

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
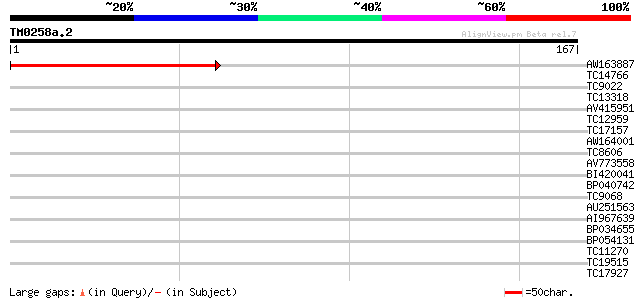
Query= TM0258a.2
(167 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW163887 125 4e-30
TC14766 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 36 0.004
TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Ara... 33 0.033
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 32 0.055
AV415951 31 0.12
TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%) 30 0.16
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 30 0.28
AW164001 30 0.28
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 30 0.28
AV773558 30 0.28
BI420041 29 0.36
BP040742 29 0.47
TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gluc... 29 0.47
AU251563 28 0.61
AI967639 28 0.80
BP034655 28 1.0
BP054131 28 1.0
TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%) 28 1.0
TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, pa... 28 1.0
TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete 27 1.4
>AW163887
Length = 260
Score = 125 bits (314), Expect = 4e-30
Identities = 62/62 (100%), Positives = 62/62 (100%)
Frame = +2
Query: 1 MENQPMQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIH 60
MENQPMQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIH
Sbjct: 68 MENQPMQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIH 247
Query: 61 QI 62
QI
Sbjct: 248 QI 253
>TC14766 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (58%)
Length = 600
Score = 35.8 bits (81), Expect = 0.004
Identities = 20/58 (34%), Positives = 30/58 (51%), Gaps = 2/58 (3%)
Frame = +2
Query: 84 YLHGYAYEDDEEQVVDDEEDDDDR--CDLDDELVPWEVGNKLGRQRMKKLGKRACSKM 139
+L DDE V DD+EDDDD D DD+ + + G+ GR + + K++ M
Sbjct: 95 HLEQQKIHDDEPVVEDDDEDDDDEDDDDEDDDNIEGQEGDASGRSKQTRSEKKSRKAM 268
>TC9022 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (75%)
Length = 1083
Score = 32.7 bits (73), Expect = 0.033
Identities = 30/114 (26%), Positives = 49/114 (42%), Gaps = 3/114 (2%)
Frame = +3
Query: 29 LVDVVDAEEEQEQEPDQNQQDAAP-VEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHG 87
L V+ AEE ++ Q Q+D AP VE++K
Sbjct: 144 LEQVLSAEETTLKKKPQPQEDDAPIVEDVK------------------------------ 233
Query: 88 YAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLG--RQRMKKLGKRACSKM 139
+DD+++ DDE++DDD D DD+ +G G + R +K ++A K+
Sbjct: 234 ---DDDKDEADDDEDEDDD--DEDDDKEDGALGGSEGSKQSRSEKKSRKAMLKL 380
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 32.0 bits (71), Expect = 0.055
Identities = 13/23 (56%), Positives = 18/23 (77%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDD+E+ D++EDDDD D DD+
Sbjct: 261 EDDDEEEDDEDEDDDDDDDDDDD 329
Score = 29.6 bits (65), Expect = 0.28
Identities = 11/23 (47%), Positives = 18/23 (77%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
E+D+++ DDE++DDD D DD+
Sbjct: 258 EEDDDEEEDDEDEDDDDDDDDDD 326
Score = 28.5 bits (62), Expect = 0.61
Identities = 11/23 (47%), Positives = 18/23 (77%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+++EE+ D+EEDD+D D DD+
Sbjct: 246 DNEEEEDDDEEEDDEDEDDDDDD 314
Score = 27.3 bits (59), Expect = 1.4
