

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258b.5
(360 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
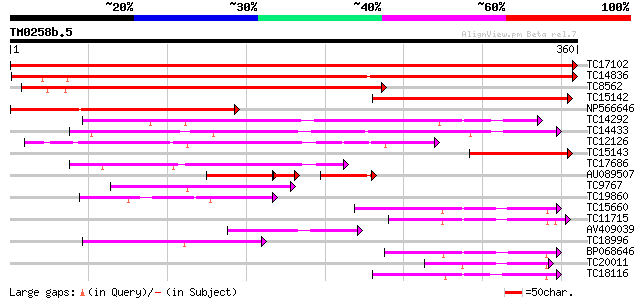
Score E
Sequences producing significant alignments: (bits) Value
TC17102 UP|CAE45588 (CAE45588) Papain-like cysteine proteinase-l... 708 0.0
TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precur... 530 e-151
TC8562 similar to UP|O24322 (O24322) Cysteine proteinase precurs... 318 1e-87
TC15142 similar to UP|Q945R8 (Q945R8) Cysteine proteinase (Fragm... 229 4e-61
NP566646 hypothetical protein [Lotus japonicus] 226 4e-60
TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precur... 196 4e-51
TC14433 similar to UP|Q41057 (Q41057) Cysteine protease , complete 194 3e-50
TC12126 similar to PIR|S49166|S49166 cysteine proteinase precur... 135 8e-33
TC15143 similar to UP|CYSP_PEA (P25804) Cysteine proteinase 15A ... 122 9e-29
TC17686 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Be... 115 9e-27
AU089507 58 1e-22
TC9767 similar to UP|Q7X750 (Q7X750) Cysteine proteinase , part... 89 9e-19
TC19860 weakly similar to UP|Q41064 (Q41064) Thiolprotease , par... 85 2e-17
TC15660 weakly similar to UP|Q38886 (Q38886) SAG12 protein, part... 83 6e-17
TC11715 weakly similar to UP|Q38886 (Q38886) SAG12 protein, part... 75 1e-14
AV409039 75 2e-14
TC18996 similar to UP|O24323 (O24323) Cysteine proteinase precur... 73 6e-14
BP068646 71 2e-13
TC20011 similar to GB|CAA40073.1|1345573|PVEPC1 endopeptidase (E... 70 7e-13
TC18116 similar to UP|AAP32193 (AAP32193) Cysteine protease 14, ... 66 1e-11
>TC17102 UP|CAE45588 (CAE45588) Papain-like cysteine proteinase-like protein
1, complete
Length = 1086
Score = 708 bits (1828), Expect = 0.0
Identities = 340/361 (94%), Positives = 350/361 (96%), Gaps = 1/361 (0%)
Frame = +1
Query: 1 MVDHRILFLMFVFFLFFSVVSSDGGVDPLIRQVVDGEGLGAEHHFLEFKRRFGKVYVSEE 60
MVDHRILFL+FVFFLFFS VSS GVDP+I QVVD EGLGAEHHFLEFKRRFGKVY +EE
Sbjct: 1 MVDHRILFLLFVFFLFFSNVSSSDGVDPMICQVVDDEGLGAEHHFLEFKRRFGKVYATEE 180
Query: 61 EHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGVGLPSDADSA 120
EHGYRFNVFKSNMHRARRHQLLDPSAVHGVT+FSDLTPMEF+HSVLGLRGVGLPSDADSA
Sbjct: 181 EHGYRFNVFKSNMHRARRHQLLDPSAVHGVTQFSDLTPMEFQHSVLGLRGVGLPSDADSA 360
Query: 121 PILRTDNLPKDFDWREHGAVTPVKNQGSCGACWSFSATGALEGAHFLSTGKLVSLSEQQL 180
PIL TDNLPKDFDWREHGAVTPVKNQGSCG+CWSFSATGALEGAHFLSTG+LVSLSEQQL
Sbjct: 361 PILPTDNLPKDFDWREHGAVTPVKNQGSCGSCWSFSATGALEGAHFLSTGELVSLSEQQL 540
