

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
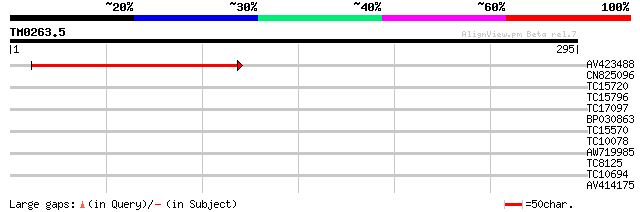
Query= TM0263.5
(295 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV423488 170 2e-43
CN825096 28 1.8
TC15720 similar to UP|O35328 (O35328) Proline-rich protein 9-1 (... 28 2.4
TC15796 similar to UP|Q948T1 (Q948T1) Two-pore calcium channel, ... 28 2.4
TC17097 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfat... 27 3.1
BP030863 27 4.1
TC15570 similar to GB|AAP21363.1|30102890|BT006555 At4g38290 {Ar... 27 4.1
TC10078 27 5.4
AW719985 27 5.4
TC8125 26 7.0
TC10694 similar to PIR|T05577|T05577 uncoupling protein homolog ... 26 9.1
AV414175 26 9.1
>AV423488
Length = 487
Score = 170 bits (431), Expect = 2e-43
Identities = 77/110 (70%), Positives = 91/110 (82%)
Frame = +3
Query: 12 NVGVTDFPEFSTNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYG 71
+V +FPEFST MTLG MTA N+ TPN+ED+TP+S+KT SPAW+T QNLVLIS WI G
Sbjct: 156 SVDAPEFPEFSTQMTLGGMTAANQFTPNQEDTTPKSKKTHSPAWSTAQNLVLISRWINRG 335
Query: 72 TSSVVGRNQKGETYWSQIAEYCHEHCSFDPPREGGACRNCFNYMSNQLNK 121
TSSVVG+NQKGET+W IAEYC+EHCSFDPPR+G ACRN +NYM+ L K
Sbjct: 336 TSSVVGKNQKGETFWRDIAEYCNEHCSFDPPRDGIACRNRWNYMNKILGK 485
>CN825096
Length = 730
Score = 28.1 bits (61), Expect = 1.8
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = +3
Query: 251 KISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQ 290
KI+ Q N+ +M L EH+DDR K+LL Q+ Q
Sbjct: 321 KIACAQAAKNKWVEMGKQLITRKVEHVDDRVKDLLLQVQQ 440
>TC15720 similar to UP|O35328 (O35328) Proline-rich protein 9-1 (Fragment),
partial (11%)
Length = 1115
Score = 27.7 bits (60), Expect = 2.4
Identities = 16/46 (34%), Positives = 19/46 (40%)
Frame = +3
Query: 168 LCNEPHYCSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGRE 213
LC S GG+ G GSS S S S C S I + + E
Sbjct: 33 LCGNDSGTSFTGGSSGGGSSSSSISSSSAGCFSGYITEVDEEVEEE 170
>TC15796 similar to UP|Q948T1 (Q948T1) Two-pore calcium channel, partial
(42%)
Length = 1524
Score = 27.7 bits (60), Expect = 2.4
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +3
Query: 155 NSHPFTLMKEWLALCNEPH 173
N PFT +KE+ +LC EPH
Sbjct: 60 NILPFTHLKEFESLCEEPH 116
>TC17097 similar to UP|Q9SE92 (Q9SE92) Adenosine-5'-phosphosulfate kinase
(Fragment) , partial (39%)
Length = 576
Score = 27.3 bits (59), Expect = 3.1
Identities = 13/31 (41%), Positives = 22/31 (70%), Gaps = 5/31 (16%)
Frame = -3
Query: 69 KYGTSSV----VGRNQKGETYWSQIAE-YCH 94
++GT++V +GRNQ G+TY + I + +CH
Sbjct: 127 QHGTANVPISSIGRNQTGDTY*TSIGK*FCH 35
>BP030863
Length = 453
Score = 26.9 bits (58), Expect = 4.1
Identities = 16/45 (35%), Positives = 21/45 (46%), Gaps = 7/45 (15%)
Frame = -3
Query: 179 GGNLGSGS-------SGSKRSHESDACGSNSIGSIPRPMGREAAK 216
G N GS S SGSK H + S GS+ +P R ++K
Sbjct: 349 GSNHGSDSPSQDAPASGSKHGHRRKSKNSEDGGSVRKPSSRRSSK 215
>TC15570 similar to GB|AAP21363.1|30102890|BT006555 At4g38290 {Arabidopsis
thaliana;}, partial (41%)
Length = 642
Score = 26.9 bits (58), Expect = 4.1
Identities = 19/73 (26%), Positives = 38/73 (52%), Gaps = 12/73 (16%)
Frame = +1
Query: 230 VVNNEWSE-YKQLRE-----------QELERLEKISSVQQEANRLEKMKMFLQLSFEEHL 277
+++ +W E Y+QL+ + ++ L+ +SSV+ + +E K +QLS EE
Sbjct: 106 LIDMQWEERYQQLQMLLRKLDQSDQVEYVQMLQSLSSVELSRHAVELEKRAIQLSLEEAK 285
Query: 278 DDRRKELLKQLSQ 290
+ +R +L L +
Sbjct: 286 ELQRVAVLNVLGK 324
>TC10078
Length = 500
Score = 26.6 bits (57), Expect = 5.4
Identities = 9/27 (33%), Positives = 14/27 (51%)
Frame = -2
Query: 96 HCSFDPPREGGACRNCFNYMSNQLNKW 122
HC + R G +C +CFN+ + W
Sbjct: 151 HC*YKLIRGGSSCCSCFNFTEEEEANW 71
>AW719985
Length = 359
Score = 26.6 bits (57), Expect = 5.4
Identities = 12/42 (28%), Positives = 19/42 (44%)
Frame = +2
Query: 177 QVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKK 218
+ G G+G S + H D CG PR RE+ +++
Sbjct: 143 RAGEGGGTGWSWEQPRHGGDGCGGGGSSCHPRSESRESLRRR 268
>TC8125
Length = 1373
Score = 26.2 bits (56), Expect = 7.0
Identities = 15/47 (31%), Positives = 20/47 (41%)
Frame = -1
Query: 28 GDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSS 74
G T N P S P S TQ+P+ + L GW++ SS
Sbjct: 812 GVSTNQNHHHPYRNLSRPLSHTTQTPSSTSRHRLQRQQGWLRQAFSS 672
>TC10694 similar to PIR|T05577|T05577 uncoupling protein homolog F22K18.230
- Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(43%)
Length = 604
Score = 25.8 bits (55), Expect = 9.1
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = -2
Query: 179 GGNLGSGSSGSKRSHESDACGSNS 202
GG+ G S G+ RS +SDA G S
Sbjct: 336 GGDAGEESGGAFRSDDSDADGDGS 265
>AV414175
Length = 226
Score = 25.8 bits (55), Expect = 9.1
Identities = 13/38 (34%), Positives = 18/38 (47%)
Frame = -2
Query: 188 GSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRD 225
G++ SHES +GS R GR ++ SRD
Sbjct: 162 GNETSHESKPITPKRVGSTGRRRGRRGLWPLTRRTSRD 49
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,619,376
Number of Sequences: 28460
Number of extensions: 86467
Number of successful extensions: 395
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 391
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 395
length of query: 295
length of database: 4,897,600
effective HSP length: 90
effective length of query: 205
effective length of database: 2,336,200
effective search space: 478921000
effective search space used: 478921000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0263.5


