

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.5
(404 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
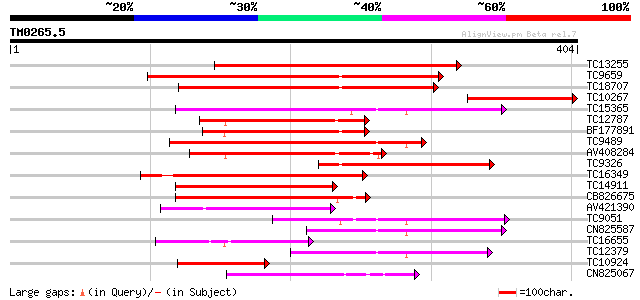
TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%) 350 2e-97
TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATP... 234 3e-62
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 169 1e-42
TC10267 similar to UP|Q7XXR9 (Q7XXR9) Katanin, partial (20%) 159 8e-40
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 146 7e-36
TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (T... 131 2e-31
BF177891 127 3e-30
TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase... 127 4e-30
AV408284 126 7e-30
TC9326 homologue to UP|Q93WX4 (Q93WX4) Suppressor of K+ transpor... 117 3e-27
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 113 5e-26
TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), pa... 108 2e-24
CB826675 103 4e-23
AV421390 80 5e-16
TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 ... 80 6e-16
CN825587 74 6e-14
TC16655 homologue to UP|O99018 (O99018) Chloroplast protease pre... 70 8e-13
TC12379 homologue to UP|O04327 (O04327) Cell division protein Ft... 67 7e-12
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 60 8e-10
CN825067 59 1e-09
>TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%)
Length = 532
Score = 350 bits (898), Expect = 2e-97
Identities = 176/176 (100%), Positives = 176/176 (100%)
Frame = +1
Query: 147 PKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVK 206
PKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVK
Sbjct: 4 PKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVK 183
Query: 207 VLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVL 266
VLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVL
Sbjct: 184 VLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVL 363
Query: 267 AATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQ 322
AATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQ
Sbjct: 364 AATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQ 531
>TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (74%)
Length = 1060
Score = 234 bits (596), Expect = 3e-62
Identities = 116/211 (54%), Positives = 154/211 (72%)
Frame = +1
Query: 99 ESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK 158
E E L L+ IIR P++KW + GLE+AK+ L+EAV++P+K+P++FTG PW+
Sbjct: 388 EDPEQAKLRAGLNSAIIREKPNIKWNDVAGLESAKQSLQEAVILPVKFPQFFTGKRRPWR 567
Query: 159 GILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPS 218
LL+GPPGTGK+ LAKAVATE +TFF++S+S + SKW G+SEKLV LFQ+AR APS
Sbjct: 568 AFLLYGPPGTGKSYLAKAVATEADSTFFSVSSSDLGSKWMGESEKLVSNLFQMARESAPS 747
Query: 219 TIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAA 278
IF+DEID++ QRGE +E EASRR+KTELL+QM G+ D+ V VLAATN P+ LD A
Sbjct: 748 IIFVDEIDSLCGQRGEG-NESEASRRIKTELLVQMQGVGNNDQKVLVLAATNTPYALDQA 924
Query: 279 MLRRLEKRILVPLPEPEARVAMFEELLPPQP 309
+ RR +KRI +PLP+ +AR MF+ L P
Sbjct: 925 IRRRFDKRIYIPLPDVKARQHMFKVHLGDTP 1017
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 169 bits (427), Expect = 1e-42
Identities = 90/187 (48%), Positives = 124/187 (66%), Gaps = 2/187 (1%)
Frame = +3
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTG-LLSPWKGILLFGPPGTGKTMLAKAVAT 179
