

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0260.10
(751 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
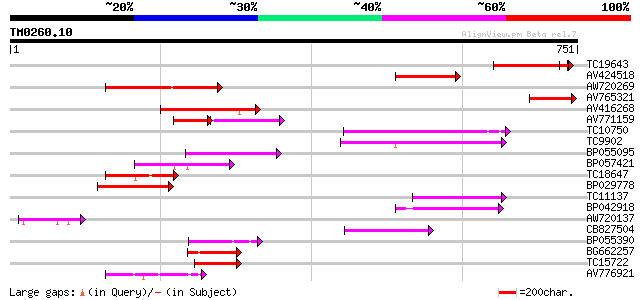
Score E
Sequences producing significant alignments: (bits) Value
TC19643 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like prote... 177 4e-45
AV424518 141 3e-34
AW720269 137 8e-33
AV765321 116 1e-26
AV416268 116 2e-26
AV771159 76 4e-21
TC10750 similar to UP|AAP80385 (AAP80385) ABC transporter, parti... 86 2e-17
TC9902 similar to UP|BAD07483 (BAD07483) PDR-type ABC transporte... 84 1e-16
BP055095 83 2e-16
BP057421 79 3e-15
TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partia... 78 5e-15
BP029778 72 2e-13
TC11137 similar to GB|CAD59563.1|27368811|OSA535041 PDR-like ABC... 61 7e-10
BP042918 61 7e-10
AW720137 60 1e-09
CB827504 55 4e-08
BP055390 49 3e-06
BG662257 49 3e-06
TC15722 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transpor... 49 4e-06
AV776921 48 5e-06
>TC19643 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like protein, partial
(12%)
Length = 550
Score = 177 bits (450), Expect = 4e-45
Identities = 87/104 (83%), Positives = 91/104 (86%)
Frame = +1
Query: 642 LVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATT 701
LVKYPYEAVLQNEFDDPVKCFVRGVQIFD+SPLRSVPYELKLKLLGSMSGTLGT+ITATT
Sbjct: 1 LVKYPYEAVLQNEFDDPVKCFVRGVQIFDHSPLRSVPYELKLKLLGSMSGTLGTDITATT 180
Query: 702 CLTTGADILQQNGVTELSKWNCLWITVAWGFFFRFLFYLALLVG 745
CLTTGADILQQNGVTELS WNCLWITV G + L + G
Sbjct: 181 CLTTGADILQQNGVTELS*WNCLWITVRLGILLQVFVLLGFVSG 312
Score = 42.4 bits (98), Expect = 3e-04
Identities = 18/19 (94%), Positives = 19/19 (99%)
Frame = +2
Query: 729 AWGFFFRFLFYLALLVGSK 747
AWGFFFRFLFYLALLVGS+
Sbjct: 263 AWGFFFRFLFYLALLVGSQ 319
>AV424518
Length = 261
Score = 141 bits (356), Expect = 3e-34
Identities = 66/86 (76%), Positives = 76/86 (87%)
Frame = +1
Query: 512 ADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGG 571
A+A+PVFLQERYIFMRETAYNAYRRSSY++SH++++LP L FLS AFA TFW+VGL GG
Sbjct: 4 AEAIPVFLQERYIFMRETAYNAYRRSSYVLSHSVIALPALIFLSFAFAATTFWSVGLAGG 183
Query: 572 ASGFLFYFLIIFASFWAGNSFVTFLS 597
SGF FYFL I ASFWAG+SFVTFLS
Sbjct: 184 TSGFFFYFLTIVASFWAGSSFVTFLS 261
>AW720269
Length = 524
Score = 137 bits (344), Expect = 8e-33
Identities = 73/158 (46%), Positives = 113/158 (71%), Gaps = 2/158 (1%)
Frame = +3
Query: 127 LLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKG-TLALNGEALESRL-LK 184
+L +S AR EI+AV+G SG+GKSTL+ +A R+ T+++N + S L+
Sbjct: 54 ILKSVSFVARSSEIVAVVGPSGTGKSTLLRIIAGRVKDRDFDPKTISINDHPMTSPAQLR 233
Query: 185 VISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVI 244
I +V Q+D L P+LTV+ETL F+A+FRL + ++ + ++ RV++L+ +LGL + + + +
Sbjct: 234 KICGFVAQEDNLLPLLTVKETLLFSAKFRL-KEMTPNDREMRVESLMQELGLFHVSDSFV 410
Query: 245 GDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGL 282
