

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258b.1
(207 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
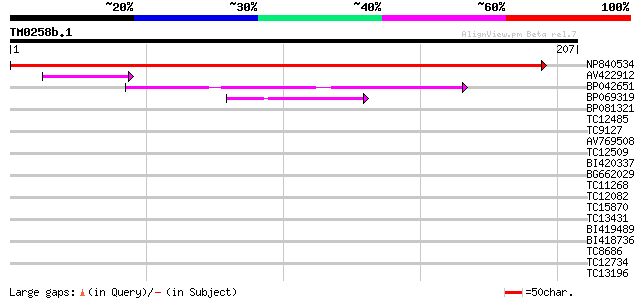
NP840534 hypothetical protein [Lotus corniculatus var. japonicus] 413 e-116
AV422912 40 4e-04
BP042651 39 5e-04
BP069319 39 8e-04
BP081321 33 0.046
TC12485 similar to UP|O80906 (O80906) Expressed protein, partial... 32 0.060
TC9127 weakly similar to UP|LYM2_ARATH (O23006) LysM-domain GPI-... 32 0.10
AV769508 31 0.13
TC12509 similar to UP|ACRO_HUMAN (P10323) Acrosin precursor , p... 30 0.23
BI420337 30 0.23
BG662029 30 0.39
TC11268 similar to UP|Q7SBH0 (Q7SBH0) Predicted protein, partial... 29 0.51
TC12082 weakly similar to UP|Q40385 (Q40385) Pistil extensin-lik... 29 0.67
TC15870 28 0.87
TC13431 28 0.87
BI419489 28 1.1
BI418736 28 1.1
TC8686 similar to UP|O23724 (O23724) GAI protein, partial (22%) 28 1.1
TC12734 similar to UP|Q9LM78 (Q9LM78) F2D10.25, partial (11%) 28 1.5
TC13196 28 1.5
>NP840534 hypothetical protein [Lotus corniculatus var. japonicus]
Length = 741
Score = 413 bits (1062), Expect = e-116
Identities = 196/196 (100%), Positives = 196/196 (100%)
Frame = +1
Query: 1 MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM 60
MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM
Sbjct: 1 MSMMLGQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLM 180
Query: 61 EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM 120
EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM
Sbjct: 181 EHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEM 360
Query: 121 AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATA 180
AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATA
Sbjct: 361 AYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTTIHSHDGATMAPPSSSATA 540
Query: 181 AHELFFPQPRSLASFR 196
AHELFFPQPRSLASFR
Sbjct: 541 AHELFFPQPRSLASFR 588
>AV422912
Length = 460
Score = 39.7 bits (91), Expect = 4e-04
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +3
Query: 13 RCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHV 45
+C++CGN A+ C Y C++CC C HV
Sbjct: 162 KCKQCGNVARSRCPYECCKSCCARNQNPCHIHV 260
>BP042651
Length = 534
Score = 39.3 bits (90), Expect = 5e-04
Identities = 31/125 (24%), Positives = 51/125 (40%)
Frame = +3
Query: 43 THVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSM 102
+H + IP+DRR H+ M T+ HH P S P +E L + +
Sbjct: 165 SHFSTFPIPIDRRNHQTMMDSENHTSEEHHH----HPLSAAAAPREEMESLALHDDDGNG 332
Query: 103 AVFSSVRVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQNLNLTT 162
V SS + S N + ++++ H S + E L + P++ H N +
Sbjct: 333 EVLSSSKSYS-----NYRSVMSTLSESRHPLSPPIVSTPAESDPLLSPPMNHHHNREFSN 497
Query: 163 IHSHD 167
+SHD
Sbjct: 498 PNSHD 512
>BP069319
Length = 494
Score = 38.5 bits (88), Expect = 8e-04
Identities = 20/52 (38%), Positives = 31/52 (59%)
Frame = -2
Query: 80 QSHNQNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQTSVNIGGH 131