V + I ++ K L+E V++P++ P F G LL P +GILLFGPPGTGKTMLA A+A
Sbjct: 192 VTFGDIGAMDEIKESLQELVMLPLRRPDLFKGGLLKPCRGILLFGPPGTGKTMLAXAIAN 371
Query: 180 ECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEH 239
E +F N+S S++ SKW G+ EK V+ LF LA AP+ IF+DE+D+++ QR EH
Sbjct: 372 EAGASFINVSMSTITSKWFGEDEKNVRALFSLAAKVAPTIIFVDEVDSMLGQRTRV-GEH 548
Query: 240 EASRRLKTELLIQMDG-LTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARV 298
EA R++K E + DG LT E + VLAATN P++LD A++RR E+RI+V LP E R
Sbjct: 549 EAMRKIKNEFMTHWDGLLTAPGEQILVLAATNRPFDLDEAIIRRFERRIMVGLPSVENRE 728
Query: 299 AMFEELL 305
+ + LL
Sbjct: 729 MILKTLL 749
>TC10267 similar to UP|Q7XXR9 (Q7XXR9) Katanin, partial (20%)
Length = 562
Score = 159 bits (402), Expect = 8e-40
Identities = 78/78 (100%), Positives = 78/78 (100%)
Frame = +1
Query: 327 SGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHL 386
SGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHL
Sbjct: 1 SGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSAHL 180
Query: 387 HAHKYDKFNADYGSQILQ 404
HAHKYDKFNADYGSQILQ
Sbjct: 181 HAHKYDKFNADYGSQILQ 234
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 146 bits (368), Expect = 7e-36
Identities = 91/241 (37%), Positives = 140/241 (57%), Gaps = 5/241 (2%)
Frame = +3
Query: 119 PDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 177
PD ++ I GL+ + +KE + +PIK+P+ F L ++ KG+LL+GPPGTGKT+LA+AV
Sbjct: 312 PDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAV 491
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARS 237
A TF +S S +V K+ G+ ++V+ LF +AR HAPS IF+DEID+I S R E+ S
Sbjct: 492 AHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARMESGS 671
Query: 238 EHEAS--RRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPE 293
S +R ELL Q+DG +++ + VL ATN LD A+LR R++++I P P
Sbjct: 672 GXGDSEVQRTMLELLNQLDGFEASNK-IKVLMATNRIDILDQALLRPGRIDRKIEFPNPN 848
Query: 294 PEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQR 353
E+R + + I + + G SG++++ +C E M LR + Q
Sbjct: 849 EESRQDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHVTQE 1028
Query: 354 E 354
+
Sbjct: 1029D 1031
>TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (TAT-binding
homolog 4), partial (20%)
Length = 389
Score = 131 bits (329), Expect = 2e-31
Identities = 68/123 (55%), Positives = 85/123 (68%), Gaps = 2/123 (1%)
Frame = +3
Query: 136 LKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSV 193
LKE V++P+K P+ F L P KGILLFGPPGTGKTMLAKAVATE F NIS SS+
Sbjct: 12 LKELVMLPLKRPELFCKGQLTKPCKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSI 191
Query: 194 VSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQM 253
SKW G+ EK VK +F LA +PS IF+DE+D ++ +R E EHEA R++ E ++
Sbjct: 192 TSKWFGEGEKYVKAVFSLASKISPSVIFVDEVDXMLXRR-ENPGEHEAMRKMXNEFMVHW 368
Query: 254 DGL 256
DGL
Sbjct: 369 DGL 377
>BF177891
Length = 390
Score = 127 bits (320), Expect = 3e-30
Identities = 64/121 (52%), Positives = 86/121 (70%), Gaps = 2/121 (1%)
Frame = +1
Query: 138 EAVVMPIKYPKYFT--GLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVS 195
E V++P++ P+ FT LL P KGILLFGPPGTGKT++AKA+ATE F ++S S++ S
Sbjct: 16 ELVILPMRRPELFTHGNLLRPVKGILLFGPPGTGKTLMAKALATEAGANFISVSGSTLTS 195