Query: 181 VDCDHE-CDPEEAGSCDSGCKGGLMNSAFEYILNNGGVMREEDYPYSGTAGGTCKFDQTK 239
VDCDH+ CDPEEAGSCDSGC GGLMNSAFEYILNNGGVMREEDYPYSGT GGTCKFD+ K
Sbjct: 541 VDCDHQQCDPEEAGSCDSGCNGGLMNSAFEYILNNGGVMREEDYPYSGTNGGTCKFDKAK 720
Query: 240 IAASVANFSVVSRDEDQIAANLVKNGPLAVAINAVYMQTYVGGVSCPYVCSKKLNHGVLL 299
IAASVANFSVVSRDEDQIAANLVKNGPLAVAINAVYMQTYVGGVSCPYVCSKKLNHGVLL
Sbjct: 721 IAASVANFSVVSRDEDQIAANLVKNGPLAVAINAVYMQTYVGGVSCPYVCSKKLNHGVLL 900
Query: 300 VGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGRNVCGVDSMVSTVAALHTTG 359
VGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGRN+CGVDSMVSTVAALHTTG
Sbjct: 901 VGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGRNICGVDSMVSTVAALHTTG 1080
Query: 360 N 360
N
Sbjct: 1081N 1083
>TC14836 similar to UP|O24322 (O24322) Cysteine proteinase precursor ,
partial (92%)
Length = 1416
Score = 530 bits (1366), Expect = e-151
Identities = 258/368 (70%), Positives = 295/368 (80%), Gaps = 9/368 (2%)
Frame = +2
Query: 2 VDHRILFLMFVFFLFFSV-----VSSDGGVDPLIRQVVD----GEGLGAEHHFLEFKRRF 52
+DHR LF + +F + + ++ D LIRQVVD L AE HF FK +F
Sbjct: 29 MDHRKLFAILLFAAVATAAAANRIDAEDHDDALIRQVVDHVHDDHLLNAERHFTTFKAKF 208
Query: 53 GKVYVSEEEHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGVG 112
GK Y + EEH YRF VFKSN+ RA+ H LDPSA HGVT+FSDLTP EFR LG++ +
Sbjct: 209 GKSYATREEHDYRFGVFKSNLRRAKVHAKLDPSAEHGVTKFSDLTPAEFRRQFLGVKALR 388
Query: 113 LPSDADSAPILRTDNLPKDFDWREHGAVTPVKNQGSCGACWSFSATGALEGAHFLSTGKL 172
LP+DA APIL T +LP DFDWR+ GAVTPVK+QG+CG+CW+FSATGALEG+H+L+TGKL
Sbjct: 389 LPADAQKAPILPTRDLPDDFDWRDKGAVTPVKDQGACGSCWAFSATGALEGSHYLTTGKL 568
Query: 173 VSLSEQQLVDCDHECDPEEAGSCDSGCKGGLMNSAFEYILNNGGVMREEDYPYSGTAGGT 232
VS+SEQQLVDCDH CDPEE SCDSGC GGLMN+AFEYIL +GGVMRE+DYPY+G GT
Sbjct: 569 VSISEQQLVDCDHVCDPEEYNSCDSGCNGGLMNNAFEYILQSGGVMREKDYPYTG-RDGT 745
Query: 233 CKFDQTKIAASVANFSVVSRDEDQIAANLVKNGPLAVAINAVYMQTYVGGVSCPYVCSKK 292
CKFD++KIAASVANFSV S DEDQIAANLV NGPLAVAINAVYMQTYVGGVSCPY+C K
Sbjct: 746 CKFDKSKIAASVANFSVGSLDEDQIAANLVMNGPLAVAINAVYMQTYVGGVSCPYICGKN 925
Query: 293 LNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGRNVCGVDSMVSTV 352
L+HGVLLVGYG YAPIR K K YWIIKNSWGE+WGENGYYKICRGRNVCGVDSMVSTV
Sbjct: 926 LDHGVLLVGYGKAGYAPIRFKNKAYWIIKNSWGESWGENGYYKICRGRNVCGVDSMVSTV 1105
Query: 353 AALHTTGN 360
AA HT+ +
Sbjct: 1106AAAHTSSS 1129
>TC8562 similar to UP|O24322 (O24322) Cysteine proteinase precursor ,
partial (57%)
Length = 776
Score = 318 bits (814), Expect = 1e-87