++ + +P S +E P + + AVF VRV +++D +E AYQ V IGGH
Sbjct: 442 ETSSSHPDASFKET-MPGQVRAPAVFKCVRVTAVEDGKDEYAYQAVVKIGGH 290
>BP081321
Length = 369
Score = 32.7 bits (73), Expect = 0.046
Identities = 16/40 (40%), Positives = 22/40 (55%)
Frame = -3
Query: 118 NEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQN 157
+E AY +V+I GH F G LYD G ++ S L +N
Sbjct: 346 DEFAYLATVHISGHVFQGFLYDHGVGGKNACVSELQLGKN 227
>TC12485 similar to UP|O80906 (O80906) Expressed protein, partial (77%)
Length = 750
Score = 32.3 bits (72), Expect = 0.060
Identities = 21/60 (35%), Positives = 28/60 (46%), Gaps = 2/60 (3%)
Frame = +3
Query: 25 CAYSRCRTCCNNKGFQCQTHVRSTWIP--VDRRRHRLMEHQPPPTTNNPHHLHEDIPQSH 82
C+ S+C+T + G RST IP V R M PPP N +HLH ++ H
Sbjct: 159 CSQSKCKTGRTSMGIS-----RSTSIP*KV*TRNQIGMLKNPPPNHRNQNHLHYNLDAYH 323
>TC9127 weakly similar to UP|LYM2_ARATH (O23006) LysM-domain GPI-anchored
protein 2 precursor, partial (46%)
Length = 1016
Score = 31.6 bits (70), Expect = 0.10
Identities = 19/47 (40%), Positives = 24/47 (50%), Gaps = 2/47 (4%)
Frame = +3
Query: 40 QCQTHVRSTWIPVDRRRHRLM--EHQPPPTTNNPHHLHEDIPQSHNQ 84
Q Q H+R PV R R L H PPP P+ LH +IP H++
Sbjct: 405 QVQVHLRER--PVSRTRRLLSPKRHHPPPH---PNPLHREIPPGHSR 530
>AV769508
Length = 386
Score = 31.2 bits (69), Expect = 0.13
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +2
Query: 39 FQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNP 71
+Q +T+ +++ P RRHR HQPPP+ + P
Sbjct: 275 YQSETNPSTSYPPTSSRRHRRR-HQPPPSPSQP 370
>TC12509 similar to UP|ACRO_HUMAN (P10323) Acrosin precursor , partial (5%)
Length = 710
Score = 30.4 bits (67), Expect = 0.23
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +3
Query: 57 HRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFP 96
H + +PPPTTN+ HH H PFT L+ P
Sbjct: 177 HYRSQTRPPPTTNHHHHHH---------LPFTPLQTFNLP 269
>BI420337
Length = 397
Score = 30.4 bits (67), Expect = 0.23
Identities = 12/30 (40%), Positives = 19/30 (63%)
Frame = +2
Query: 137 LYDQGPEQQSLNASPLDQHQNLNLTTIHSH 166
+Y Q QQ++ + L HQ+L +TT+H H
Sbjct: 59 IYCQFKSQQTITSIHLHHHQSLTITTLHHH 148
>BG662029
Length = 161
Score = 29.6 bits (65), Expect = 0.39
Identities = 12/43 (27%), Positives = 20/43 (45%), Gaps = 1/43 (2%)
Frame = -2
Query: 38 GFQCQT-HVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIP 79
GF+C + +++ +P + HQPP + H H D P
Sbjct: 142 GFRCHSVQIKTLIVPTSLSSSSVYHHQPPHQLHRRRHFHRDPP 14
>TC11268 similar to UP|Q7SBH0 (Q7SBH0) Predicted protein, partial (5%)
Length = 1097
Score = 29.3 bits (64), Expect = 0.51
Identities = 16/53 (30%), Positives = 24/53 (45%)
Frame = -1
Query: 35 NNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPF 87
N K QTH S + H+ + P N+ HHLH P ++ ++PF
Sbjct: 827 NKKSISSQTHSPSNTL------HKNINLLHPHDLNHLHHLHPHHPLNNTKSPF 687
>TC12082 weakly similar to UP|Q40385 (Q40385) Pistil extensin-like protein,
partial (11%)
Length = 308
Score = 28.9 bits (63), Expect = 0.67
Identities = 18/52 (34%), Positives = 22/52 (41%), Gaps = 3/52 (5%)