Query: 196 KWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDG 255
KW GD+EKL K LF A AP IF+DE+D+++ RG A EHEA+RR++ E + DG
Sbjct: 196 KWFGDAEKLTKALFSFASKLAPVIIFVDEVDSLLGARGGA-FEHEATRRMRNEFMAAWDG 372
Query: 256 L 256
L
Sbjct: 373 L 375
>TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase subunit
RPT2a, partial (69%)
Length = 916
Score = 127 bits (318), Expect = 4e-30
Identities = 74/187 (39%), Positives = 116/187 (61%), Gaps = 4/187 (2%)
Frame = +1
Query: 115 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTML 173
+ +P + I GL+ + +KEAV +P+ +P+ + + + P KG++L+G PGTGKT+L
Sbjct: 355 VEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLL 534
Query: 174 AKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG 233
AKAVA TF + S ++ K+ GD KLV+ LF++A +PS +F+DEIDA+ ++R
Sbjct: 535 AKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRY 714
Query: 234 EARSEHEAS-RRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVP 290
+A S E +R ELL Q+DG + V V+ ATN LD A+LR R++++I P
Sbjct: 715 DAHSGGEREIQRTMLELLNQLDGFDSRGD-VKVILATNRIESLDPALLRPGRIDRKIEFP 891
Query: 291 LPEPEAR 297
LP+ + R
Sbjct: 892 LPDTKTR 912
>AV408284
Length = 425
Score = 126 bits (316), Expect = 7e-30
Identities = 70/144 (48%), Positives = 95/144 (65%), Gaps = 4/144 (2%)
Frame = +1
Query: 129 LENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVATECKTTFF 186
LE+ K L E V++P+K P F+ LL P KG+LL+GPPGTGKTMLAKA+A E F
Sbjct: 1 LESIKEALFELVILPLKRPDLFSHGKLLGPQKGVLLYGPPGTGKTMLAKAIAKESGAVFI 180
Query: 187 NISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLK 246
N+ S+++SKW GD++KLV +F LA P+ IF+DE+D+ + QR ++HEA +K
Sbjct: 181 NVRISNLMSKWFGDAQKLVAAVFSLAHKLQPAIIFIDEVDSFLGQR--RTTDHEALLNMK 354
Query: 247 TELLIQMDGLTRTDE--LVFVLAA 268
TE + DG T TD+ V VLAA
Sbjct: 355 TEFMALWDGFT-TDQNARVMVLAA 423
>TC9326 homologue to UP|Q93WX4 (Q93WX4) Suppressor of K+ transport growth
defect-like protein (Fragment), partial (73%)
Length = 894
Score = 117 bits (293), Expect = 3e-27
Identities = 62/126 (49%), Positives = 87/126 (68%), Gaps = 1/126 (0%)
Frame = +2
Query: 221 FLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAML 280
F+DEID++ QRGE +E EASRR+KTELL+QM G+ D+ V VLAATN P+ LD A+
Sbjct: 20 FVDEIDSLCGQRGEG-NESEASRRIKTELLVQMQGVGNNDQKVLVLAATNTPYALDQAIR 196
Query: 281 RRLEKRILVPLPEPEARVAMFEELLPPQPDE-ESIPYDLLVNQTEGYSGSDIRLLCKEVA 339
RR +KRI +PLP+ +AR MF+ L P ++ L ++TEG+SGSDI + K+V
Sbjct: 197 RRFDKRIYIPLPDLKARQHMFKVHLGDTPHNLNESDFEYLASRTEGFSGSDISVCVKDVL 376
Query: 340 MQPLRR 345
+P+R+
Sbjct: 377 FEPVRK 394
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 113 bits (283), Expect = 5e-26
Identities = 62/164 (37%), Positives = 102/164 (61%), Gaps = 2/164 (1%)
Frame = +3
Query: 94 LLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL 153
+LPP + + L++S PDV + I G + K+ ++EAV +P+ + + + +
Sbjct: 429 VLPPEADSSISLLSQS-------EKPDVTYNDIGGCDIQKQEIREAVELPLTHHELYKQI 587
Query: 154 -LSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLA 212
+ P +G+LL+GPPGTGKTMLAKAVA F + S V K+ G+ ++V+ +F+LA
Sbjct: 588 GIDPPRGVLLYGPPGTGKTMLAKAVANHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLA 767
Query: 213 RHHAPSTIFLDEIDAIISQRGEARSEHEAS-RRLKTELLIQMDG 255
+ +AP+ IF+DE+DAI + R +A++ + +R+ ELL QMDG
Sbjct: 768 KENAPAIIFIDEVDAIATARFDAQTGADREVQRILMELLNQMDG 899
>TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), partial (32%)
Length = 1418
Score = 108 bits (270), Expect = 2e-24
Identities = 50/115 (43%), Positives = 74/115 (63%)
Frame = +2
Query: 119 PDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVA 178
P+VKWE + GLE+ K+ + + V +P+ + F L G+LL+GPPGTGKT+LAKAVA
Sbjct: 209 PNVKWEDVGGLEDVKKSILDTVQLPLLHKDLFASGLRKRSGVLLYGPPGTGKTLLAKAVA 388
Query: 179 TECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG 233
TEC F ++ +++ + G+SEK V+ +FQ AR P IF DE+D++ G
Sbjct: 389 TECSLNFLSVKGPELINMYIGESEKNVRDIFQKARSARPCVIFFDELDSLAPASG 553
>CB826675
Length = 551
Score = 103 bits (258), Expect = 4e-23
Identities = 58/142 (40%), Positives = 87/142 (60%), Gaps = 3/142 (2%)
Frame = +1
Query: 119 PDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 177
P + + GLE + L EA+V+P+ + + F L + P KG+LL+GPPGTGKT++A+A
Sbjct: 121 PTEDYNDVGGLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLMARAC 300
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQR--GEA 235
A + TF ++ +V + GD KLV+ FQLA+ +P IF+DEIDAI ++R E
Sbjct: 301 AAQTNATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEV 480
Query: 236 RSEHEASRRLKTELLIQMDGLT 257
+ E R + ELL Q+DG +
Sbjct: 481 SGDREVQRTM-LELLNQLDGFS 543
>AV421390
Length = 409
Score = 80.5 bits (197), Expect = 5e-16
Identities = 51/126 (40%), Positives = 75/126 (59%), Gaps = 1/126 (0%)
Frame = +2
Query: 108 ESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPW-KGILLFGPP 166
+ L+++++ ++ +KG ++AK+ L+E VV ++ P FT L KGILL G P
Sbjct: 23 KELNKEVMPEKNVKTFKDVKGCDDAKQELEE-VVEYLRNPAKFTRLGGKLPKGILLTGAP 199
Query: 167 GTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEID 226
GTGKT+LAKA+A E FF + S + G + V+ LFQ A+ AP IF+DEID
Sbjct: 200 GTGKTLLAKAIAGEAGVPFFYRAGSEFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEID 379
Query: 227 AIISQR 232
A+ S R
Sbjct: 380 AVGSTR 397
>TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (46%)
Length = 867
Score = 80.1 bits (196), Expect = 6e-16
Identities = 54/173 (31%), Positives = 91/173 (52%), Gaps = 4/173 (2%)
Frame = +2
Query: 188 ISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGE--ARSEHEASRRL 245
+ S +V K+ G+ ++V+ LFQ+AR +F DE+DAI R + ++E R +
Sbjct: 8 VIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAIGGARFDDGVGGDNEVQRTM 187
Query: 246 KTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPEPEARVAMFEE 303
E++ Q+DG + VL ATN P LD A+LR RL++++ LP+ E+R +F+
Sbjct: 188 -LEIVNQLDGFDARGN-IKVLMATNRPDTLDPALLRPGRLDRKVEFGLPDLESRTQIFKI 361
Query: 304 LLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDL 356
E I ++LL +G+DIR +C E M +R + +++ L
Sbjct: 362 HTRTMNCERDIRFELLSRLCPNSTGADIRSVCTEAGMYAIRARRKTVTEKDFL 520
>CN825587
Length = 663
Score = 73.6 bits (179), Expect = 6e-14
Identities = 46/146 (31%), Positives = 81/146 (54%), Gaps = 3/146 (2%)
Frame = +2
Query: 212 ARHHAPSTIFLDEIDAIISQRGEA-RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATN 270
A+ AP +F+DEIDA+ QRG ++ + +LL +MDG + V VLAATN
Sbjct: 2 AKSKAPCIVFIDEIDAVGRQRGAGLGGGNDEREQTINQLLTEMDGFSGNSG-VIVLAATN 178
Query: 271 LPWELDAAMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSG 328
P LD+A+LR R ++++ V P+ RV + + + + + +D + +T G++G