Identities = 157/239 (65%), Positives = 180/239 (74%), Gaps = 7/239 (2%)
Frame = +2
Query: 8 FLMFVFFLFFSVVSS----DGGVDPLIRQVV---DGEGLGAEHHFLEFKRRFGKVYVSEE 60
FL+ F + V++ D D LIRQVV + L AEHHF FK RFGK Y +EE
Sbjct: 59 FLLLALIFFSAAVATAAVIDDAEDLLIRQVVPEVEDHLLNAEHHFSTFKARFGKTYATEE 238
Query: 61 EHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGVGLPSDADSA 120
EH YRF VFK N+ A+ HQ LDPSAVHGVTRFSDLTP EFR LGL+ DA A
Sbjct: 239 EHNYRFGVFKKNLILAKSHQKLDPSAVHGVTRFSDLTPSEFRRQFLGLKAPRHLKDAQKA 418
Query: 121 PILRTDNLPKDFDWREHGAVTPVKNQGSCGACWSFSATGALEGAHFLSTGKLVSLSEQQL 180
PIL T++LP DFDWRE GAVT VKNQGSCG+CWSFSATGALEGAH+L+TG+LVSLSEQQL
Sbjct: 419 PILPTNDLPTDFDWREKGAVTEVKNQGSCGSCWSFSATGALEGAHYLATGELVSLSEQQL 598
Query: 181 VDCDHECDPEEAGSCDSGCKGGLMNSAFEYILNNGGVMREEDYPYSGTAGGTCKFDQTK 239
VDCDHECDPE++ SCDSGC GGLM +AFEY L GGVMRE+DYPY+G GTCKF+++K
Sbjct: 599 VDCDHECDPEDSRSCDSGCNGGLMTNAFEYTLKAGGVMREKDYPYTGRDRGTCKFEKSK 775
>TC15142 similar to UP|Q945R8 (Q945R8) Cysteine proteinase (Fragment),
partial (36%)
Length = 646
Score = 229 bits (585), Expect = 4e-61
Identities = 103/127 (81%), Positives = 119/127 (93%)
Frame = +1
Query: 231 GTCKFDQTKIAASVANFSVVSRDEDQIAANLVKNGPLAVAINAVYMQTYVGGVSCPYVCS 290
GTCKF+++KIAAS +NFSVVS DEDQIAANLVK+GPLA+ INAV+MQTY+GGVSCPY+C
Sbjct: 1 GTCKFEKSKIAASASNFSVVSVDEDQIAANLVKHGPLAIGINAVFMQTYIGGVSCPYICG 180
Query: 291 KKLNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGRNVCGVDSMVS 350
K L+HGVLLVGYG+ +Y+PIR+K+KPYWIIKNSWGE WGENGYYKICRGRNVCGVDSMVS
Sbjct: 181 KHLDHGVLLVGYGAGAYSPIRLKEKPYWIIKNSWGETWGENGYYKICRGRNVCGVDSMVS 360
Query: 351 TVAALHT 357
TVAA+ T
Sbjct: 361 TVAAIQT 381
>NP566646 hypothetical protein [Lotus japonicus]
Length = 996
Score = 226 bits (577), Expect = 4e-60
Identities = 118/146 (80%), Positives = 123/146 (83%)
Frame = -1
Query: 1 MVDHRILFLMFVFFLFFSVVSSDGGVDPLIRQVVDGEGLGAEHHFLEFKRRFGKVYVSEE 60
MVDHRILFL+FVFFLFFS VSS GVDP+I QVVD EGLG L + V EE
Sbjct: 489 MVDHRILFLLFVFFLFFSNVSSSDGVDPMICQVVDDEGLGGRTP-LSGV*ASVREGVREE 313
Query: 61 EHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGVGLPSDADSA 120
EHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGVGLPSDADSA
Sbjct: 312 EHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGVGLPSDADSA 133
Query: 121 PILRTDNLPKDFDWREHGAVTPVKNQ 146
PIL TDNLPKDFDWREHGAVT ++Q
Sbjct: 132 PILPTDNLPKDFDWREHGAVTRDEDQ 55
Score = 39.3 bits (90), Expect = 0.001
Identities = 18/19 (94%), Positives = 19/19 (99%)
Frame = -1
Query: 250 VSRDEDQIAANLVKNGPLA 268
