

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0243.9
(310 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
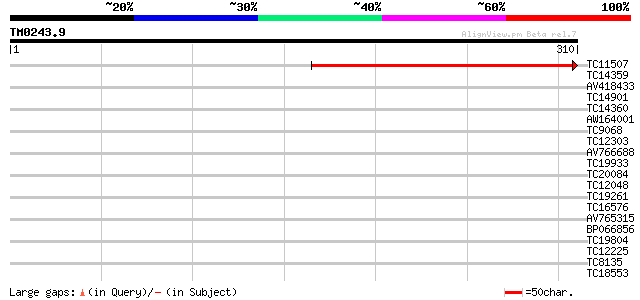
Score E
Sequences producing significant alignments: (bits) Value
TC11507 weakly similar to UP|Q9HD28 (Q9HD28) Retinitis pigmentos... 298 1e-81
TC14359 similar to UP|AAQ88099 (AAQ88099) NADPH-dependent cinnam... 30 0.52
AV418433 30 0.52
TC14901 similar to UP|CAE47100 (CAE47100) 4,5-DOPA dioxygenase e... 30 0.52
TC14360 similar to UP|O04391 (O04391) Cinnamyl alcohol dehydroge... 30 0.68
AW164001 29 0.89
TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gluc... 29 0.89
TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transpo... 29 1.2
AV766688 29 1.2
TC19933 28 1.5
TC20084 similar to UP|Q8LI88 (Q8LI88) OJ1641_C04.24 protein, par... 28 1.5
TC12048 weakly similar to UP|Q9LGA3 (Q9LGA3) ESTs D22655(C0749),... 28 1.5
TC19261 similar to UP|BAD06577 (BAD06577) Cell wall protein Awa1... 28 1.5
TC16576 weakly similar to UP|O76728 (O76728) Oligopeptidase B, p... 28 2.0
AV765315 28 2.0
BP066856 28 2.0
TC19804 similar to UP|Q9M2K2 (Q9M2K2) Phenylalanine-tRNA synthet... 28 2.6
TC12225 28 2.6
TC8135 similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-binding pr... 28 2.6
TC18553 similar to UP|AAR83868 (AAR83868) 60S ribosomal protein ... 28 2.6
>TC11507 weakly similar to UP|Q9HD28 (Q9HD28) Retinitis pigmentosa GTPase
regulator (Fragment), partial (3%)
Length = 744
Score = 298 bits (762), Expect = 1e-81
Identities = 143/145 (98%), Positives = 143/145 (98%)
Frame = +3
Query: 166 QTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQIWCE 225
QTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQIWCE
Sbjct: 15 QTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQIWCE 194
Query: 226 SKGRVPDLVRQWLYGSDPALNGEEQWWRNLGVSGSVSGSLLHLLHRLCFSLSESGAWLRK 285
SKGRVPDLVRQW YGSDPALNGEEQWWRNLGVSGSVSGSLLHLLHRLCFSLSESG WLRK
Sbjct: 195 SKGRVPDLVRQWFYGSDPALNGEEQWWRNLGVSGSVSGSLLHLLHRLCFSLSESGPWLRK 374
Query: 286 EGCDGDGFESRLELFVVFVAWTNFL 310
EGCDGDGFESRLELFVVFVAWTNFL
Sbjct: 375 EGCDGDGFESRLELFVVFVAWTNFL 449
>TC14359 similar to UP|AAQ88099 (AAQ88099) NADPH-dependent cinnamyl alcohol
dehydrogenase, partial (98%)
Length = 1403
Score = 30.0 bits (66), Expect = 0.52
Identities = 23/96 (23%), Positives = 32/96 (32%)
Frame = +1
Query: 157 EKGVLEGAFQTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGR 216
E+G + Q H + PF L T + S R V
Sbjct: 250 EEGSFDSVVQGCHGVFHTASPFYHDVKDPQVELLDPAVKGTLNVLKSCVNSPTLKRVVLT 429
Query: 217 SEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQWW 252
S + K R PD+V + +DP LN E + W
Sbjct: 430 SSIAAVAYNGKPRTPDVVVDETWFTDPVLNREAKMW 537
>AV418433
Length = 401
Score = 30.0 bits (66), Expect = 0.52
Identities = 18/48 (37%), Positives = 27/48 (55%)
Frame = +3
Query: 252 WRNLGVSGSVSGSLLHLLHRLCFSLSESGAWLRKEGCDGDGFESRLEL 299
WR L ++G + ++L L F + ES WL K GC D FE+ L++
Sbjct: 222 WRALALTGLIPTAVLLLG---LFFIPESPRWLAKRGCRED-FEAALQI 353
>TC14901 similar to UP|CAE47100 (CAE47100) 4,5-DOPA dioxygenase extradiol,
partial (71%)
Length = 1276
Score = 30.0 bits (66), Expect = 0.52
Identities = 21/56 (37%), Positives = 29/56 (51%), Gaps = 5/56 (8%)