Identities = 12/23 (52%), Positives = 18/23 (78%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDDE++ DD++DDDD D ++E
Sbjct: 279 EDDEDE--DDDDDDDDDDDGEEE 341
Score = 26.9 bits (58), Expect = 1.8
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+D+EE D+++DDDD D D E
Sbjct: 267 DDEEEDDEDEDDDDDDDDDDDGE 335
Score = 25.4 bits (54), Expect = 5.2
Identities = 9/23 (39%), Positives = 17/23 (73%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+D++E DD++DDDD + +D+
Sbjct: 282 DDEDEDDDDDDDDDDDGEEEEDD 350
Score = 25.4 bits (54), Expect = 5.2
Identities = 9/23 (39%), Positives = 16/23 (69%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+ D E+ DD+E++DD + DD+
Sbjct: 240 QTDNEEEEDDDEEEDDEDEDDDD 308
Score = 25.0 bits (53), Expect = 6.8
Identities = 13/25 (52%), Positives = 17/25 (68%), Gaps = 3/25 (12%)
Frame = +3
Query: 92 DDEEQVVDDEEDDD---DRCDLDDE 113
D+EE+ DDEE+DD D D DD+
Sbjct: 246 DNEEEEDDDEEEDDEDEDDDDDDDD 320
Score = 25.0 bits (53), Expect = 6.8
Identities = 14/49 (28%), Positives = 25/49 (50%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKM 139
+DD++ ++EEDDDD + ++ EV K G + +A + M
Sbjct: 309 DDDDDDDGEEEEDDDDDDEEEEGSEEVEVEGKDGAKGAAAQTTKAAAAM 455
Score = 25.0 bits (53), Expect = 6.8
Identities = 9/23 (39%), Positives = 16/23 (69%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
++DE+ DD++DDD + DD+
Sbjct: 285 DEDEDDDDDDDDDDDGEEEEDDD 353
>AV415951
Length = 366
Score = 30.8 bits (68), Expect = 0.12
Identities = 12/30 (40%), Positives = 22/30 (73%)
Frame = +2
Query: 86 HGYAYEDDEEQVVDDEEDDDDRCDLDDELV 115
HGYAYE+++ D+EE++D D +D+++
Sbjct: 263 HGYAYEEED----DEEEEEDKWVDWEDQIL 340
>TC12959 similar to UP|AAS52042 (AAS52042) ADR122Cp, partial (6%)
Length = 517
Score = 30.4 bits (67), Expect = 0.16
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +1
Query: 87 GYAYEDDEEQVVDDEEDDDDRCDLDDELVPW 117
G + +DD++ ++EDDDD D DDE W
Sbjct: 118 GSSSDDDDDDDDSEDEDDDDNEDDDDEW*SW 210
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 29.6 bits (65), Expect = 0.28
Identities = 18/56 (32%), Positives = 28/56 (49%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
++DE+ D+EE+DDD D DDE G+ + K K K + ++R P
Sbjct: 800 DEDEDDDDDEEEEDDDDDDEDDE-------EGEGKSKSKSGSKARPGKDQPTERPP 946
Score = 26.2 bits (56), Expect = 3.0
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = +2
Query: 87 GYAYEDDEEQVVDDEEDDDDRCDLDDE 113
G A + D E + +D+ED D+ D DD+
Sbjct: 740 GEAEQSDFEDIEEDDEDGDEDEDEDDD 820
Score = 25.4 bits (54), Expect = 5.2
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = +2
Query: 90 YEDDEEQVVDDEEDDDDRCDLDDE 113
+ED EE D +ED+D+ D D+E
Sbjct: 761 FEDIEEDDEDGDEDEDEDDDDDEE 832
>AW164001
Length = 447
Score = 29.6 bits (65), Expect = 0.28
Identities = 12/23 (52%), Positives = 17/23 (73%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
E+D ++ DD++DDDD D DDE
Sbjct: 67 EEDVDEDFDDDDDDDDDDDDDDE 135
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 29.6 bits (65), Expect = 0.28
Identities = 12/23 (52%), Positives = 18/23 (78%)
Frame = -2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
ED+E++ +D+EDDDD + DDE