V+RDEDQIAANLVKNGPLA
Sbjct: 75 VTRDEDQIAANLVKNGPLA 19
>TC14292 similar to UP|O24323 (O24323) Cysteine proteinase precursor ,
partial (95%)
Length = 1792
Score = 196 bits (499), Expect = 4e-51
Identities = 116/302 (38%), Positives = 164/302 (53%), Gaps = 10/302 (3%)
Frame = +3
Query: 47 EFKRRFGKVYVSEEEHGYRFNVFKSNMHRARRHQLLDPSAVH--GVTRFSDLTPMEFRHS 104
E+ + GKVY + E RF +FK N+ H D + + G+ RF+DLT E+R
Sbjct: 186 EWLVKHGKVYNALGEKEKRFEIFKDNLKFIDEHNNADLNRSYKLGLNRFADLTNEEYRSK 365
Query: 105 VLGLRG------VGLPSDADSAPILRTDNLPKDFDWREHGAVTPVKNQGSCGACWSFSAT 158
G R L + +D D LP+ DWR+ GA+ VK+QGSCG+CW+FSA
Sbjct: 366 YFGTRVDPNRRMAKLRTKSDRYAPRVGDKLPESVDWRKEGALVGVKDQGSCGSCWAFSAV 545
Query: 159 GALEGAHFLSTGKLVSLSEQQLVDCDHECDPEEAGSCDSGCKGGLMNSAFEYILNNGGVM 218
A+E + + TG LVSLS Q+LVDCD S + GC GGLM+ AF++I+NNGG+
Sbjct: 546 TAVESINKIVTGDLVSLSVQELVDCDR--------SYNEGCNGGLMDYAFDFIINNGGID 701
Query: 219 REEDYPYSGTAGGTCKFDQTKIAASVANFSVVSRDEDQIAANLVKNGPLAVAI--NAVYM 276
EEDYPY G G ++ + S+ ++ V ++ V N P++VAI
Sbjct: 702 SEEDYPYKGVDGRCDQYRKNAKVVSIDDYEDVPTYDEIALKKAVANQPISVAIEGGGREF 881
Query: 277 QTYVGGVSCPYVCSKKLNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKI 336
Q Y G+ C L+HGV+ VGYG+E+ YWI++NSWG +WGE GY ++
Sbjct: 882 QLYDSGIFTGR-CGTALDHGVVAVGYGTEN-------GLDYWIVRNSWGASWGEGGYIRL 1037
Query: 337 CR 338
R
Sbjct: 1038ER 1043
>TC14433 similar to UP|Q41057 (Q41057) Cysteine protease , complete
Length = 1392
Score = 194 bits (492), Expect = 3e-50
Identities = 124/326 (38%), Positives = 173/326 (53%), Gaps = 14/326 (4%)
Frame = +1
Query: 39 LGAEHHFLEFKR---RFGKVYVSEEEHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSD 95
+G H L F R R+GK Y S EE +RF +F ++ + S G+ F+D
Sbjct: 205 IGQTRHALSFARFATRYGKRYDSVEEIQHRFRIFSESLELIKSTNKKRLSYKLGLNHFAD 384
Query: 96 LTPMEFRHSVLGLRGVGLPSDADSAPILRTDNL-----PKDFDWREHGAVTPVKNQGSCG 150
L+ EFR LG + SA ++ L P + DWR+ V+ VK+Q CG
Sbjct: 385 LSWDEFRTQKLGA------AQNCSATLIGNHKLTDAVLPAEKDWRKESIVSEVKDQAHCG 546
Query: 151 ACWSFSATGALEGAHFLSTGKLVSLSEQQLVDCDHECDPEEAGSCDS-GCKGGLMNSAFE 209
+CW+FS TGALE A+ + GK +SLSEQQLVDC AG+ ++ GC GGL + AFE
Sbjct: 547 SCWTFSTTGALEAAYAQAHGKNISLSEQQLVDC--------AGAFNNFGCNGGLPSQAFE 702
Query: 210 YILNNGGVMREEDYPYSGTAGGTCKFDQTKIAASVA-NFSVVSRDEDQIAANLVKNGPLA 268
YI NGG+ E++YPY+ CKF +A V + ++ ED++ + P++
Sbjct: 703 YIKYNGGIALEKEYPYT-AKDEACKFTAENVAVRVLDSVNITLGAEDELKHAVAFARPVS 879
Query: 269 VAINAV-YMQTYVGGVSCPYVCSK---KLNHGVLLVGYGSESYAPIRMKQKPYWIIKNSW 324
