

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0243.12
(1626 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
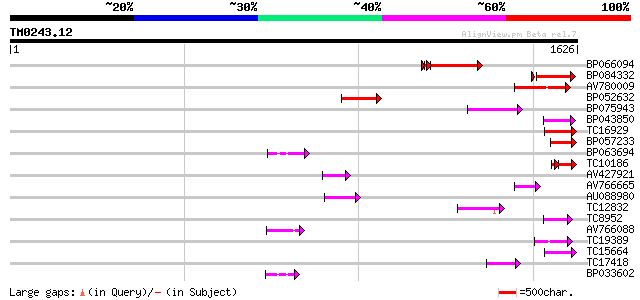
Score E
Sequences producing significant alignments: (bits) Value
BP066094 279 2e-80
BP084332 226 4e-59
AV780009 133 3e-31
BP052632 105 4e-23
BP075943 76 5e-14
BP043850 74 2e-13
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 73 4e-13
BP057233 68 1e-11
BP063694 67 2e-11
TC10186 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprot... 50 5e-11
AV427921 65 7e-11
AV766665 62 6e-10
AU088980 61 1e-09
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 59 5e-09
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 58 1e-08
AV766088 54 2e-07
TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 54 2e-07
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 54 3e-07
TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partia... 53 3e-07
BP033602 52 6e-07
>BP066094
Length = 532
Score = 279 bits (714), Expect(3) = 2e-80
Identities = 137/149 (91%), Positives = 142/149 (94%)
Frame = +2
Query: 1207 ARLEAIRLLISFSVNHNIVLHQMDVKSAFLNGYISEEVYVHQPPGFEDEKKPDHVFKLKK 1266
++ EAIRLLISFSVNHNI+LHQM+VKSAFLNGYISEEVYVHQPPG EDEK DH+FKLKK
Sbjct: 86 SKTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKK 265
Query: 1267 SLYGLKQAPRAWYERLSSFLLENEFVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANQ 1326
SLYGLKQAPRAWYERLSSFLLENE VRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSAN
Sbjct: 266 SLYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANP 445
Query: 1327 SLCKEFSEMMQAEFEMSMMGELKYFLGIQ 1355
SLCKEFSEMMQAEFEM MMGELKYFLGIQ
Sbjct: 446 SLCKEFSEMMQAEFEMRMMGELKYFLGIQ 532
Score = 37.4 bits (85), Expect(3) = 2e-80
Identities = 17/18 (94%), Positives = 17/18 (94%)
Frame = +3
Query: 1191 YSQQEGIDYTETFAPVAR 1208
YSQQ GIDYTETFAPVAR
Sbjct: 39 YSQQ*GIDYTETFAPVAR 92
Score = 23.1 bits (48), Expect(3) = 2e-80
Identities = 10/10 (100%), Positives = 10/10 (100%)
Frame = +1
Query: 1181 RNKARLVAQG 1190
RNKARLVAQG
Sbjct: 10 RNKARLVAQG 39
>BP084332
Length = 368
Score = 226 bits (575), Expect(2) = 4e-59
Identities = 111/112 (99%), Positives = 111/112 (99%)
Frame = +2
Query: 1511 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS 1570
QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYCDNTAAISLSKNPILHS 211
Query: 1571 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFILKNL 1622
RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTK LAEDRFNFILKNL
Sbjct: 212 RAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFNFILKNL 367
Score = 21.6 bits (44), Expect(2) = 4e-59
Identities = 8/8 (100%), Positives = 8/8 (100%)
Frame = +1
Query: 1496 CQFLGSNL 1503
CQFLGSNL
Sbjct: 4 CQFLGSNL 27
>AV780009
Length = 529
Score = 133 bits (334), Expect = 3e-31
Identities = 68/167 (40%), Positives = 105/167 (62%), Gaps = 6/167 (3%)
Frame = -1
Query: 1448 AVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSWA 1507