Sbjct: 179 RPDVLDSALLRPGRFDRQVTVDRPDVAGRVKILQVHSRGKALAKDVDFDKIARRTPGFTG 358
Query: 329 SDIRLLCKEVAMQPLRRLMSQLEQRE 354
+D++ L E A+ RR + ++ + E
Sbjct: 359 ADLQNLMNEAAILAARRDLKEISKDE 436
>TC16655 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (39%)
Length = 934
Score = 69.7 bits (169), Expect = 8e-13
Identities = 44/114 (38%), Positives = 66/114 (57%), Gaps = 2/114 (1%)
Frame = +3
Query: 105 TLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFT--GLLSPWKGILL 162
T +S ++ + + V ++ + G++ AK+ E V +K P+ FT G P KG+LL
Sbjct: 597 TFGQSKAKFQMEPNTGVTFDDVAGVDEAKQDFMEVVEF-LKKPERFTAVGARIP-KGVLL 770
Query: 163 FGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHA 216
GPPGTGKT+LAKA+A E FF+IS S V + G V+ LF+ A+ +A
Sbjct: 771 IGPPGTGKTLLAKAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENA 932
>TC12379 homologue to UP|O04327 (O04327) Cell division protein FtsH isolog,
partial (15%)
Length = 505
Score = 66.6 bits (161), Expect = 7e-12
Identities = 42/147 (28%), Positives = 78/147 (52%), Gaps = 3/147 (2%)
Frame = +1
Query: 201 SEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASR-RLKTELLIQMDGLTRT 259
S L + L+Q A+ +APS +F+DE+DA+ +RG + R +LL+ +DG
Sbjct: 46 SLSLSRALYQEAKENAPSVVFIDELDAVGRERGLIKGSGGQERDATLNQLLVCLDGFEGR 225
Query: 260 DELVFVLAATNLPWELDAAMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYD 317
E V +A+TN P LD A++R R +++I +P P R+ + + +P + Y
Sbjct: 226 GE-VITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRIEILKVHARKKPIAGDVDYM 402
Query: 318 LLVNQTEGYSGSDIRLLCKEVAMQPLR 344
+ + T+G G+++ + + A+ +R
Sbjct: 403 AVASMTDGMVGAELANIVEVAAINMMR 483
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 59.7 bits (143), Expect = 8e-10
Identities = 27/67 (40%), Positives = 46/67 (68%), Gaps = 1/67 (1%)
Frame = +2
Query: 120 DVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAVA 178
+V + ++ GL + R L+E++ +P+ P+ F + + P KG+LL+GPPGTGKT+LA+A+A
Sbjct: 446 NVSYSAVGGLSDQIRELRESIELPLMNPELFLRVGIKPPKGVLLYGPPGTGKTLLARAIA 625
Query: 179 TECKTTF 185
+ F
Sbjct: 626 SNIDANF 646
>CN825067
Length = 687
Score = 58.9 bits (141), Expect = 1e-09
Identities = 39/138 (28%), Positives = 71/138 (51%)
Frame = +1
Query: 155 SPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARH 214
+P++ +L +GPPGTGKTM A+ +A + + ++ V K+ ++ +
Sbjct: 184 APFRNMLFYGPPGTGKTMAARELARKSGLDYALMTGGDVAPLGSQAVTKIHELFDWSKKS 363
Query: 215 HAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWE 274
+F+DE DA + +R + EA R LL + ++ ++V L ATN P +
Sbjct: 364 KRGLLLFIDEADAFLCERNKTYMS-EAQRSALNALLFRTG--DQSKDIVLAL-ATNRPGD 531
Query: 275 LDAAMLRRLEKRILVPLP 292
LD+A+ R+++ + PLP
Sbjct: 532 LDSAVADRIDEVLEFPLP 585
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.315 0.133 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,243,018
Number of Sequences: 28460
Number of extensions: 77957
Number of successful extensions: 461
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 443
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 444
length of query: 404
length of database: 4,897,600
effective HSP length: 92
effective length of query: 312
effective length of database: 2,279,280
effective search space: 711135360
effective search space used: 711135360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0265.5