VA V + Y GV C +NH VL VGYG E+ PYWIIKNSW
Sbjct: 880 VAFQVVDGFRLYKEGVYTSDTCGNTPMDVNHAVLAVGYGVEN-------NVPYWIIKNSW 1038
Query: 325 GENWGENGYYKICRGRNVCGVDSMVS 350
G WG++GY+K+ G+N+CGV + S
Sbjct: 1039GSTWGDHGYFKMELGKNMCGVATCAS 1116
>TC12126 similar to PIR|S49166|S49166 cysteine proteinase precursor -
spring vetch {Vicia sativa;} , partial (71%)
Length = 839
Score = 135 bits (341), Expect = 8e-33
Identities = 97/273 (35%), Positives = 133/273 (48%), Gaps = 9/273 (3%)
Frame = +1
Query: 10 MFVFFLFFSVVSSDGGVDPLIRQVVDGEGLGAEHHFLEFKRRFGKVYVSEEEHGYRFNVF 69
+F LF SV S D + + EGL E R V S +E RFNVF
Sbjct: 43 LFSLALFLSVAES---FDYHEKDLASEEGLW---DLYERWRSHHTVSRSLDEKHKRFNVF 204
Query: 70 KSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGV-------GLPSDADSAPI 122
K+N+ LD + +F+DLT EF+ S+ + G+P S
Sbjct: 205 KANVMHVHETNKLDKPYKLKLNKFADLTNYEFK-SIYASSKIDHHRMFHGIPRGNGSFMY 381
Query: 123 LRTDNLPKDFDWREHGAVTPVKNQGSCGACWSFSATGALEGAHFLSTGKLVSLSEQQLVD 182
D++P DWR GAVTPVK+QG CG+CW+FSA A+EG + + T LVSLSEQ+LVD
Sbjct: 382 ENVDSVPISKDWRTSGAVTPVKDQGQCGSCWAFSAIAAVEGINQIRTHHLVSLSEQELVD 561
Query: 183 CDHECDPEEAGSCDSGCKGGLMNSAFEYILNNGGVMREEDYPYSGTAGGTCKFDQ--TKI 240
CD E + GC GG M A E+I G+ E YPY+ T GTC + +
Sbjct: 562 CDVE--------VNEGCNGGFMAKALEFI-KEKGITSESIYPYTAT-DGTCNTQKMVNEP 711
Query: 241 AASVANFSVVSRDEDQIAANLVKNGPLAVAINA 273
A S+ + +V + + N P++V I+A
Sbjct: 712 AVSIDGYEIVPENNEVALLKAAANQPISVGIDA 810
>TC15143 similar to UP|CYSP_PEA (P25804) Cysteine proteinase 15A precursor
(Turgor-responsive protein 15A) , partial (18%)
Length = 477
Score = 122 bits (306), Expect = 9e-29
Identities = 51/65 (78%), Positives = 62/65 (94%)
Frame = +1
Query: 293 LNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGRNVCGVDSMVSTV 352
L+HGVLLVGYG+ +Y+P+R+K+KPYWIIKNSWGE WGENGYY+ICRGR+VCGVDS+VSTV
Sbjct: 1 LDHGVLLVGYGAGAYSPLRLKEKPYWIIKNSWGETWGENGYYQICRGRHVCGVDSLVSTV 180
Query: 353 AALHT 357
AA+ T
Sbjct: 181 AAIQT 195
>TC17686 similar to UP|CYSP_VIGMU (P12412) Vignain precursor (Bean
endopeptidase) (Cysteine proteinase)
(Sulfhydryl-endopeptidase) (SH-EP) , partial (54%)
Length = 624
Score = 115 bits (289), Expect = 9e-27
Identities = 71/190 (37%), Positives = 100/190 (52%), Gaps = 13/190 (6%)
Frame = +3
Query: 39 LGAEHHFLEFKRRFGKVYV---SEEEHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSD 95
L +E + R+ V+ S +E RFNVFK+N+ LD + +F+D
Sbjct: 87 LESEESLWDLYERWRSVHTVSRSLDEKRKRFNVFKANVMHVHDTNKLDMPYKLELNKFAD 266
Query: 96 LTPMEFR----------HSVLGLRGVGLPSDADSAPILRTDNLPKDFDWREHGAVTPVKN 145