Frame = +2
Query: 39 FQCQTH--VRSTWIPVDRRRHRLMEHQPP-PTTNNPHHLHEDIPQSHNQNPF 87
F Q H S +P +R H + H P PT +PHH H Q PF
Sbjct: 104 FAAQHHHGTHSLPLPPPQRHHHPLHHGPSIPTIQHPHHHHH-----QQQQPF 244
>TC15870
Length = 512
Score = 28.5 bits (62), Expect = 0.87
Identities = 12/41 (29%), Positives = 20/41 (48%)
Frame = -1
Query: 49 WIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTS 89
W+P L + Q NNP +L++++PQ + F S
Sbjct: 434 WLPFKTMEKTLQQWQQGEQNNNPFYLYQNLPQLYRPPFFHS 312
>TC13431
Length = 552
Score = 28.5 bits (62), Expect = 0.87
Identities = 12/35 (34%), Positives = 18/35 (51%), Gaps = 2/35 (5%)
Frame = +3
Query: 3 MMLGQEVKGSRCQECGN--QAKKGCAYSRCRTCCN 35
+ L + K RC +C Q + GC + +CR CN
Sbjct: 138 LKLAKRKKWQRCPKCSFYVQRRSGCEHMKCRCGCN 242
>BI419489
Length = 479
Score = 28.1 bits (61), Expect = 1.1
Identities = 16/52 (30%), Positives = 21/52 (39%), Gaps = 1/52 (1%)
Frame = +3
Query: 46 RSTWIPVDRRRHRLMEHQPPPTTNNPHHLHE-DIPQSHNQNPFTSLEELKFP 96
R T P+ ++ HQPP + PHHL H P EE + P
Sbjct: 315 RGTLRPLPPSPPGILHHQPPHCQDEPHHLRRLAAAPHHTLQPHPHREESQQP 470
>BI418736
Length = 544
Score = 28.1 bits (61), Expect = 1.1
Identities = 18/41 (43%), Positives = 23/41 (55%), Gaps = 3/41 (7%)
Frame = +3
Query: 55 RRHRLMEHQPPPTTNNPHHLHEDIPQ---SHNQNPFTSLEE 92
RRH H PPPT PH+LH P+ +H Q+PF + E
Sbjct: 210 RRHL---HLPPPT-RPPHNLHRQ-PRHRTTHPQDPFLQIPE 317
>TC8686 similar to UP|O23724 (O23724) GAI protein, partial (22%)
Length = 786
Score = 28.1 bits (61), Expect = 1.1
Identities = 16/39 (41%), Positives = 20/39 (51%), Gaps = 6/39 (15%)
Frame = +3
Query: 51 PVDRRRHRLM------EHQPPPTTNNPHHLHEDIPQSHN 83
P+ RRHR M + Q P + N HHL PQSH+
Sbjct: 303 PLQPRRHRHMA*NHALQFQFRPHSGNHHHLRLRRPQSHS 419
>TC12734 similar to UP|Q9LM78 (Q9LM78) F2D10.25, partial (11%)
Length = 936
Score = 27.7 bits (60), Expect = 1.5
Identities = 11/29 (37%), Positives = 15/29 (50%)
Frame = -3
Query: 22 KKGCAYSRCRTCCNNKGFQCQTHVRSTWI 50
++GC Y+ R C C TH+R WI
Sbjct: 553 ERGCCYTHLRGC-------CYTHLRGCWI 488
>TC13196
Length = 490
Score = 27.7 bits (60), Expect = 1.5
Identities = 14/44 (31%), Positives = 23/44 (51%)
Frame = -2
Query: 43 THVRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNP 86
T + S+++P+ L +H P +PHH H SH++NP
Sbjct: 315 TTLSSSYVPLS-----LSDHHSTPR*TSPHHHHH---SSHHRNP 208
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.317 0.129 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,756,427
Number of Sequences: 28460
Number of extensions: 91346
Number of successful extensions: 933
Number of sequences better than 10.0: 98
Number of HSP's better than 10.0 without gapping: 897
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 922
length of query: 207
length of database: 4,897,600
effective HSP length: 86
effective length of query: 121
effective length of database: 2,450,040
effective search space: 296454840
effective search space used: 296454840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0258b.1


