

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
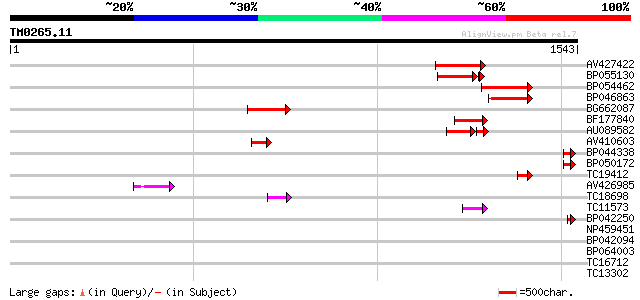
Query= TM0265.11
(1543 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV427422 241 8e-64
BP055130 140 3e-36
BP054462 147 1e-35
BP046863 115 7e-26
BG662087 99 4e-21
BF177840 96 6e-20
AU089582 75 7e-18
AV410603 75 6e-14
BP044338 60 2e-09
BP050172 60 2e-09
TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%) 54 2e-07
AV426985 52 9e-07
TC18698 45 7e-05
TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase regi... 42 6e-04
BP042250 42 7e-04
NP459451 NDX3 protein [Lotus japonicus] 41 0.002
BP042094 38 0.011
BP064003 36 0.053
TC16712 UP|ATPI_LOTJA (Q9BBS5) Chloroplast ATP synthase a chain ... 35 0.12
TC13302 32 0.77
>AV427422
Length = 417
Score = 241 bits (614), Expect = 8e-64
Identities = 102/137 (74%), Positives = 127/137 (92%)
Frame = +1
Query: 1158 GLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDP 1217
GLPKSKGYEA+LVVVDRLSK+SHF+ LKHPYTAK IA++FVRE+VRLHG+P ++ SDRDP
Sbjct: 7 GLPKSKGYEAVLVVVDRLSKFSHFVPLKHPYTAKVIADIFVREVVRLHGVPLSIVSDRDP 186
Query: 1218 IFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPW 1277
+F+S+FW ELFK+QGTKLKMS++YHPE+DGQTEV+NRCLE+YLRCF +D PK+W+ WVPW
Sbjct: 187 LFMSNFWKELFKMQGTKLKMSTAYHPESDGQTEVVNRCLETYLRCFIADQPKSWAHWVPW 366
Query: 1278 AEFWYNSTFHCSIGQTP 1294
AE+WYN+++H S GQTP
Sbjct: 367 AEYWYNTSYHVSTGQTP 417
>BP055130
Length = 567
Score = 140 bits (353), Expect(2) = 3e-36
Identities = 63/108 (58%), Positives = 76/108 (70%)
Frame = +2
Query: 1164 GYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHF 1223
G+ I+VVVDRLSKY HF + YT+ +AE FV IV+LHG+P + SDRD F S F
Sbjct: 179 GFTVIIVVVDRLSKYGHFAPHRANYTSSQVAETFVSTIVKLHGMPRAIVSDRDKAFTSAF 358
Query: 1224 WSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTW 1271
W FKL GT L MSSSYHP+TDGQTE +N+CLE YLRCF + P+ W
Sbjct: 359 WKHFFKLHGTTLNMSSSYHPQTDGQTEALNKCLELYLRCFVHETPRLW 502
Score = 30.0 bits (66), Expect(2) = 3e-36
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +3
Query: 1275 VPWAEFWYNSTFHCSIG 1291
+PWAE+ YNS+F SIG
Sbjct: 516 LPWAEYCYNSSFQTSIG 566
>BP054462
Length = 422
Score = 147 bits (371), Expect = 1e-35
Identities = 70/140 (50%), Positives = 102/140 (72%)
Frame = +1
Query: 1283 NSTFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQE 1342
NS ++ S +PF+ +YGR+ P +++ T +K+ AV RDE L LR +L ++Q+
Sbjct: 1 NSNYNRSAKMSPFQALYGREPPVLLQGTTIPSKIAAVNDLQVGRDELLSDLRANLLKSQD 180
Query: 1343 QMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAY 1402
M +YANKKRRD+ + IG+ VFLKL+P+R+ S+ K++N+KL+ R+YGP+ I KIG VAY