Sbjct: 431 EDEEDEDGEDQEDDDDDDEDDDE 363
Score = 27.7 bits (60), Expect = 1.0
Identities = 10/23 (43%), Positives = 19/23 (82%)
Frame = -2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
ED+EE+ V++E+++D+ D +DE
Sbjct: 347 EDEEEEGVEEEDNEDEEEDEEDE 279
Score = 26.2 bits (56), Expect = 3.0
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = -2
Query: 87 GYAYEDDEEQVVDDEEDDDDRCDLDDELVPWE 118
G EDD++ DD+E+DD D ++E V E
Sbjct: 410 GEDQEDDDDDDEDDDEEDDGGEDEEEEGVEEE 315
Score = 25.4 bits (54), Expect = 5.2
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = -2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
++++E D E+DDDD D D+E
Sbjct: 428 DEEDEDGEDQEDDDDDDEDDDEE 360
>AV773558
Length = 469
Score = 29.6 bits (65), Expect = 0.28
Identities = 11/24 (45%), Positives = 17/24 (70%)
Frame = -2
Query: 90 YEDDEEQVVDDEEDDDDRCDLDDE 113
Y +D++ DD++DDDD D +DE
Sbjct: 372 YSEDDDDDDDDDDDDDDYTDDEDE 301
Score = 26.9 bits (58), Expect = 1.8
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = -2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDD++ DD++DDD D D+E
Sbjct: 366 EDDDDDDDDDDDDDDYTDDEDEE 298
Score = 25.0 bits (53), Expect = 6.8
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = -2
Query: 92 DDEEQVVDDEEDDDDRCDLDDE 113
DD+ +V E+DDDD D DD+
Sbjct: 393 DDDFVLVYSEDDDDDDDDDDDD 328
>BI420041
Length = 452
Score = 29.3 bits (64), Expect = 0.36
Identities = 19/73 (26%), Positives = 35/73 (47%)
Frame = +2
Query: 45 QNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDD 104
+N D+ PV E I+ ++ ++ E D Y ED +E DDE+++
Sbjct: 188 RNSSDSEPVLEPTIVQEV--------SMDEEEDDEFDYEEPLNDEDEDGDEYYDDDEQEE 343
Query: 105 DDRCDLDDELVPW 117
++ D D++ VP+
Sbjct: 344 EEFEDEDEDSVPY 382
>BP040742
Length = 465
Score = 28.9 bits (63), Expect = 0.47
Identities = 13/24 (54%), Positives = 17/24 (70%)
Frame = -1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDEL 114
EDD+E DD++D+DD DL EL
Sbjct: 462 EDDDEDDEDDDDDEDDD*DLCAEL 391
>TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (6%)
Length = 1209
Score = 28.9 bits (63), Expect = 0.47
Identities = 11/23 (47%), Positives = 17/23 (73%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+DD++ DD++DDDD D DD+
Sbjct: 86 DDDDDDDDDDDDDDDDDDDDDDD 154
Score = 28.9 bits (63), Expect = 0.47
Identities = 11/23 (47%), Positives = 17/23 (73%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+DD++ DD++DDDD D DD+
Sbjct: 83 DDDDDDDDDDDDDDDDDDDDDDD 151
Score = 26.9 bits (58), Expect = 1.8
Identities = 11/26 (42%), Positives = 17/26 (65%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVP 116
+DD++ DD++DDDD D D+ P
Sbjct: 95 DDDDDDDDDDDDDDDDDDDDGDKKQP 172
Score = 26.9 bits (58), Expect = 1.8
Identities = 12/33 (36%), Positives = 19/33 (57%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKL 123
+DD++ DD++DDDD D D + + G L
Sbjct: 98 DDDDDDDDDDDDDDDDDDDGDKKQPDSDTGTAL 196
Score = 26.2 bits (56), Expect = 3.0
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
+DD++ DD++DDDD D D +
Sbjct: 92 DDDDDDDDDDDDDDDDDDDDDGD 160
>AU251563
Length = 350
Score = 28.5 bits (62), Expect = 0.61