+T EFR H +L G P + D++P DWR+ GAVT +K+
Sbjct: 267 MTNHEFRSIYASSKIDHHKMLR----GTPHGKGTFMYENVDSVPPSIDWRKKGAVTAIKD 434
Query: 146 QGSCGACWSFSATGALEGAHFLSTGKLVSLSEQQLVDCDHECDPEEAGSCDSGCKGGLMN 205
Q C +CW+FSA A+EG + + T KLVSLSEQ+LVDCD E ++GC GG MN
Sbjct: 435 QRDCSSCWAFSAVAAVEGINQIKTQKLVSLSEQELVDCDIE--------HNNGCNGGYMN 590
Query: 206 SAFEYILNNG 215
+AF++I NG
Sbjct: 591 NAFDFIKQNG 620
>AU089507
Length = 379
Score = 58.2 bits (139), Expect(3) = 1e-22
Identities = 23/45 (51%), Positives = 30/45 (66%)
Frame = +3
Query: 126 DNLPKDFDWREHGAVTPVKNQGSCGACWSFSATGALEGAHFLSTG 170
D LP DWR GAV P+K+QGSCG+CW+FS +E + + TG
Sbjct: 33 DRLPAHVDWRLQGAVAPIKDQGSCGSCWAFSTIATVEAVNKIVTG 167
Score = 53.1 bits (126), Expect(3) = 1e-22
Identities = 23/36 (63%), Positives = 27/36 (74%)
Frame = +2
Query: 198 GCKGGLMNSAFEYILNNGGVMREEDYPYSGTAGGTC 233
GC GGLM+ AFE+I+ NGG+ EEDYPY G GTC
Sbjct: 227 GCNGGLMDYAFEFIIGNGGIDTEEDYPYKG-VDGTC 331
Score = 31.6 bits (70), Expect(3) = 1e-22
Identities = 14/17 (82%), Positives = 15/17 (87%)
Frame = +1
Query: 168 STGKLVSLSEQQLVDCD 184
S GKLV LSEQ+LVDCD
Sbjct: 160 SQGKLVPLSEQELVDCD 210
>TC9767 similar to UP|Q7X750 (Q7X750) Cysteine proteinase , partial (50%)
Length = 564
Score = 89.4 bits (220), Expect = 9e-19
Identities = 49/123 (39%), Positives = 69/123 (55%), Gaps = 6/123 (4%)
Frame = +3
Query: 65 RFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVLGLRGV------GLPSDAD 118
RFNVFK+N+ +D + +F+D+T EFR + G + G
Sbjct: 195 RFNVFKANVMHVHNTNKMDKPYKLKLNKFADMTNHEFRSAYAGSKVNHHRMFRGATRGNG 374
Query: 119 SAPILRTDNLPKDFDWREHGAVTPVKNQGSCGACWSFSATGALEGAHFLSTGKLVSLSEQ 178
S + + +P DWR+ GAVT VK+QG CG+CW+FS A+EG + + T KLVSLSEQ
Sbjct: 375 SFMYEKINRVPPSVDWRKKGAVTDVKDQGQCGSCWAFSTVVAVEGINQIKTNKLVSLSEQ 554
Query: 179 QLV 181
+LV
Sbjct: 555 ELV 563
>TC19860 weakly similar to UP|Q41064 (Q41064) Thiolprotease , partial (16%)
Length = 571
Score = 84.7 bits (208), Expect = 2e-17
Identities = 47/136 (34%), Positives = 79/136 (57%), Gaps = 10/136 (7%)
Frame = +3
Query: 45 FLEFKRRFGKVYVSEEEHGYRFNVFKSNMH-------RARRHQLLDPSAVHGVTRFSDLT 97
F ++++ G+ Y + EE RF+++++N+ + R + L D +F+DLT
Sbjct: 183 FESWQKQHGRKYENPEEWQVRFDIYQTNVEFIECINSQNRSYHLTD-------NKFADLT 341
Query: 98 PMEFRHSVLGLRGVGLPSDADSAPILRTD---NLPKDFDWREHGAVTPVKNQGSCGACWS 154
EF+ +G L +A A +LR + +LP+ DWR+ GAVT +K+QG+CG+CW+
Sbjct: 342 NEEFKRIYMGYGKTSLSCNA-GAGLLRYNGHGDLPESIDWRKKGAVTDIKDQGTCGSCWA 518
Query: 155 FSATGALEGAHFLSTG 170