Frame = +2
Query: 173 FRLFPFSFFS-----LPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQIW 223
FR P F S PLSPPLT I +T +++S ++ TS + + EL IW
Sbjct: 308 FRRDPLPFLSSPAIGTPLSPPLTLLIPPTTPSMTSMASPNPCTSLNI-QHLELLIW 472
>TC14360 similar to UP|O04391 (O04391) Cinnamyl alcohol dehydrogenase ,
partial (68%)
Length = 821
Score = 29.6 bits (65), Expect = 0.68
Identities = 23/96 (23%), Positives = 30/96 (30%)
Frame = +3
Query: 157 EKGVLEGAFQTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGR 216
E+G Q H + PF + L T + S R V
Sbjct: 255 EEGSFNSVVQGCHGVFHTASPFYYDVEDPQAELLDPAVKGTLNVLKSCANSPSLKRVVLT 434
Query: 217 SEELQIWCESKGRVPDLVRQWLYGSDPALNGEEQWW 252
S + K R PD+V + SDP N E W
Sbjct: 435 SSIAAVAYNGKSRTPDVVVDETWFSDPHFNLEANMW 542
>AW164001
Length = 447
Score = 29.3 bits (64), Expect = 0.89
Identities = 19/58 (32%), Positives = 30/58 (50%), Gaps = 5/58 (8%)
Frame = -2
Query: 159 GVLEGAFQT-----LHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTS 211
G+L G+ Q L Y+R S F L P + + SSS+ + SSSS++K ++
Sbjct: 248 GILRGSSQAGTPPALLGKYWRQRDHSSFGLSTKPRVAGSSSSSSSSSSSSSSSKSSST 75
>TC9068 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (6%)
Length = 1209
Score = 29.3 bits (64), Expect = 0.89
Identities = 18/49 (36%), Positives = 30/49 (60%)
Frame = -1
Query: 173 FRLFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQ 221
F+ P S LSP +S+ SSS+ + SSSS++ +S G+G ++ L+
Sbjct: 198 FKAVPVSESGCFLSPSSSSS-SSSSSSSSSSSSSSSSSSSGLGLNKRLK 55
>TC12303 similar to GB|CAB10424.1|2245004|ATFCA6 membrane transporter like
protein {Arabidopsis thaliana;}, partial (25%)
Length = 545
Score = 28.9 bits (63), Expect = 1.2
Identities = 14/33 (42%), Positives = 17/33 (51%)
Frame = +3
Query: 82 FFDCLSIFFTGDVQGKVEEADSRARRSGRSSRW 114
FF C S+ F ++G V EAD A R S W
Sbjct: 75 FFLCFSVLFFNTMEGGVPEADMSAFRECLSLSW 173
>AV766688
Length = 608
Score = 28.9 bits (63), Expect = 1.2
Identities = 13/27 (48%), Positives = 19/27 (70%)
Frame = +2
Query: 182 SLPLSPPLTSTISSSTGTISSSSTTKK 208
SL ++PP TS IS STG +S S++ +
Sbjct: 494 SLQVTPPATSQISDSTGALSPFSSSSE 574
>TC19933
Length = 472
Score = 28.5 bits (62), Expect = 1.5
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -3
Query: 251 WWRNLGVSGSVSGSLLHLLHRLCFSLSESGAWLRKEGCDGDG 292
WW ++ VS + H L +LC + LR+ GC DG
Sbjct: 146 WWSSI*-KAKVSNKVKHFLWKLCHNAIPLKGNLRRRGCATDG 24
>TC20084 similar to UP|Q8LI88 (Q8LI88) OJ1641_C04.24 protein, partial (14%)
Length = 461
Score = 28.5 bits (62), Expect = 1.5
Identities = 16/42 (38%), Positives = 22/42 (52%), Gaps = 3/42 (7%)
Frame = +1
Query: 177 PFSFFSLPLSPPLTSTISSSTGTISSS---STTKKKTSRGVG 215
PFS S P PP T++ +S T T+ S T S+G+G
Sbjct: 247 PFSTSSSPSPPPATTSSTSITDTVKGSHRFKITGYSLSKGIG 372
>TC12048 weakly similar to UP|Q9LGA3 (Q9LGA3) ESTs D22655(C0749), partial
(20%)
Length = 697
Score = 28.5 bits (62), Expect = 1.5
Identities = 17/49 (34%), Positives = 29/49 (58%)
Frame = -2
Query: 157 EKGVLEGAFQTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSSST 205
E + + A T+ G+F F F+FF+ + P ++T SSS+ + SS S+
Sbjct: 480 EASIFDAASATVTLGFFGFFTFTFFT-SSTFPFSATSSSSS*SPSSYSS 337
>TC19261 similar to UP|BAD06577 (BAD06577) Cell wall protein Awa1p, partial
(3%)
Length = 495
Score = 28.5 bits (62), Expect = 1.5
