

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0263.7
(787 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
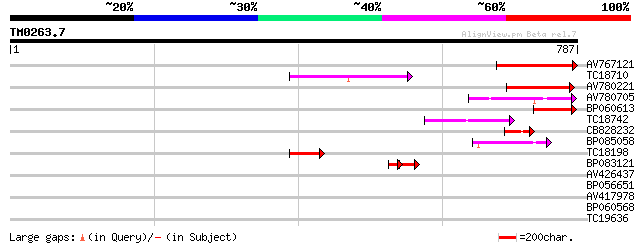
Sequences producing significant alignments: (bits) Value
AV767121 126 2e-29
TC18710 114 6e-26
AV780221 97 8e-21
AV780705 66 2e-11
BP060613 65 5e-11
TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%) 59 4e-09
CB828232 57 9e-09
BP085058 44 1e-04
TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, p... 42 4e-04
BP083121 33 7e-04
AV426437 36 0.021
BP056651 34 0.10
AV417978 30 1.1
BP060568 30 1.9
TC19636 28 5.6
>AV767121
Length = 525
Score = 126 bits (316), Expect = 2e-29
Identities = 64/113 (56%), Positives = 82/113 (71%), Gaps = 1/113 (0%)
Frame = +2
Query: 676 LSL*KHKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRG- 734
LS K K++SFGGRI LI+SVL++LPLFF SFFK P + C ++ R FLWGG E
Sbjct: 2 LSKWKQKTVSFGGRIFLIQSVLTALPLFFLSFFKLPIGVGKSCVRLMRNFLWGGSENENK 181
Query: 735 VAWVKWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
+A VKW ++CK KE GGLG+KDL +FNKALL K RWR L E + LW +V+E++
Sbjct: 182 IA*VKWTDVCKPKELGGLGIKDLFTFNKALLGK*RWRYLTEPDSLWRRVIEAQ 340
>TC18710
Length = 843
Score = 114 bits (285), Expect = 6e-26
Identities = 59/174 (33%), Positives = 96/174 (54%), Gaps = 3/174 (1%)
Frame = +2
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAFIGGR 448
N + +TLIPKV +P+ + +RPIS +YK+++K+ RL++ + +I Q+ FI GR
Sbjct: 302 NETLVTLIPKVPHPESINQFRPISGCTFLYKVISKIFVARLKDAMPDLISPMQSGFIQGR 481
Query: 449 FMLDSVVVANEVVEEAKRRK---KECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIH 505
+ D++++ E R + + K+D KAYD V+W FL + GFS W+
Sbjct: 482 QIQDNLLIVQEAFHAINRPGALGRNHSIIKLDMNKAYDRVEWKFLESSLLAFGFSTNWVK 661
Query: 506 WIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQA 559
I + S + +NG + RGLRQGDP +P+LFL E L+ L++ +
Sbjct: 662 MIMILVSGVSYNYKINGVVGPKLLPQRGLRQGDPFSPYLFLFTMEVLSLLIQNS 823
>AV780221
Length = 461
Score = 97.4 bits (241), Expect = 8e-21
Identities = 50/97 (51%), Positives = 65/97 (66%), Gaps = 2/97 (2%)
Frame = +3
Query: 690 ICLIKSVLSSLPLFFFSFFKAPK--CIIDQCNKIQRKFLWGGEEGRGVAWVKWEEICKKK 747
+CLI+SVL +LPLF FFK P + N F E G+ +AWVKW +C+ K
Sbjct: 3 LCLIRSVLIALPLFHLPFFKLPTGGFYTNVNN**DPFFGREVEGGKKIAWVKWSTVCRPK 182
Query: 748 EEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVL 784
+EGGLGV++L FNKALL KWRWR+L+ER+ LW +VL
Sbjct: 183 DEGGLGVQNLGLFNKALLGKWRWRMLKERDGLWYKVL 293
>AV780705
Length = 524
Score = 65.9 bits (159), Expect = 2e-11
Identities = 45/156 (28%), Positives = 74/156 (46%), Gaps = 7/156 (4%)
Frame = -2
