

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0253.9
(677 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
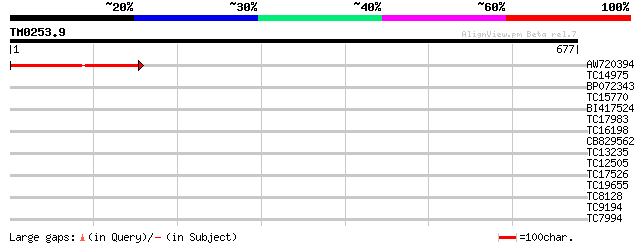
Score E
Sequences producing significant alignments: (bits) Value
AW720394 305 2e-83
TC14975 homologue to UP|Q8LG37 (Q8LG37) AtPH1-like protein, part... 34 0.088
BP072343 33 0.12
TC15770 similar to UP|ENPL_CATRO (P35016) Endoplasmin homolog pr... 31 0.57
BI417524 30 1.7
TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , part... 30 1.7
TC16198 29 2.8
CB829562 28 3.7
TC13235 similar to UP|Q9LQB6 (Q9LQB6) F4N2.1, partial (38%) 28 3.7
TC12505 similar to UP|BAD00698 (BAD00698) Sericin 1A, partial (5%) 28 4.8
TC17526 similar to UP|ADF_VITVI (Q8SAG3) Actin-depolymerizing fa... 28 4.8
TC19655 28 6.3
TC8128 homologue to UP|TBA3_ELEIN (O22349) Tubulin alpha-3 chain... 28 6.3
TC9194 similar to UP|Q9LYP1 (Q9LYP1) Membrane protein (AT5g07250... 27 8.3
TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomeras... 27 8.3
>AW720394
Length = 587
Score = 305 bits (780), Expect = 2e-83
Identities = 157/159 (98%), Positives = 157/159 (98%)
Frame = +1
Query: 1 MEKNKIDETVAEQQQPLDPLSNYRGLSLFPSTSSALPLSVADDLDAIHNHLRSMALRSPA 60
MEKNKIDETVAEQQQPLDPLSNYRGLSLFPSTSSALPLSVADDLDAIHNHLRSMALRSPA
Sbjct: 115 MEKNKIDETVAEQQQPLDPLSNYRGLSLFPSTSSALPLSVADDLDAIHNHLRSMALRSPA 294
Query: 61 KLVEQTKAILEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLGLKRARFSLKP 120
KLVEQTKAILEENPGLFNSQDDERNDV VDVDVAEDGQDFPRKRRPGLGLKRARFSLKP
Sbjct: 295 KLVEQTKAILEENPGLFNSQDDERNDV--VDVDVAEDGQDFPRKRRPGLGLKRARFSLKP 468
Query: 121 STSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQ 159
STSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQ
Sbjct: 469 STSVSLESLLPNLDIENLKDPVEFFKAHERMENARREIQ 585
>TC14975 homologue to UP|Q8LG37 (Q8LG37) AtPH1-like protein, partial (86%)
Length = 892
Score = 33.9 bits (76), Expect = 0.088
Identities = 22/57 (38%), Positives = 29/57 (50%), Gaps = 1/57 (1%)
Frame = +3
Query: 308 RGSLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPF-APLSSLLN 363
+ S SKP A + N N +KTP SP+ +SP+PP SP AP +S N
Sbjct: 201 KASTSKPGAAAGSFSNKENSSGSKTP--PSRASPSHAASSPSPPVSPSRAPKTSSTN 365
>BP072343
Length = 517
Score = 33.5 bits (75), Expect = 0.12
Identities = 16/63 (25%), Positives = 34/63 (53%)
Frame = +3
Query: 608 EQKRVKSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGERLVYGRIH 667
E++R +++++ K +R +R+ + +RRS R + R LE W+ ++ R+H
Sbjct: 69 EKRRGNAKAKQAWKGRRSKQRKRQPKQLSKHKRKLRRSLRRQRRKLELWRNRQMKSSRLH 248
Query: 668 QSK 670
+ K
Sbjct: 249 RKK 257
>TC15770 similar to UP|ENPL_CATRO (P35016) Endoplasmin homolog precursor
(GRP94 homolog), partial (43%)