Sbjct: 181 MMRTYANKKRRDVDYQIGDEVFLKLQPYRRRSLAKKMNEKLSPRYYGPYPIVAKIGAVAY 360
Query: 1403 KLKLPVESRIHPVFHVSLLK 1422
+L+LP SR+HPVFHVSLLK
Sbjct: 361 RLELPAHSRVHPVFHVSLLK 420
>BP046863
Length = 580
Score = 115 bits (287), Expect = 7e-26
Identities = 57/120 (47%), Positives = 85/120 (70%)
Frame = +1
Query: 1304 PPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWV 1363
PP F + + V AV + + ERD L +LR +L +AQ+QM + ANK RR + + +G WV
Sbjct: 226 PP--HFCSIKLSVEAVNMLIEERDALLLELRGNLLKAQDQMRAQANKHRRYVDYQVGNWV 399
Query: 1364 FLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKR 1423
FLKL+P++ ++ +R NQKL+ RFYGPF++ +++ VAY L L ESR+HPVFH+SLL++
Sbjct: 400 FLKLQPYKLQNLAQRKNQKLSPRFYGPFKVLERVVQVAY*LDLXSESRVHPVFHLSLLEK 579
>BG662087
Length = 373
Score = 99.4 bits (246), Expect = 4e-21
Identities = 48/117 (41%), Positives = 71/117 (60%)
Frame = +1
Query: 646 WRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTT 705
WRM VDY LNKA D YP+P +D+L+D + + S +D SGYHQI++H D KT
Sbjct: 22 WRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDEDKTA 201
Query: 706 FRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEH 762
F T +Y Y +PFGL NA AT+Q +M+ +F + + + V+ D++++ S H
Sbjct: 202 FMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BF177840
Length = 410
Score = 95.5 bits (236), Expect = 6e-20
Identities = 43/91 (47%), Positives = 58/91 (63%)
Frame = +2
Query: 1210 TVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPK 1269
++ SDRD F+SHFW L+ GTKL S++ HP+TDGQTEV+N+ L + LR + K
Sbjct: 14 SIVSDRDTKFISHFWRTLWGKVGTKLLYSTTCHPQTDGQTEVVNKTLSTLLRSVLERNLK 193
Query: 1270 TWSFWVPWAEFWYNSTFHCSIGQTPFEVVYG 1300
W W+P EF YN H + +PFE+VYG
Sbjct: 194 MWETWLPHIEFAYNRVVHSTTKHSPFEIVYG 286
>AU089582
Length = 383
Score = 75.1 bits (183), Expect(2) = 7e-18
Identities = 35/77 (45%), Positives = 49/77 (63%)
Frame = +2
Query: 1190 AKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQT 1249
A A++++ EIV LHG+P ++ SDR F SHFW GT+LKMS+++HP+TDGQ+
Sbjct: 11 ASQYAKIYLDEIVSLHGVPVSIISDRGAQFTSHFWRSFQTALGTRLKMSTAFHPQTDGQS 190
Query: 1250 EVINRCLESYLRCFASD 1266
E + LE LR D
Sbjct: 191 ERTIQILEDMLRACVXD 241
Score = 33.9 bits (76), Expect(2) = 7e-18
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 1270 TWSFWVPWAEFWYNSTFHCSIGQTPFEVVYGRK 1302
+W ++ EF YN+++ SI PFE +YGR+
Sbjct: 252 SWDQYLSLMEFAYNNSYRSSI*MAPFEALYGRR 350
>AV410603
Length = 162
Score = 75.5 bits (184), Expect = 6e-14
Identities = 34/54 (62%), Positives = 42/54 (76%)
Frame = +1
Query: 658 ATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNG 711
+T+ D +P+P VDELLDEL G+ FSK+DL+SGYHQI V ED KT FRTH+G
Sbjct: 1 STVKDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>BP044338