Query: 638 MLNCKVMGVPFTYLGIPV--GANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKS 695
+L+ + P YLG+P G N + W + +K++ KL K L+ G+ LIK+
Sbjct: 520 LLHMPIWDDPGHYLGLPAIWGRNKSHSLVW--IEEKVKEKLEGWKETLLNQAGKEVLIKA 347
Query: 696 VLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-----GGEEGRGVAWVKWEEICKKKEEG 750
++ ++P + + PK + + F W G E G K KE+G
Sbjct: 346 IIQAIPSYAMTIVHFPKTFCNSLYALVADFWWKSQGEGAVEYTGAVGTK*P---LSKEKG 176
Query: 751 GLGVKDLESFNKALLVKWRWRLLREREWLWCQVLES 786
G+G +DL + N A L + WR+L E LW +V++S
Sbjct: 175 GVGFRDLRTQNLAFLARQAWRVLTNPEALWVRVMKS 68
>BP060613
Length = 378
Score = 64.7 bits (156), Expect = 5e-11
Identities = 28/60 (46%), Positives = 39/60 (64%)
Frame = +1
Query: 727 WGGEEGRGVAWVKWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLES 786
W + RGV W KW+ + + K+EGGLG KD E N+ALL K WR+L + LW Q+L++
Sbjct: 49 WSSKGTRGVHWRKWDLLTELKDEGGLGFKDFEIQNQALLAKQAWRILHNPDALWVQILKA 228
>TC18742 similar to UP|Q9FZG5 (Q9FZG5) T2E6.4, partial (3%)
Length = 912
Score = 58.5 bits (140), Expect = 4e-09
Identities = 40/125 (32%), Positives = 67/125 (53%)
Frame = -3
Query: 576 EISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIF 635
EIS L FAD+ + F ++I ++ ++L F+ +SGL N KS++ + + + QI
Sbjct: 376 EISHLCFADNLMVFSNGYLESIAIINNVLHIFQHLSGLTPNPAKSEVFILVKETTKNQI- 200
Query: 636 AAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKS 695
ML K +P LG+P KA+ + ++ + KL +SLS+ GR+ LI S
Sbjct: 199 TNMLGYKEGKLPVR*LGVPQLTTKLKASDCKVLVDRKPSKLQHWTGRSLSYAGRLQLINS 20
Query: 696 VLSSL 700
+L S+
Sbjct: 19 ILFSM 5
>CB828232
Length = 532
Score = 57.4 bits (137), Expect = 9e-09
Identities = 26/42 (61%), Positives = 30/42 (70%)
Frame = +1
Query: 687 GGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWG 728
G R+C K VLSSLPLFF SFFK KC+ C +IQ +FLWG
Sbjct: 409 GARVCCFKMVLSSLPLFFLSFFKI-KCVAKVCKQIQSRFLWG 531
>BP085058
Length = 452
Score = 43.5 bits (101), Expect = 1e-04
Identities = 31/115 (26%), Positives = 52/115 (44%), Gaps = 5/115 (4%)
Frame = +2
Query: 643 VMGVPFT-----YLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKSVL 697
+ +PFT Y+ P+ + + ++ +L+ *K L+ R+ L K VL
Sbjct: 101 ITSIPFTDNIGKYMSFPIHQGHV*KEDFVFTLDRVISRLAC*KAYLLNKPARLTLAKPVL 280
Query: 698 SSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRGVAWVKWEEICKKKEEGGL 752
S++ ++ P+ I DQ + R F+W E G+ V WE I +K GL
Sbjct: 281 SAISMYVMQLNWLPQSICDQICTVVRHFVW--ENVNGIPLVAWETIQSRKSMVGL 439
>TC18198 weakly similar to UP|Q9XGZ5 (Q9XGZ5) T1N24.20 protein, partial
(14%)
Length = 592
Score = 42.0 bits (97), Expect = 4e-04
Identities = 16/49 (32%), Positives = 32/49 (64%)
Frame = -3
