

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0252a.5
(150 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
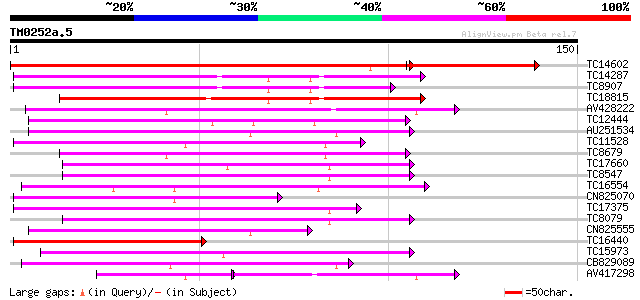
Score E
Sequences producing significant alignments: (bits) Value
TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinas... 188 9e-59
TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein k... 86 3e-18
TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein ki... 86 3e-18
TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein ki... 79 4e-16
AV428222 78 7e-16
TC12444 similar to UP|O22981 (O22981) Ser/Thr protein kinase iso... 75 5e-15
AU251534 67 1e-12
TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial... 66 2e-12
TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partia... 65 4e-12
TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, part... 65 6e-12
TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated pro... 62 4e-11
TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase ki... 62 5e-11
CN825070 62 5e-11
TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corn... 61 7e-11
TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete 59 3e-10
CN825555 57 1e-09
TC16440 similar to UP|Q93Y18 (Q93Y18) SNF1 related protein kinas... 56 3e-09
TC15973 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F... 55 5e-09
CB829089 55 5e-09
AV417298 43 7e-09
>TC14602 UP|Q8GS56 (Q8GS56) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1256
Score = 188 bits (478), Expect(2) = 9e-59
Identities = 96/126 (76%), Positives = 97/126 (76%), Gaps = 19/126 (15%)
Frame = +2
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDG 60
G IPEVE A LMKQLLEA+ HCHRLGVAH DVKPDNVLFG GGDLKLADFGSAEWFGDG
Sbjct: 353 GTSIPEVEAAGLMKQLLEAVAHCHRLGVAHRDVKPDNVLFGGGGDLKLADFGSAEWFGDG 532
Query: 61 RRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIW 101
RRMSGVVGTPYYVAPEVLMG E GEKVDVWSCGV IFEAVI
Sbjct: 533 RRMSGVVGTPYYVAPEVLMGREYGEKVDVWSCGVILYIMLSGTPPFYGDSAAEIFEAVIR 712
Query: 102 GNLRFP 107
GNLRFP
Sbjct: 713 GNLRFP 730
Score = 53.5 bits (127), Expect(2) = 9e-59
Identities = 30/35 (85%), Positives = 31/35 (87%)
Frame = +3
Query: 106 FPLESSGMFLLLLRIS*GR*FAGTLLTESLQNKH* 140
F LESSGMFLLLL+IS*GR*FA TLLTES NKH*
Sbjct: 726 FLLESSGMFLLLLKIS*GR*FAETLLTESQLNKH* 830
>TC14287 similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein kinase-like
protein (CBL-interacting protein kinase 5), partial
(75%)
Length = 1999
Score = 85.9 bits (211), Expect = 3e-18
Identities = 49/115 (42%), Positives = 69/115 (59%), Gaps = 6/115 (5%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E + +QL+ A+ CH GV H D+KP+N+L DLK++DFG + + R
Sbjct: 708 GKLNEDDARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDENEDLKVSDFGLSA-LPEQR 884
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
R G++ GTP YVAPEVL G++ G K D+WSCGVI A++ G L F E+
Sbjct: 885 RDDGMLVTPCGTPAYVAPEVLKKKGYD-GSKADIWSCGVILYALLSGYLPFQGEN 1046
>TC8907 similar to UP|Q8H0X3 (Q8H0X3) Serine/threonine protein kinase-like
protein, partial (44%)
Length = 941
Score = 85.9 bits (211), Expect = 3e-18
Identities = 47/107 (43%), Positives = 65/107 (59%), Gaps = 6/107 (5%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + EV +QL+ A+ CH GV H D+KP+N+L DLK++DFG + + R
Sbjct: 618 GKMTEVAARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDDNEDLKVSDFGLSS-LPEQR 794