FSA A+EG H + +G
Sbjct: 519 FSAVAAVEGIHQIKSG 566
>TC15660 weakly similar to UP|Q38886 (Q38886) SAG12 protein, partial (34%)
Length = 618
Score = 83.2 bits (204), Expect = 6e-17
Identities = 49/137 (35%), Positives = 66/137 (47%), Gaps = 6/137 (4%)
Frame = +1
Query: 220 EEDYPYSGTAGGTCKFDQTKIAASVANFSVVSRDEDQIAANLVKNGPLAVAINA--VYMQ 277
E DYPY G G K A +++ V + + + P++V I+A Q
Sbjct: 7 ERDYPYKGKDGTCNKEKAAHHAVTISGHKKVPASNEAMLKAAAAHQPVSVGIDAGGFLFQ 186
Query: 278 TYVGGVSCPYVCSKKLNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKIC 337
Y GGV + C K+LNH V +VGYG E P YW++KNSWG +WGE GY KI
Sbjct: 187 LYSGGVFSGF-CGKELNHAVTVVGYGREVNGP------KYWLVKNSWGADWGEAGYMKIL 345
Query: 338 RGR----NVCGVDSMVS 350
R CG+ + S
Sbjct: 346 RDTQDKDGTCGIAMVAS 396
>TC11715 weakly similar to UP|Q38886 (Q38886) SAG12 protein, partial (32%)
Length = 566
Score = 75.5 bits (184), Expect = 1e-14
Identities = 44/124 (35%), Positives = 65/124 (51%), Gaps = 8/124 (6%)
Frame = +1
Query: 241 AASVANFSVVSRDEDQIAANLVKNGPLAVAINA--VYMQTYVGGVSCPYVCSKKLNHGVL 298
A S++ V + + N P++VAI+A Q Y GGV + C K+LNH V
Sbjct: 16 AVSISGHEKVPASNEAMLKAAAANQPVSVAIDAGGFLFQLYSGGVFFGF-CGKQLNHAVT 192
Query: 299 LVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRGR----NVCGV--DSMVSTV 352
+VGYG + P YW++KNSWG +WGE+GY KI R CG+ D+ T+
Sbjct: 193 IVGYGRQVNGP------KYWLVKNSWGADWGESGYMKILRDTLDKDGTCGIAMDASYPTL 354
Query: 353 AALH 356
+ ++
Sbjct: 355 STIN 366
>AV409039
Length = 344
Score = 75.1 bits (183), Expect = 2e-14
Identities = 37/86 (43%), Positives = 49/86 (56%)
Frame = +3
Query: 139 AVTPVKNQGSCGACWSFSATGALEGAHFLSTGKLVSLSEQQLVDCDHECDPEEAGSCDSG 198
++ P + GACW+F+A ALEGA +++G LVSLSEQQL+DC G G
Sbjct: 108 SIIPAQEDLPSGACWAFAAVAALEGALQIASGTLVSLSEQQLIDC-------VKGGRSDG 266
Query: 199 CKGGLMNSAFEYILNNGGVMREEDYP 224
KGG AF+Y+ GG+ E YP
Sbjct: 267 FKGGFSEEAFDYVTRRGGISTEAGYP 344
>TC18996 similar to UP|O24323 (O24323) Cysteine proteinase precursor ,
partial (29%)
Length = 549
Score = 73.2 bits (178), Expect = 6e-14
Identities = 45/128 (35%), Positives = 65/128 (50%), Gaps = 11/128 (8%)
Frame = +2
Query: 47 EFKRRFGKVYVSEEEHGYRFNVFKSNMHRARRHQLLDPSAVHGVTRFSDLTPMEFRHSVL 106
E+ + GKVY + E RF +FK N+ H + + G+ RF+DL+ E+R L
Sbjct: 155 EWAMKQGKVYNALGEKEKRFEIFKDNLKFIDEHNAENRTYKVGLNRFADLSNEEYRAKFL 334
Query: 107 GLR----------GVGLPSDADSAPILRT-DNLPKDFDWREHGAVTPVKNQGSCGACWSF 155
G+R S+ AP + + LP+ DWR+ GAV VK+QG CG+ W+
Sbjct: 335 GIRVDSNRRRRRMAKSTTSNHRYAPRVGDGEKLPESVDWRKEGAVVGVKDQGECGSAWAL 514