Query: 389 NASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVI 437
N + +TLIPK+ + + + +RPIS +YK+++K++ RL+ + +I
Sbjct: 158 NETLVTLIPKIPHAEAIQQFRPISCCSFIYKVISKIIVARLKPDMVNLI 12
>BP083121
Length = 527
Score = 33.5 bits (75), Expect(2) = 7e-04
Identities = 15/27 (55%), Positives = 18/27 (66%)
Frame = +3
Query: 542 PFLFLIVAEGLNGLMRQAVQQKLFTGY 568
P FLIVAEGLNGL++Q F G+
Sbjct: 312 PSFFLIVAEGLNGLLKQVATINKFKGF 392
Score = 26.6 bits (57), Expect(2) = 7e-04
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = +2
Query: 526 EEFNMGRGLRQGDPLAPFLF 545
+EF M + L Q DP+A FLF
Sbjct: 263 KEFKMWKSLHQVDPMAAFLF 322
>AV426437
Length = 303
Score = 36.2 bits (82), Expect = 0.021
Identities = 15/27 (55%), Positives = 18/27 (66%)
Frame = +1
Query: 730 EEGRGVAWVKWEEICKKKEEGGLGVKD 756
E R +AW+ WE ICK K GLGVK+
Sbjct: 214 ENQRKIAWINWETICKPKWLRGLGVKN 294
>BP056651
Length = 564
Score = 33.9 bits (76), Expect = 0.10
Identities = 17/35 (48%), Positives = 20/35 (56%)
Frame = +1
Query: 79 NTRGSGPMDNLRVDWTDSWFLKNG*LIGPIVHNMS 113
N G M+ V W DSW+L NG L+G IV N S
Sbjct: 460 NILGFDRMEKQEVVWIDSWYLINGILLGLIVLNTS 564
>AV417978
Length = 374
Score = 30.4 bits (67), Expect = 1.1
Identities = 22/77 (28%), Positives = 33/77 (42%), Gaps = 19/77 (24%)
Frame = -2
Query: 721 IQRKFLWGGEEGRGVA-------------WVKWEEICKKKEEGGLGVKDLESFNKALLV- 766
+QR FL E RG+ WV EE + +GGLGVK++E + + +
Sbjct: 262 LQRSFLGSVERERGLLLSSVVVDRG*GCWWVNEEE----ETQGGLGVKNVEKGERVVKLA 95
Query: 767 -----KWRWRLLREREW 778
W+W+ E W
Sbjct: 94 L**EGTWQWKRAIEEFW 44
>BP060568
Length = 286
Score = 29.6 bits (65), Expect = 1.9
Identities = 12/29 (41%), Positives = 19/29 (65%), Gaps = 1/29 (3%)
Frame = +1
Query: 4 IVRWRTRGEH-GRKSYSGGQPVTCKHGAW 31
+++WR++ +H GR S G P+ HGAW
Sbjct: 157 LIKWRSKKDHKGRISPLPGSPLGYTHGAW 243
>TC19636
Length = 529
Score = 28.1 bits (61), Expect = 5.6
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +2
Query: 603 ILRWFEAMSGLKVNFFKSKLAG 624
+LRWF ++GLK N + LAG
Sbjct: 56 LLRWFATVAGLKTNHRRHSLAG 121
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.341 0.150 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,017,280
Number of Sequences: 28460
Number of extensions: 204715
Number of successful extensions: 1732
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1702
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1727
length of query: 787
length of database: 4,897,600
effective HSP length: 98
effective length of query: 689
effective length of database: 2,108,520
effective search space: 1452770280
effective search space used: 1452770280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0263.7