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWG 102
R G++ GTP YVAPEVL G++ G K D+WSCGVI A++ G
Sbjct: 795 RSDGMLLTPCGTPAYVAPEVLKKKGYD-GSKADIWSCGVILYALLCG 932
>TC18815 similar to UP|Q8W1D5 (Q8W1D5) CBL-interacting protein kinase
CIPK25, partial (58%)
Length = 979
Score = 78.6 bits (192), Expect = 4e-16
Identities = 45/103 (43%), Positives = 64/103 (61%), Gaps = 6/103 (5%)
Frame = +2
Query: 14 KQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVV----GT 69
+QL+ A+ +CH GV+H D+KP+N+L +LK++DFG + R G++ GT
Sbjct: 350 QQLISAVDYCHSRGVSHRDLKPENLLLDENENLKVSDFG-LSGLPEQLRQDGLLHTQCGT 526
Query: 70 PYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
P YVAPEVL G++ G K D WSCGVI A++ G L F E+
Sbjct: 527 PAYVAPEVLRKKGYD-GFKTDTWSCGVILYALLAGCLPFQHEN 652
>AV428222
Length = 416
Score = 77.8 bits (190), Expect = 7e-16
Identities = 45/119 (37%), Positives = 70/119 (58%), Gaps = 4/119 (3%)
Frame = +3
Query: 5 PEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLF---GAGGDLKLADFGSAEWFGDGR 61
PE + +++ Q+L + CH GV H D+KP+N LF A +K+ DFG +++ +
Sbjct: 9 PEDDAKAILLQILNVVAFCHLHGVVHRDLKPENFLFVSKEADSVMKVIDFGLSDFVRPDQ 188
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF-PLESSGMFLLLLR 119
R++ +VG+ YYVAPEVL + E D+WS GVI ++ G+ F SG+F +LR
Sbjct: 189 RLNDIVGSAYYVAPEVLHRSYSVE-ADLWSIGVISYILLCGSRPFWARTESGIFRSVLR 362
>TC12444 similar to UP|O22981 (O22981) Ser/Thr protein kinase isolog,
partial (37%)
Length = 709
Score = 75.1 bits (183), Expect = 5e-15
Identities = 45/106 (42%), Positives = 61/106 (57%), Gaps = 5/106 (4%)
Frame = +2
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFG-SAEWFGDGRRM- 63
E + S++K+ L+A+ + HR G H DVK N+L G+ G +KLADFG SA F G R
Sbjct: 380 EEAIGSILKETLKALEYLHRHGHIHRDVKAGNILLGSDGQVKLADFGVSASMFDAGERQR 559
Query: 64 --SGVVGTPYYVAPEVLM-GWENGEKVDVWSCGVIFEAVIWGNLRF 106
+ VGTP ++APEVL G K DVWS G+ + G+ F
Sbjct: 560 NRNTFVGTPCWMAPEVLQPGTGYNFKADVWSFGITALELAHGHAPF 697
>AU251534
Length = 391
Score = 67.0 bits (162), Expect = 1e-12
Identities = 40/105 (38%), Positives = 56/105 (53%), Gaps = 3/105 (2%)
Frame = +2
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR--M 63
E +V M+QLL + HCH V H D+K N+L G LK+ADFG A F + M
Sbjct: 11 EPQVKCYMRQLLSGLEHCHHRRVLHRDIKGSNLLIDNEGVLKIADFGLASVFDPNHKHPM 190
Query: 64 SGVVGTPYYVAPEVLMGWENGE-KVDVWSCGVIFEAVIWGNLRFP 107
+ V T +Y PE+L+G + + VD+WS G I ++ G P
Sbjct: 191 TSRVVTLWYRPPELLLGATDYDVGVDLWSAGCILGELLAGKPIMP 325
>TC11528 homologue to UP|Q39868 (Q39868) Protein kinase , partial (56%)
Length = 640
Score = 66.2 bits (160), Expect = 2e-12
Identities = 36/96 (37%), Positives = 52/96 (53%), Gaps = 3/96 (3%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD--LKLADFGSAEWFGD 59
G E E +QL+ +++CH + + H D+K +N L LK+ DFG ++
Sbjct: 345 GRFSEDEARYFFQQLISGVSYCHSMEICHRDLKLENTLLDGNPSPRLKICDFGYSKSAIL 524
Query: 60 GRRMSGVVGTPYYVAPEVLMGWE-NGEKVDVWSCGV 94
+ VGTP Y+APEVL E +G+ DVWSCGV
Sbjct: 525 HSQPKSTVGTPAYIAPEVLSRKEYDGKVADVWSCGV 632
>TC8679 homologue to UP|Q43466 (Q43466) Protein kinase 3 , partial (70%)
Length = 972
Score = 65.5 bits (158), Expect = 4e-12
Identities = 37/96 (38%), Positives = 52/96 (53%), Gaps = 3/96 (3%)
Frame = +2
Query: 14 KQLLEAITHCHRLGVAHHDVKPDNVLF--GAGGDLKLADFGSAEWFGDGRRMSGVVGTPY 71
+QL+ + +CH L + H D+K +N L LK+ DFG ++ R VGTP
Sbjct: 2 EQLISGVHYCHALQICHRDLKLENTLLDGSPAPRLKICDFGYSKSSLLHSRPKSTVGTPA 181