GDE +RG+SGGER+RVSIG+D+IH+P +L LDEPTSGL
Sbjct: 411 GDEENRGISGGERKRVSIGVDMIHNPPILLLDEPTSGL 524
>AV765321
Length = 455
Score = 116 bits (291), Expect = 1e-26
Identities = 51/62 (82%), Positives = 59/62 (94%)
Frame = -3
Query: 689 MSGTLGTNITATTCLTTGADILQQNGVTELSKWNCLWITVAWGFFFRFLFYLALLVGSKN 748
MS TLG NIT++TC+TTGADIL+Q G+T+LSKWNCLWIT+AWGFFFRFLFYLALL+GSKN
Sbjct: 453 MSKTLGVNITSSTCVTTGADILRQQGITQLSKWNCLWITIAWGFFFRFLFYLALLLGSKN 274
Query: 749 KR 750
KR
Sbjct: 273 KR 268
>AV416268
Length = 424
Score = 116 bits (290), Expect = 2e-26
Identities = 62/136 (45%), Positives = 90/136 (65%), Gaps = 3/136 (2%)
Frame = +3
Query: 200 LTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRR 259
LTV E + ++A +LP ++SKS+K+ R I ++GL++A T IG G +GVSGG++RR
Sbjct: 12 LTVGEAVYYSAHLQLPDSMSKSEKRERADFTIKEMGLQDAINTRIGGWGSKGVSGGQKRR 191
Query: 260 VSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSI---VIMSIHQPSYRIL 316
VSI I+I+ P LLFLDEPTSGLDS +++ V+ + + + I ++ SIHQPS I
Sbjct: 192 VSICIEILTHPRLLFLDEPTSGLDSAASYHVISRISSLNKKDGIQRTIVASIHQPSNEIF 371
Query: 317 GLLDRMIFLSRGQTVY 332
L + LS G+TVY
Sbjct: 372 QLFHSLCLLSSGKTVY 419
>AV771159
Length = 486
Score = 75.9 bits (185), Expect(2) = 4e-21
Identities = 39/98 (39%), Positives = 58/98 (58%)
Frame = -3
Query: 266 IIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFL 325
+IH P LL LDEPTSGLDST+A ++ + R+A G V+ +IHQPS R+ + D+++ L
Sbjct: 337 LIH-PSLLLLDEPTSGLDSTTALRILNTIPRLATGGRTVVTTIHQPSSRLYYMFDKVVLL 161
Query: 326 SRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALDL 363
S G +Y G + +F+ G + N + LDL
Sbjct: 160 SEGCPIYYGPASTALEYFSSVGFSTCVTVNPADLLLDL 47
Score = 43.1 bits (100), Expect(2) = 4e-21
Identities = 21/51 (41%), Positives = 32/51 (62%)
Frame = -2
Query: 218 LSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIH 268
L++ +K V+ +I +LGL ++IG RG+SG E+RRVSIG + H
Sbjct: 485 LTRDEKVQHVERVITELGLSGCRSSMIGGPLLRGISGAEKRRVSIGQETAH 333
>TC10750 similar to UP|AAP80385 (AAP80385) ABC transporter, partial (38%)
Length = 807
Score = 85.9 bits (211), Expect = 2e-17
Identities = 52/221 (23%), Positives = 109/221 (48%)
Frame = +1
Query: 443 FWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLGFFAF 502
F ++ TL+KRSF + R + +RL +V + T++ N+ + R +F
Sbjct: 142 FLMQSYTLTKRSFINMSRDFGYYWLRLVIYIVVTICIGTIYLNVGTGYNSILARGSCASF 321
Query: 503 AMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVIT 562
+ + P F+++ +F RE Y +++++S+ + + P L ++ I
Sbjct: 322 VFGFVTFMSIGGFPSFVEDMKVFQRERLNGHYGVTAFVVSNTVSATPFLVLITFLSGTIC 501
Query: 563 FWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSG 622
++ V L G S ++F+ L ++AS S + ++ +VP+ ++G I I F+L+SG
Sbjct: 502 YFMVRLHPGFSHYVFFVLCLYASVTVVESLMMAIASIVPNFLMGIIIGAGIQGIFMLVSG 681
Query: 623 FFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFV 663
+F IP +W + +S + + + A LQ ++ + ++ V