Length = 581
Score = 60.5 bits (145), Expect = 2e-09
Identities = 27/33 (81%), Positives = 30/33 (90%)
Frame = -1
Query: 1508 ERDSNTKWGMGLGKPKIWKVYVRKKGKGAEVSK 1540
+RDSNTK MGLGKPK+WKVY RKKGKGAEVS+
Sbjct: 329 DRDSNTKGEMGLGKPKVWKVYERKKGKGAEVSQ 231
>BP050172
Length = 594
Score = 60.5 bits (145), Expect = 2e-09
Identities = 27/33 (81%), Positives = 30/33 (90%)
Frame = -1
Query: 1508 ERDSNTKWGMGLGKPKIWKVYVRKKGKGAEVSK 1540
+RDSNTK MGLGKPK+WKVY RKKGKGAEVS+
Sbjct: 264 DRDSNTKGEMGLGKPKVWKVYERKKGKGAEVSQ 166
>TC19412 similar to UP|Q84KB0 (Q84KB0) Pol protein, partial (7%)
Length = 519
Score = 53.5 bits (127), Expect = 2e-07
Identities = 22/43 (51%), Positives = 36/43 (83%), Gaps = 1/43 (2%)
Frame = +3
Query: 1382 KLAARFYGPFQIEDKIGTVAYKLKLPVE-SRIHPVFHVSLLKR 1423
KL+ RF GPF++ +++G+V+Y+L LP + S +HPVFHVS+L++
Sbjct: 48 KLSPRFIGPFEVLERVGSVSYRLALPPDLSAVHPVFHVSMLRK 176
>AV426985
Length = 422
Score = 51.6 bits (122), Expect = 9e-07
Identities = 30/114 (26%), Positives = 52/114 (45%)
Frame = +1
Query: 336 HKCPERAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGEHQTQTM 395
HKCP + + +L L D D+ ++ I ++ VE + G T+
Sbjct: 91 HKCPNKYLMMLQLTDENDSDDTSN----QQVTTTISDDSPVEDHHFSLNAMRGFTGVGTI 258
Query: 396 KLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVT 449
+ GQ+ N+ + +L+D G+S F+ P I L L I + +G+G K+ T
Sbjct: 259 RFTGQIGNISVQVLVDGGSSDCFLQPHIAKFLQLPIESKPNFKVLVGNGQKMET 420
>TC18698
Length = 808
Score = 45.4 bits (106), Expect = 7e-05
Identities = 24/64 (37%), Positives = 36/64 (55%)
Frame = -2
Query: 703 KTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEH 762
KTT + + +Y Y VMP GL N T+Q +M+ IF + K V V+ +D+++ S H
Sbjct: 801 KTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE*FH 622
Query: 763 RDHL 766
R L
Sbjct: 621 RGDL 610
>TC11573 similar to UP|Q9M6N4 (Q9M6N4) Pol protein integrase region
(Fragment), partial (10%)
Length = 572
Score = 42.4 bits (98), Expect = 6e-04
Identities = 22/69 (31%), Positives = 37/69 (52%)
Frame = +2
Query: 1232 GTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTFHCSIG 1291
G +++ SS HP+T+GQTE N+ + ++ + W +P + YN+ SI
Sbjct: 62 GIQMRFSSVKHPQTNGQTEAANKVILKGIKRRLYEAEGRWIDELPIVLWSYNTMPQSSIK 241
Query: 1292 QTPFEVVYG 1300
+TPF + YG
Sbjct: 242 ETPF*LTYG 268
>BP042250
Length = 370
Score = 42.0 bits (97), Expect = 7e-04
Identities = 18/24 (75%), Positives = 20/24 (83%)
Frame = +1
Query: 1517 MGLGKPKIWKVYVRKKGKGAEVSK 1540
+GLGKP IWKVY RKK K AE+SK
Sbjct: 295 LGLGKPTIWKVYTRKKEKAAELSK 366
>NP459451 NDX3 protein [Lotus japonicus]
Length = 665
Score = 40.8 bits (94), Expect = 0.002