Query: 156 SATGALEG 163
SA A EG
Sbjct: 515 SAVAAGEG 538
>BP068646
Length = 491
Score = 71.2 bits (173), Expect = 2e-13
Identities = 44/118 (37%), Positives = 58/118 (48%), Gaps = 6/118 (5%)
Frame = -3
Query: 239 KIAASVANFSVVSRDEDQIAANLVKNGPLAVAINAV--YMQTYVGGVSCPYVCSKKLNHG 296
K AAS+ F V + V N P++VAI+A Q Y GV C +L+HG
Sbjct: 489 KDAASLKGFEDVPAHSESALLKAVANQPISVAIDASGSEFQCYSSGVFTGS-CGTELDHG 313
Query: 297 VLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRG----RNVCGVDSMVS 350
V VGYGS+ YW++KNSWGE WGE GY ++ R +CG+ S
Sbjct: 312 VTAVGYGSDGGTQ-------YWLVKNSWGEQWGEQGYIRMQRDVAAEEGLCGIAMQAS 160
>TC20011 similar to GB|CAA40073.1|1345573|PVEPC1 endopeptidase (EP-C1)
{Phaseolus vulgaris;} , partial (29%)
Length = 472
Score = 69.7 bits (169), Expect = 7e-13
Identities = 39/89 (43%), Positives = 50/89 (55%), Gaps = 7/89 (7%)
Frame = +1
Query: 264 NGPLAVAINA-VYMQTYVGGVSCP--YVCSKKLNHGVLLVGYGSESYAPIRMKQKPYWII 320
N P++VA+ A V MQ Y GV C +LNHGV +VGYG+ Y YWI+
Sbjct: 40 NQPVSVAVEAGVDMQLYSEGVFMGGGVDCGTQLNHGVAIVGYGATQYGT------QYWIV 201
Query: 321 KNSWGENWGENGYYKICRG----RNVCGV 345
KNSWG WGE GY ++ R R +CG+
Sbjct: 202 KNSWGAKWGEEGYIRMKRDISDKRGLCGI 288
>TC18116 similar to UP|AAP32193 (AAP32193) Cysteine protease 14, partial
(37%)
Length = 555
Score = 65.9 bits (159), Expect = 1e-11
Identities = 43/127 (33%), Positives = 63/127 (48%), Gaps = 7/127 (5%)
Frame = +3
Query: 231 GTCKFDQTKI-AASVANFSVVSRDEDQIAANLVKNGPLAVAINAVY--MQTYVGGVSCPY 287
GTC+ + + +++ + V ++ +Q + N PL+VAI A Q Y GGV +
Sbjct: 30 GTCEMSKEESQVVTISGYHDVPQNNEQSLLKALANQPLSVAIEASGRDFQFYSGGVFDGH 209
Query: 288 VCSKKLNHGVLLVGYGSESYAPIRMKQKPYWIIKNSWGENWGENGYYKICRG----RNVC 343
C +L+HGV VGYG+ K Y +KNSWG WGE GY + R +C
Sbjct: 210 -CGTQLDHGVAAVGYGTS-------KGLDYITVKNSWGTKWGEKGYIRFRRNNGKPEGMC 365
Query: 344 GVDSMVS 350
G+ M S
Sbjct: 366 GLYKMAS 386
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.320 0.138 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,696,781
Number of Sequences: 28460
Number of extensions: 99601
Number of successful extensions: 568
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 534
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 539
length of query: 360
length of database: 4,897,600
effective HSP length: 91
effective length of query: 269
effective length of database: 2,307,740
effective search space: 620782060
effective search space used: 620782060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0258b.5