Sbjct: 682 YFRLPHDIPKP-VWRYPMSYISFHFWA-LQGQYQNDLRGLV 798
>TC9902 similar to UP|BAD07483 (BAD07483) PDR-type ABC transporter 1,
partial (19%)
Length = 1207
Score = 83.6 bits (205), Expect = 1e-16
Identities = 53/223 (23%), Positives = 91/223 (40%), Gaps = 4/223 (1%)
Frame = +1
Query: 439 FANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNSPKGVQERLG 498
F+ PF I+ + R P +R + T+FW+L K Q+ L
Sbjct: 94 FSQPFLIQCQACLWKQRWSYWRNPPYTAVRFFFTTFIAVMFGTIFWDLGGKHKRRQDLLN 273
Query: 499 FFAFAMSTTFY----TTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFL 554
S + ++ PV ER +F RE A Y Y + LV LP + F
Sbjct: 274 AVGSMYSAVLFLGVQNSSSVQPVVSVERTVFYREKAAGMYSALPYAFAQILVELPYIFFQ 453
Query: 555 SLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAIL 614
++ + VI + +G D A F +Y ++ + + V P+ + + A
Sbjct: 454 AVTYGVIVYAMIGFDWTAEKFFWYLFFMYFTLLYFTFYGMMGVAVTPNHHVASIVAAAFY 633
Query: 615 AYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDD 657
A + L SGF + R IP +W W+++ V + ++ ++F D
Sbjct: 634 AIWNLFSGFVVPRPSIPVWWRWYYWACPVAWTIYGLIASQFGD 762
>BP055095
Length = 538
Score = 82.8 bits (203), Expect = 2e-16
Identities = 44/127 (34%), Positives = 72/127 (56%), Gaps = 1/127 (0%)
Frame = +3
Query: 234 LGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKV 293
LGL A +IGDE RG+SGG+++RV+ G ++ LF+DE ++GLDS++ F + K
Sbjct: 48 LGLDICADVMIGDEMRRGISGGQKKRVTTGEMLVGPAKALFMDEISTGLDSSTTFQICKF 227
Query: 294 LQRIAQSGSI-VIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPD 352
++++ + +++S+ QP+ L D +I LS GQ VY G + FF G P+
Sbjct: 228 MRQMVHIMDVTMVISLLQPAPETFELFDDIILLSEGQIVYQGPRENVLEFFEYMGFKCPE 407
Query: 353 SDNRTEF 359
+F
Sbjct: 408 RKGAADF 428
>BP057421
Length = 547
Score = 79.0 bits (193), Expect = 3e-15
Identities = 54/163 (33%), Positives = 86/163 (52%), Gaps = 31/163 (19%)
Frame = +3
Query: 166 RLKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRT-------L 218
R+ G + NG L + + +AY+ Q+D+ +TV+ETL F+A + T L
Sbjct: 42 RVTGEITYNGHKLNEFVPRKTAAYISQNDVHVGEMTVKETLDFSARCQGVGTRYDXLSEL 221
Query: 219 SKSKKKARV--QALIDQ----------------------LGLRNAAKTVIGDEGHRGVSG 254
+ +K+A + +A +D LGL T++GDE HRGVSG
Sbjct: 222 GRREKEAGIFPEAELDLFMKATALKGTESSLITDYTLKILGLDICKDTIVGDEMHRGVSG 401
Query: 255 GERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRI 297
G+++RV+ G I+ LF+DE ++GLDS++ F +VK LQ+I
Sbjct: 402 GQKKRVTTGEMIVGPTKTLFMDEISTGLDSSTTFQIVKCLQQI 530
>TC18647 similar to UP|Q9ASR9 (Q9ASR9) At2g01320/F10A8.20, partial (17%)
Length = 430
Score = 78.2 bits (191), Expect = 5e-15
Identities = 43/100 (43%), Positives = 61/100 (61%), Gaps = 3/100 (3%)
Frame = +1
Query: 127 LLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKG---RLKGTLALNGEALESRLL 183
LL ++SGEA+ G ++A++G SGSGK+TL++ LA ++A L G L NG+
Sbjct: 106 LLKNVSGEAKPGRLLAIMGPSGSGKTTLLNVLAGQLAASPRLHLSGLLEFNGKPGSKNAY 285