Identities = 23/77 (29%), Positives = 31/77 (39%), Gaps = 16/77 (20%)
Frame = +3
Query: 184 PLSPPLTSTISSSTG-------TISSSSTTKKKTSRGVGRSEELQIWCES----KGRVPD 232
P PP TS +SST T+S+SS + S G + C S KG++P
Sbjct: 93 PTPPPATSGYTSSTSTNTLSRITLSTSSASTSSASTGSASTSSASRSCGSSNSRKGKLPG 272
Query: 233 LVRQ-----WLYGSDPA 244
R+ W PA
Sbjct: 273 YQRRNGQRGWQQAGSPA 323
>TC16576 weakly similar to UP|O76728 (O76728) Oligopeptidase B, partial
(30%)
Length = 1202
Score = 28.1 bits (61), Expect = 2.0
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = -2
Query: 172 YFRLFPFSFFSLPLSPP 188
+F FPFS+ LP+SPP
Sbjct: 238 FFSSFPFSYHCLPISPP 188
>AV765315
Length = 510
Score = 28.1 bits (61), Expect = 2.0
Identities = 12/24 (50%), Positives = 15/24 (62%)
Frame = +1
Query: 177 PFSFFSLPLSPPLTSTISSSTGTI 200
P S F +PLSPPL + S S G +
Sbjct: 352 PISVFMIPLSPPLFTLFSCSLGLL 423
>BP066856
Length = 485
Score = 28.1 bits (61), Expect = 2.0
Identities = 16/37 (43%), Positives = 20/37 (53%), Gaps = 2/37 (5%)
Frame = +2
Query: 184 PLSPPLTSTI--SSSTGTISSSSTTKKKTSRGVGRSE 218
P PP TS S+STGTIS+ S + SR S+
Sbjct: 62 PTGPPTTSRSMSSASTGTISTGSASTTSASRSSSSSD 172
>TC19804 similar to UP|Q9M2K2 (Q9M2K2) Phenylalanine-tRNA synthetase-like
protein, partial (24%)
Length = 612
Score = 27.7 bits (60), Expect = 2.6
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +3
Query: 260 SVSGSLLHLLHRLCFSLSESGAWLRKEGCD 289
SVSGS+ HLL +C LSE + + C+
Sbjct: 75 SVSGSMNHLLKTICVKLSEE*PGILQRRCN 164
>TC12225
Length = 552
Score = 27.7 bits (60), Expect = 2.6
Identities = 18/44 (40%), Positives = 22/44 (49%)
Frame = -1
Query: 160 VLEGAFQTLHSGYFRLFPFSFFSLPLSPPLTSTISSSTGTISSS 203
+L G LH + FP F SLP P S+ +S G ISSS
Sbjct: 372 MLSGFLHLLHVLFSSCFP-PFLSLPCEPRSFSSSASLPGLISSS 244
>TC8135 similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-binding protein,
chloroplast precursor (Genomes uncoulped 4), partial
(51%)
Length = 1213
Score = 27.7 bits (60), Expect = 2.6
Identities = 17/47 (36%), Positives = 23/47 (48%)
Frame = +3
Query: 175 LFPFSFFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQ 221
LFP PL PPL+ +S+G+ S T+ K+ R S LQ
Sbjct: 297 LFPRQLPPPPLPPPLSPPPLTSSGSFSQPETSGKRMMRPGASS*PLQ 437
>TC18553 similar to UP|AAR83868 (AAR83868) 60S ribosomal protein L12,
partial (33%)
Length = 469
Score = 27.7 bits (60), Expect = 2.6
Identities = 18/52 (34%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Frame = -1
Query: 172 YFRLFPFS-FFSLPLSPPLTSTISSSTGTISSSSTTKKKTSRGVGRSEELQI 222
+ L P S FFS P+ PP +SS+T T + + ++ RS LQI
Sbjct: 316 FLLLLPLSIFFSQPIRPPPNPPLSSNTTTTTETPQIHNQSKIITTRSLWLQI 161
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.323 0.139 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,034,989
Number of Sequences: 28460
Number of extensions: 122322
Number of successful extensions: 1075
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 1003
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1054
length of query: 310
length of database: 4,897,600
effective HSP length: 90
effective length of query: 220
effective length of database: 2,336,200
effective search space: 513964000
effective search space used: 513964000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0243.9