Query: 72 YVAPEVLMGWE-NGEKVDVWSCGVIFEAVIWGNLRF 106
Y+APEVL E +G+ DVWSCGV ++ G F
Sbjct: 182 YIAPEVLSRREYDGKLADVWSCGVTLYVMLVGAYPF 289
>TC17660 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, partial (64%)
Length = 717
Score = 64.7 bits (156), Expect = 6e-12
Identities = 37/95 (38%), Positives = 52/95 (53%), Gaps = 2/95 (2%)
Frame = +2
Query: 15 QLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEW-FGDGRRMSGVVGTPYYV 73
QLL + + H + H D+KP N+L A DLK+ DFG A + M+ V T +Y
Sbjct: 257 QLLRGLKYLHSANILHRDLKPGNLLINANCDLKICDFGLARINCSKNQFMTEYVVTRWYR 436
Query: 74 APEVLMGWEN-GEKVDVWSCGVIFEAVIWGNLRFP 107
APE+L+ +N G +DVWS G IF ++ FP
Sbjct: 437 APELLLCCDNYGTSIDVWSVGCIFAELLGRKPIFP 541
>TC8547 homologue to UP|MMK1_MEDSA (Q07176) Mitogen-activated protein
kinase homolog MMK1 (MAP kinase MSK7) (MAP kinase ERK1)
, partial (71%)
Length = 1098
Score = 62.0 bits (149), Expect = 4e-11
Identities = 35/94 (37%), Positives = 50/94 (52%), Gaps = 1/94 (1%)
Frame = +1
Query: 15 QLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVVGTPYYVA 74
Q+L + + H V H D+KP N+L A DLK+ DFG A + M+ V T +Y A
Sbjct: 160 QILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVTSETDFMTEYVVTRWYRA 339
Query: 75 PEVLMGWEN-GEKVDVWSCGVIFEAVIWGNLRFP 107
PE+L+ + +DVWS G IF ++ FP
Sbjct: 340 PELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFP 441
>TC16554 homologue to UP|AAN71719 (AAN71719) Glycogen synthase kinase 3 beta
protein kinase DWARF12 , partial (71%)
Length = 927
Score = 61.6 bits (148), Expect = 5e-11
Identities = 35/111 (31%), Positives = 57/111 (50%), Gaps = 3/111 (2%)
Frame = +1
Query: 4 IPEVEVASLMKQLLEAITHCHRL-GVAHHDVKPDNVLFGA-GGDLKLADFGSAEWFGDGR 61
+P + V M Q+ + + H + GV H D+KP N+L +KL DFGSA+ G
Sbjct: 526 MPIIYVKLYMYQIFRGLAYIHTVPGVCHRDLKPQNILVDPLTHQVKLCDFGSAKMLVKGE 705
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLESS 111
+ + +Y APE++ G E +D+WS G + ++ G FP E++
Sbjct: 706 ANISYICSRFYRAPELIFGATEYSTSIDIWSAGCVLAELLLGQPLFPGENA 858
>CN825070
Length = 634
Score = 61.6 bits (148), Expect = 5e-11
Identities = 30/74 (40%), Positives = 41/74 (54%), Gaps = 3/74 (4%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSAEWFG 58
G E E A L++ ++E + CH LGV H D+KP+N LF LK DFG + ++
Sbjct: 411 GQYSEREAAKLIRTIVEVVEACHSLGVMHRDLKPENFLFDTVEEDAKLKTTDFGLSVFYK 590
Query: 59 DGRRMSGVVGTPYY 72
G VVG+PYY
Sbjct: 591 PGETFGDVVGSPYY 632
>TC17375 mitogen-activated kinase kinase kinase alpha [Lotus corniculatus
var. japonicus]
Length = 1422
Score = 61.2 bits (147), Expect = 7e-11
Identities = 30/93 (32%), Positives = 48/93 (51%), Gaps = 1/93 (1%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G E + + +Q++ + + H H D+K N+L G++KLADFG ++
Sbjct: 216 GAFKEPVIQNYTRQIVSGLAYLHSRNTVHRDIKGANILVDPNGEIKLADFGMSKHINSAA 395
Query: 62 RMSGVVGTPYYVAPEVLMGWEN-GEKVDVWSCG 93
M G+PY++APEV+M G VD+ S G
Sbjct: 396 SMLSFKGSPYWMAPEVVMNTNGYGLPVDISSLG 494
>TC8079 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, complete
Length = 1550
Score = 59.3 bits (142), Expect = 3e-10
Identities = 33/94 (35%), Positives = 50/94 (53%), Gaps = 1/94 (1%)
Frame = +3
Query: 15 QLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVVGTPYYVA 74
Q+L + + H + H D+KP N+L + DLK+ DFG A + M+ V T +Y A
Sbjct: 618 QVLRGLKYIHSANIIHRDLKPSNLLLNSNCDLKIIDFGLARPTVENDFMTEYVVTRWYRA 797
Query: 75 PEVLMGWEN-GEKVDVWSCGVIFEAVIWGNLRFP 107