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKK 223
K AYV Q+DL F LTV ETL+ A E +LP S ++
Sbjct: 286 KF--AYVRQEDLFFSQLTVRETLSLATELQLPNISSAEER 399
>BP029778
Length = 438
Score = 72.4 bits (176), Expect = 2e-13
Identities = 36/100 (36%), Positives = 61/100 (61%)
Frame = -2
Query: 117 EESVFSRTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGE 176
++ + LL+++SG G + A++G+SG+GK+TL+D LA R G ++G + ++G
Sbjct: 308 KQGIIETKLKLLSNVSGVFAPGVLTALMGSSGAGKTTLMDVLAGRKTGGYIEGDIKISGY 129
Query: 177 ALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPR 216
++ YV Q+D+ P +TVEE+L F+A RLPR
Sbjct: 128 PKVQHTFARVAGYVEQNDIHSPQVTVEESLLFSALLRLPR 9
>TC11137 similar to GB|CAD59563.1|27368811|OSA535041 PDR-like ABC
transporter {Oryza sativa (japonica cultivar-group);},
partial (15%)
Length = 786
Score = 60.8 bits (146), Expect = 7e-10
Identities = 29/124 (23%), Positives = 56/124 (44%)
Frame = +1
Query: 534 YRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFV 593
Y Y I+ +P + + +++I + V D F +YF + F SF +
Sbjct: 253 YAPLPYAIAQVFTEIPYVFAQTTFYSLIVYAMVSFDWKVDKFFWYFFVSFFSFLYFTYYG 432
Query: 594 TFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQN 653
+ P+ + A F L SGFFI R +IP +W+W++++ V + ++ +
Sbjct: 433 MMTVSITPNHQVASIFAAAFYGLFNLFSGFFIPRPKIPGWWVWYYWICPVAWTVYGLIVS 612
Query: 654 EFDD 657
++ D
Sbjct: 613 QYRD 624
>BP042918
Length = 412
Score = 60.8 bits (146), Expect = 7e-10
Identities = 37/144 (25%), Positives = 67/144 (45%), Gaps = 1/144 (0%)
Frame = -3
Query: 511 TADALPVFLQER-YIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLD 569
T LPVF + R ++F + +Y + + L+ +P F SL + IT++ G
Sbjct: 410 TIQRLPVFYKHRDHLF--------HPAWTYTVPNFLLRIPISIFESLVWVAITYYTTGFA 255
Query: 570 GGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDR 629
AS F L++F +SGV +++ T +L LL GF + +
Sbjct: 254 PEASRFFKQLLVVFLIQQMAAGMFRLISGVCRTMIIANTGGALMLLLVFLLGGFLLPKRA 75
Query: 630 IPSYWIWFHYLSLVKYPYEAVLQN 653
IP +W+W +++S + Y + ++ N
Sbjct: 74 IPDWWVWAYWISPLSYAFNSLTVN 3
>AW720137
Length = 563
Score = 60.1 bits (144), Expect = 1e-09
Identities = 39/102 (38%), Positives = 56/102 (54%), Gaps = 13/102 (12%)
Frame = +3
Query: 12 DDATSY----LDLMELSDLTRRPASGDLPTLGQLLKHVGDARKEAAGDGSETPL------ 61
DD+ ++ ++L+ L R + PTLG+LLK V DA+ + + T +
Sbjct: 246 DDSLTFFNQSMELVNLPPTRTRTRAHISPTLGELLKRVEDAQHQHHHHNTTTTIAESDEK 425
Query: 62 HHALDVVGMEPRSLP---FVLSFSNLTYSVKVKRKLSFSSIF 100
HH LD+ SLP F LSF+NLTYSVK++RK+S S F
Sbjct: 426 HHVLDLSSSSSSSLPSHPFYLSFTNLTYSVKLRRKMSLSRCF 551
>CB827504
Length = 536
Score = 55.1 bits (131), Expect = 4e-08
Identities = 38/121 (31%), Positives = 61/121 (50%), Gaps = 3/121 (2%)
Frame = +2
Query: 444 WIEMV-TLSKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNS-PKGVQERLG-FF 500
W+E L R F + RR +R+ V+ T IL ++W D+S PKG+Q++ G F
Sbjct: 176 WLEQFFILFSRGFKE-RRHDYFSWLRITQVLSTAIILGLLWWQSDSSNPKGLQDQAGLLF 352