Identities = 12/16 (75%), Positives = 13/16 (81%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDD 106
ED E+ VVDDEEDD D
Sbjct: 233 EDSEDAVVDDEEDDYD 280
>AI967639
Length = 335
Score = 28.1 bits (61), Expect = 0.80
Identities = 18/50 (36%), Positives = 28/50 (56%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMK 140
+ DE + D+EEDDDD D DD+L E + +++K+L R + K
Sbjct: 127 QGDEYNLEDEEEDDDD--DDDDDLSISEDSDSDKPRKVKQLPGRTRRETK 270
Score = 26.9 bits (58), Expect = 1.8
Identities = 17/57 (29%), Positives = 30/57 (51%)
Frame = +1
Query: 88 YAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKR 144
Y ED+EE DD++DD + D P +V GR R ++ +R+ ++++ R
Sbjct: 139 YNLEDEEEDDDDDDDDDLSISEDSDSDKPRKVKQLPGRTR-RETKRRSVDEIQSGLR 306
>BP034655
Length = 517
Score = 27.7 bits (60), Expect = 1.0
Identities = 11/26 (42%), Positives = 20/26 (76%)
Frame = +1
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVP 116
E++EE+ +DEEDD++ D D++ +P
Sbjct: 193 EEEEEEEEEDEEDDEE--DEDEDYIP 264
>BP054131
Length = 500
Score = 27.7 bits (60), Expect = 1.0
Identities = 11/23 (47%), Positives = 17/23 (73%)
Frame = -2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
++DE++V +DEE DDD +DE
Sbjct: 262 DEDEDEVEEDEEMDDDLVYTEDE 194
>TC11270 similar to UP|Q9LM88 (Q9LM88) F2D10.15, partial (9%)
Length = 735
Score = 27.7 bits (60), Expect = 1.0
Identities = 12/41 (29%), Positives = 23/41 (55%)
Frame = +2
Query: 67 PDPDAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDR 107
P A + + R+ + + Y D+E++ D++EDDD+R
Sbjct: 164 PATTAQSTLRRSRRPRNVRYNIDYLDEEDEDEDEDEDDDER 286
Score = 25.8 bits (55), Expect = 4.0
Identities = 17/78 (21%), Positives = 31/78 (38%)
Frame = +3
Query: 36 EEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEE 95
EEE+E E D+ +Q P + +A E+DEE
Sbjct: 339 EEEEEDEEDEEEQTPHPTRGRSRVE---------------------------HAQEEDEE 437
Query: 96 QVVDDEEDDDDRCDLDDE 113
+ +EE++++ ++DE
Sbjct: 438 EENGEEEEEEEEVVVEDE 491
>TC19515 similar to UP|Q9YGK0 (Q9YGK0) Vitellogenin precursor, partial (3%)
Length = 504
Score = 27.7 bits (60), Expect = 1.0
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +3
Query: 92 DDEEQVVDDEEDDDDRCDLDDE 113
D ++ +DEEDDDD D DDE
Sbjct: 69 DPDDAESEDEEDDDDDDDGDDE 134
>TC17927 UP|Q8GRY7 (Q8GRY7) bZIP with a Ring-finger motif, complete
Length = 1793
Score = 27.3 bits (59), Expect = 1.4
Identities = 13/38 (34%), Positives = 21/38 (55%), Gaps = 2/38 (5%)
Frame = +1
Query: 88 YAYEDDEEQVVDDEEDDDDRCDLDD--ELVPWEVGNKL 123
Y + Q+ D++DDDD D+DD V + GN++
Sbjct: 715 YESHKESSQMEGDDDDDDDEDDVDDLENEVKYGQGNRM 828
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,874,070
Number of Sequences: 28460
Number of extensions: 37340
Number of successful extensions: 572
Number of sequences better than 10.0: 85
Number of HSP's better than 10.0 without gapping: 289
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 413
length of query: 167
length of database: 4,897,600
effective HSP length: 84
effective length of query: 83
effective length of database: 2,506,960
effective search space: 208077680
effective search space used: 208077680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0258a.2