Length = 1065
Score = 31.2 bits (69), Expect = 0.57
Identities = 16/61 (26%), Positives = 33/61 (53%)
Frame = +3
Query: 33 SSALPLSVADDLDAIHNHLRSMALRSPAKLVEQTKAILEENPGLFNSQDDERNDVVDVDV 92
S LPL+V+ ++ H+ L+++ + K ++ + I +E+P S D E+ + D+
Sbjct: 318 SDTLPLNVSREMLQAHSSLKTIKKKLIRKALDMIRRIADEDPD--ESTDKEKKEETSSDI 491
Query: 93 D 93
D
Sbjct: 492 D 494
>BI417524
Length = 462
Score = 29.6 bits (65), Expect = 1.7
Identities = 20/61 (32%), Positives = 27/61 (43%), Gaps = 2/61 (3%)
Frame = +3
Query: 303 DLKSLRGSLSKPRKALSN--IDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSS 360
D RGS PR+ + +L+ + T LR Q SP PP +P +P SS
Sbjct: 78 DEDETRGSSGNPRRLHQQRELGQVLHHSRFTTTLRMVRRMAPTQNPSPLPPPNPSSPSSS 257
Query: 361 L 361
L
Sbjct: 258 L 260
>TC17983 similar to UP|Q9XEX5 (Q9XEX5) Cytidine deaminase , partial (42%)
Length = 519
Score = 29.6 bits (65), Expect = 1.7
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = +3
Query: 313 KPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSV 372
+PR+A+ L +R NK+PL + P L P PPR+P +P S +H+ R +P +
Sbjct: 249 RPRRAVPPHQPLPQQR-NKSPLLRRLRRPLWPLP-PVPPRTPRSPGDSNPHHL-RTRPEI 419
>TC16198
Length = 572
Score = 28.9 bits (63), Expect = 2.8
Identities = 28/101 (27%), Positives = 44/101 (42%), Gaps = 10/101 (9%)
Frame = +2
Query: 276 LLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRGSLSKPRKALSNIDNL------LNRRK 329
+LQ L + + VSV P P S R S +KP + L+ NL +R K
Sbjct: 158 ILQSSLVEMAMAISSVSVALRPHFPPSPFHSRRYSFAKPLRCLATKQNLPETLEIESRPK 337
Query: 330 NKTPLRQDAGSPAEQLASPTP----PRSPFAPLSSLLNHIS 366
++T L + P A+ P RS FAP+ ++ ++
Sbjct: 338 SETVLHSFSPLPLLYAAALLPGGGAVRSVFAPVVEIVKTLN 460
>CB829562
Length = 530
Score = 28.5 bits (62), Expect = 3.7
Identities = 14/60 (23%), Positives = 29/60 (48%)
Frame = +1
Query: 406 PSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKEDNSGKSSNKLDAPLIEDILAVS 465
P+N+ + +T++++ ++ PD S SEN K D K D +++++S
Sbjct: 88 PNNKESTEVTRESLIAISSSYSPDKNLESNSVSENKKSDGVAKPKCDQDDDFRSELISIS 267
>TC13235 similar to UP|Q9LQB6 (Q9LQB6) F4N2.1, partial (38%)
Length = 467
Score = 28.5 bits (62), Expect = 3.7
Identities = 11/28 (39%), Positives = 16/28 (56%)
Frame = +3
Query: 327 RRKNKTPLRQDAGSPAEQLASPTPPRSP 354
RR+N+ P P ++SP PPR+P
Sbjct: 114 RRRNRKPSPAPPPPPGSPISSPKPPRNP 197
>TC12505 similar to UP|BAD00698 (BAD00698) Sericin 1A, partial (5%)
Length = 569
Score = 28.1 bits (61), Expect = 4.8
Identities = 21/78 (26%), Positives = 33/78 (41%), Gaps = 6/78 (7%)
Frame = +2
Query: 575 VIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDTKSKRLAKR------ 628
V +A+S K +T +E S++ D S E S S + +
Sbjct: 230 VSRASSTGKPSTTKHEKSDKL*PRVNDASCDGFENATTSSSSSSSSSRSSASSSLFLFGC 409