Identities = 20/45 (44%), Positives = 25/45 (55%)
Frame = +3
Query: 1477 DDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIERDSNTKWGMGLGK 1521
D+ TWEDN + QFP F LEDK GGI R + + G + K
Sbjct: 492 DEATWEDNITIKSQFPSFNLEDKVDLSAGGIVRPWDFQAGPNVTK 626
>BP042094
Length = 409
Score = 38.1 bits (87), Expect = 0.011
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 8/67 (11%)
Frame = -1
Query: 1479 VTWEDNEVLAGQFPEFCLEDKAVSKEGGIER--------DSNTKWGMGLGKPKIWKVYVR 1530
++W D L QFP F LEDKA + G +R + G +P W VY R
Sbjct: 409 ISWVDEPALKCQFPSFSLEDKAAAIGGXSDRIPGPVDYENQGEVLGQSSKRPTTWLVYSR 230
Query: 1531 KKGKGAE 1537
++ G +
Sbjct: 229 RRKIGTD 209
>BP064003
Length = 509
Score = 35.8 bits (81), Expect = 0.053
Identities = 30/104 (28%), Positives = 52/104 (49%), Gaps = 7/104 (6%)
Frame = +1
Query: 949 LGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLG---YQFEVKYKPGQENKAADALSR 1005
LG F + TD+ + FL QK +P + W L G + V+ + Q NK ADALSR
Sbjct: 112 LGSRFVIKTDNVATSSFLTQK-RAPTRARWQDFLSGGVRHGPRVQAREDQSNKVADALSR 288
Query: 1006 RGD----ESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDVQ 1045
+ + + + + +S +++ E + DP + LL++V+
Sbjct: 289 KAELASNKLEEIADMSQLKGAIPERIREGLEKDPVAQTLLKNVR 420
>TC16712 UP|ATPI_LOTJA (Q9BBS5) Chloroplast ATP synthase a chain precursor
(ATPase subunit IV) , complete
Length = 851
Score = 34.7 bits (78), Expect = 0.12
Identities = 16/58 (27%), Positives = 32/58 (54%), Gaps = 1/58 (1%)
Frame = +3
Query: 1256 LESYLRCFASDHPKTWSFWVPWAEF-WYNSTFHCSIGQTPFEVVYGRKAPPIIRFLTN 1312
+ S CF S +++ +VPW + WY+S++ C ++ + ++GR + + RF N
Sbjct: 594 ISSSCSCFFSTFSGSYTCYVPWIIYKWYSSSYFCYFSRSLYR*IHGRSS--LTRFQNN 761
>TC13302
Length = 533
Score = 32.0 bits (71), Expect = 0.77
Identities = 21/65 (32%), Positives = 31/65 (47%)
Frame = -2
Query: 1339 RAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIG 1398
+A QM + RRD+ +G+ V P +R+N KL RFYG ++ +KI
Sbjct: 271 KATNQMRQQV-RHRRDIQLHVGDMV*SN-NPTL*TRAYQRVNHKLCPRFYGTHEMVEKIN 98
Query: 1399 TVAYK 1403
V K
Sbjct: 97 PVV*K 83
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.324 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,015,517
Number of Sequences: 28460
Number of extensions: 388972
Number of successful extensions: 2210
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 2067
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2136
length of query: 1543
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1441
effective length of database: 1,994,680
effective search space: 2874333880
effective search space used: 2874333880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0265.11