Query: 501 AFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAV 560
A+ F+ A+ F QER + +E A + YR S+Y ++ LP L + F +
Sbjct: 353 FIAVFWGFFPVFTAIFTFPQERAMLAKERASDMYRLSAYFLARTTSDLPLDLVLPVLFLL 532
Query: 561 I 561
+
Sbjct: 533 V 535
>BP055390
Length = 488
Score = 48.9 bits (115), Expect = 3e-06
Identities = 34/100 (34%), Positives = 55/100 (55%), Gaps = 2/100 (2%)
Frame = +3
Query: 238 NAAKTVIGDEGHRGV--SGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQ 295
N ++ G RGV SGG+++R++I I+ +P +L LDE TS LD+ S +V + L+
Sbjct: 183 N*CSNLLTQVGQRGVQLSGGQKQRIAIARAILKNPPILLLDEATSALDTESEKLVQEALE 362
Query: 296 RIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGS 335
A G VI+ H+ S + D + + GQ V +G+
Sbjct: 363 -TAMHGRTVILIAHRLSTVVNA--DVIAVVENGQVVETGT 473
>BG662257
Length = 336
Score = 48.9 bits (115), Expect = 3e-06
Identities = 26/72 (36%), Positives = 44/72 (61%)
Frame = +1
Query: 236 LRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQ 295
L+ TV+GD G + +SGG+++RV+I I+ P +L LDE TS LD+ S +V L
Sbjct: 94 LQQGYDTVVGDRGIQ-LSGGQKQRVAIARAIVKSPKILLLDEATSALDAESEKVVQDALD 270
Query: 296 RIAQSGSIVIMS 307
R+ + ++++
Sbjct: 271 RVRVDRTTIVVA 306
>TC15722 similar to UP|Q9FMU0 (Q9FMU0) Similarity to ABC transporter,
partial (37%)
Length = 624
Score = 48.5 bits (114), Expect = 4e-06
Identities = 22/62 (35%), Positives = 42/62 (67%)
Frame = +3
Query: 246 DEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVI 305
D+ + +SGG +RR+++ I ++ P LL LDEP +GLD + VVK+L+ + + ++++
Sbjct: 51 DKNPQSLSGGYKRRLALAIQLVQVPDLLILDEPLAGLDWKARADVVKLLKHLKKELTVLV 230
Query: 306 MS 307
+S
Sbjct: 231 VS 236
>AV776921
Length = 435
Score = 48.1 bits (113), Expect = 5e-06
Identities = 40/137 (29%), Positives = 76/137 (55%), Gaps = 3/137 (2%)
Frame = +2
Query: 127 LLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNG---EALESRLL 183
+LN + + G+I+A++G SGSGKST+I +L R + L G + L+G L+ + L
Sbjct: 41 ILNKLCLDIPSGKIVALVGGSGSGKSTVI-SLIERFYE-PLSGDILLDGNDIRDLDLKWL 214
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
+ V Q+ LF +++E + + + L ++ K + Q+ I+ L R +T
Sbjct: 215 RQQIGLVNQEPALF-ATSIKENILYGKDNATLEELKRAVKLSDAQSFINNLPER--LETQ 385
Query: 244 IGDEGHRGVSGGERRRV 260
+G+ G + +SGG+++R+
Sbjct: 386 VGERGIQ-LSGGQKQRI 433
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,335,743
Number of Sequences: 28460
Number of extensions: 149641
Number of successful extensions: 1077
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 1062
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1071
length of query: 751
length of database: 4,897,600
effective HSP length: 97
effective length of query: 654
effective length of database: 2,136,980
effective search space: 1397584920
effective search space used: 1397584920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0260.10