Query: 629 QSLAAAGTSWTSGIRRST 646
+ +AAAG++WTS ST
Sbjct: 410 EEVAAAGSTWTSSSSSST 463
>TC17526 similar to UP|ADF_VITVI (Q8SAG3) Actin-depolymerizing factor (ADF),
partial (71%)
Length = 433
Score = 28.1 bits (61), Expect = 4.8
Identities = 12/39 (30%), Positives = 18/39 (45%)
Frame = -3
Query: 319 SNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAP 357
S ++ N + KTP + + P Q + P PP S P
Sbjct: 335 SQTQHIYNHSQAKTPRNRHSSQPGHQSSQPQPPFSSHQP 219
>TC19655
Length = 473
Score = 27.7 bits (60), Expect = 6.3
Identities = 14/52 (26%), Positives = 24/52 (45%)
Frame = -2
Query: 476 NSTSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGETVMDDTVAK 527
NS S + + D + P N +E H D+ +NGG + + + + AK
Sbjct: 379 NSNSATAVASTDALVNPPKATNGSTDEKHVDLFTENGGVSVWDEINEALNAK 224
>TC8128 homologue to UP|TBA3_ELEIN (O22349) Tubulin alpha-3 chain (Alpha-3
tubulin), complete
Length = 1615
Score = 27.7 bits (60), Expect = 6.3
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = +2
Query: 613 KSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLEYWKGER 660
+SR+ + +S R +L T G RR ++R RPL W+G R
Sbjct: 311 RSRTHRHRRSPRRHLPPTLPPRTTHLRQG-RRRQQLRQRPLHRWQGNR 451
>TC9194 similar to UP|Q9LYP1 (Q9LYP1) Membrane protein
(AT5g07250/T28J14_190), partial (78%)
Length = 1467
Score = 27.3 bits (59), Expect = 8.3
Identities = 17/46 (36%), Positives = 21/46 (44%)
Frame = +2
Query: 325 LNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKP 370
L ++ NK L A + A PP PF LS L + RLKP
Sbjct: 47 LKKKNNKFKLLLQAYNEAVTNLRSLPPPFPFPFLSQFLKNSFRLKP 184
>TC7994 similar to UP|Q8RU73 (Q8RU73) Ribose-5-phosphate isomerase
precursor , partial (81%)
Length = 1355
Score = 27.3 bits (59), Expect = 8.3
Identities = 19/66 (28%), Positives = 32/66 (47%), Gaps = 1/66 (1%)
Frame = +1
Query: 34 SALPLSVADDLDAIHNHLRS-MALRSPAKLVEQTKAILEENPGLFNSQDDERNDVVDVDV 92
++L LS L + H++ S + LR+P L +T A + + +QDD + D V
Sbjct: 259 ASLSLSSPPSLSSAHHNASSHLILRTPTSLKLRTSATPKPIRAITLTQDDLKRLAADKAV 438
Query: 93 DVAEDG 98
D + G
Sbjct: 439 DYVKSG 456
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.309 0.128 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,390,847
Number of Sequences: 28460
Number of extensions: 114707
Number of successful extensions: 633
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 623
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 632
length of query: 677
length of database: 4,897,600
effective HSP length: 96
effective length of query: 581
effective length of database: 2,165,440
effective search space: 1258120640
effective search space used: 1258120640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0253.9


