTWCRCAGLPHHYIYKLTTNKRPKSCSNYSFSFF 130
>AW720255
Length = 532
Score = 27.7 bits (60), Expect = 7.2
Identities = 18/60 (30%), Positives = 26/60 (43%), Gaps = 9/60 (15%)
Frame = +2
Query: 449 GENKKKQKEKEEEKRKVELEKKFTKVP---------FPPFPTNIAKRRLEKQFSKFISMF 499
GEN + K K++ K K +LEK+ K PP +I R + QF +F
Sbjct: 206 GENNSESKSKKKLKMKKKLEKETEKADKRGLCYMSRIPPHMDHIKLRHILSQFGDIQRIF 385
>TC13678
Length = 548
Score = 27.3 bits (59), Expect = 9.4
Identities = 17/55 (30%), Positives = 22/55 (39%)
Frame = -2
Query: 55 WRICH*RCTCSSRAVHQKLQYYMSSQPRACSFATFSFFTKGHN*GVAQLPTSRQH 109
W IC T + + +HQ L PR+ F+F H Q P S QH
Sbjct: 499 WWICKATSTTNKKELHQDLSSLPGPSPRSMILYKFNFSHSSHG-LKEQSPLSVQH 338
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.337 0.147 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,058,103
Number of Sequences: 28460
Number of extensions: 167483
Number of successful extensions: 1161
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 1149
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1158
length of query: 765
length of database: 4,897,600
effective HSP length: 97
effective length of query: 668
effective length of database: 2,136,980
effective search space: 1427502640
effective search space used: 1427502640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0265.8