PE+L+ + +DVWS G IF ++ FP
Sbjct: 798 PELLLNSSDYTSAIDVWSVGCIFMELMNKKPLFP 899
>CN825555
Length = 714
Score = 57.0 bits (136), Expect = 1e-09
Identities = 31/77 (40%), Positives = 42/77 (54%), Gaps = 2/77 (2%)
Frame = +1
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR--M 63
E +V M QL + HCH V H D+K N+L G LK+ADFG A +F + M
Sbjct: 478 ESQVKCYMHQLFTGLEHCHNRHVLHRDIKGSNLLIDNEGVLKIADFGLASFFDPDHKHPM 657
Query: 64 SGVVGTPYYVAPEVLMG 80
+ V T +Y PE+L+G
Sbjct: 658 TSRVVTLWYRPPELLLG 708
>TC16440 similar to UP|Q93Y18 (Q93Y18) SNF1 related protein kinase, partial
(30%)
Length = 549
Score = 55.8 bits (133), Expect = 3e-09
Identities = 25/51 (49%), Positives = 34/51 (66%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFG 52
G +PE +QL+ A+ CHR GVAH D+KP N+L A G+LK++DFG
Sbjct: 373 GRLPEPLARRYFQQLVSALCFCHRNGVAHRDLKPQNLLLDAEGNLKVSDFG 525
>TC15973 homologue to GB|AAL15278.1|16323087|AY057647 AT3g01490/F4P13_4
{Arabidopsis thaliana;}, partial (49%)
Length = 944
Score = 55.1 bits (131), Expect = 5e-09
Identities = 33/100 (33%), Positives = 50/100 (50%), Gaps = 1/100 (1%)
Frame = +3
Query: 9 VASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAE-WFGDGRRMSGVV 67
V L L +++ H + H DVK +N+L +K+ADFG A + M+G
Sbjct: 63 VIQLALDLARGLSYLHSQKIVHRDVKTENMLLDKTRTVKIADFGVARVEASNPNDMTGET 242
Query: 68 GTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFP 107
GT Y+APEVL G K DV+S G+ + ++ +P
Sbjct: 243 GTLGYMAPEVLNGNPYNRKCDVYSFGICLWEIYCCDMPYP 362
>CB829089
Length = 278
Score = 55.1 bits (131), Expect = 5e-09
Identities = 27/90 (30%), Positives = 53/90 (58%), Gaps = 2/90 (2%)
Frame = +2
Query: 4 IPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFG-AGGDLKLADFGSAEWFGDGRR 62
+ + ++ + +LL+A+ +CH G+ H DVKP NV+ L L D+G AE++ G+
Sbjct: 5 LSDYDIRYYIYELLKALDYCHSQGIMHRDVKPHNVMIDHEQRKLHLIDWGLAEFYHPGKE 184
Query: 63 MSGVVGTPYYVAPEVLMGWENGE-KVDVWS 91
+ + + Y+ PE+L+ ++ + +D+WS
Sbjct: 185YNDRLASRYFKGPELLVDLQDYDYSLDLWS 274
>AV417298
Length = 424
Score = 42.7 bits (99), Expect(2) = 7e-09
Identities = 25/61 (40%), Positives = 36/61 (58%), Gaps = 1/61 (1%)
Frame = +1
Query: 60 GRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF-PLESSGMFLLLL 118
G++ +VG+ YYVAPEVL ++G + DVWS GVI ++ G F G+F +L
Sbjct: 145 GKKFQDIVGSAYYVAPEVLKR-KSGPESDVWSIGVITYILLCGRRPFWDKTEDGIFKEVL 321
Query: 119 R 119
R
Sbjct: 322 R 324
Score = 31.6 bits (70), Expect(2) = 7e-09
Identities = 15/40 (37%), Positives = 23/40 (57%), Gaps = 3/40 (7%)
Frame = +2
Query: 24 HRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFGDG 60
H G+ H D+KP+N LF + + LK DFG +++ G
Sbjct: 2 HLHGLVHRDMKPENFLFKSNREDSALKATDFGLSDFIKPG 121
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.334 0.150 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,941,559
Number of Sequences: 28460
Number of extensions: 38770
Number of successful extensions: 440
Number of sequences better than 10.0: 165
Number of HSP's better than 10.0 without gapping: 389
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 392
length of query: 150
length of database: 4,897,600
effective HSP length: 82
effective length of query: 68
effective length of database: 2,563,880
effective search space: 174343840
effective search space used: 174343840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 51 (24.3 bits)
Lotus: description of TM0252a.5