A R+LRY+KG GL + S KL Y D+D+AG R+S +G FLG++L+SW
Sbjct: 526 AATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWR 347
Query: 1508 SKRQSTIALSTAEAEY--ISAAICSTQMLWMKHQLEDYQILESN----IPIYCDNTAAIS 1561
+K+Q+T++ S++EAEY ++A +C Q W+ + +Q L+ N +P++CDN +A+
Sbjct: 346 TKKQTTVSRSSSEAEYRALAATVCEVQ--WLSYL---FQFLKLNVPLPVPLFCDNQSALH 182
Query: 1562 LSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTK 1608
++ NP H R KHIE+ H +R +Q G++ L + T HQ ADIFTK
Sbjct: 181 IAHNPTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTK 41
>BP052632
Length = 489
Score = 105 bits (263), Expect = 4e-23
Identities = 50/114 (43%), Positives = 71/114 (61%)
Frame = -3
Query: 952 KTPYELWKNIKPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGFRFYNT 1011
KTPYELW+ +P +SYFHPFGC ++LNT D + KFD K K +LLGYS SK + +N+
Sbjct: 487 KTPYELWRGRRPTVSYFHPFGCK*FILNTNDNIGKFDNKFDKRILLGYSSNSKAYIVFNS 308
Query: 1012 DAKTIEESIHVRFDDKLDSDQSKLVEKFADLSINVSDKGKAPEEVEPEEDEPEE 1065
+ +EESI+V+FDD +L + F +L ++ D+ E +P E EE
Sbjct: 307 RTQVVEESINVKFDD*SIEATYRLEKDFLNLDLSDQDEEVPTNEGQPSETSHEE 146
>BP075943
Length = 547
Score = 75.9 bits (185), Expect = 5e-14
Identities = 45/159 (28%), Positives = 78/159 (48%), Gaps = 2/159 (1%)
Frame = -3
Query: 1314 IYVDDIIFGSANQSLCKEFSEMMQAEFEMSMMGELKYFLGIQVDQTPEGTYIHQSKYTKE 1373
+YVDD++ + + + +F + +GE KYFLG+++ ++ G ++Q KY +
Sbjct: 515 LYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYALQ 336
Query: 1374 LLKKFNMLESTVAKTPMHPTCILEKEDKSGKVCQKL--YRGMIGSLLYLTASRPDILFSV 1431
L+ + H T +G + YR ++G LLYL +RPDI F+V
Sbjct: 335 LISDSGHF-GFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPDITFAV 159
Query: 1432 HLCARFQSDPRETHLTAVKRILRYLKGTTNLGLMYKKTS 1470
+ ++F S P + H + L+YL G+ GL Y +S
Sbjct: 158 NQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGLFYPASS 42
>BP043850
Length = 515
Score = 73.6 bits (179), Expect = 2e-13
Identities = 35/93 (37%), Positives = 56/93 (59%), Gaps = 1/93 (1%)
Frame = -1
Query: 1531 TQMLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKG 1589
++++W++ L + L+S P++ DNT+AI ++ NP+ H +HIEV H +R+ +
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 1590 VLLLKFVDTDHQWADIFTKSLAEDRFNFILKNL 1622
V+ L V T Q ADI TKSL R NF++ L
Sbjct: 335 VITLPHVSTSVQIADILTKSLTRQRHNFLVSKL 237
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 72.8 bits (177), Expect = 4e-13
Identities = 32/91 (35%), Positives = 55/91 (60%), Gaps = 1/91 (1%)
Frame = +1
Query: 1535 WMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLL 1593
W+ + L+D ++ E +YCDN +A ++ NP+ H R KHIE+ H +R+ +QKG++ L
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 1594 KFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+ + ADI+TK+L+ F+ I L +
Sbjct: 184 LPISSSEPLADIYTKALSPQNFHQICAKLGL 276
>BP057233
Length = 473
Score = 67.8 bits (164), Expect = 1e-11
Identities = 30/73 (41%), Positives = 49/73 (67%)
Frame = -2
Query: 1552 IYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLA 1611
++CDN +A +L+ NP+LH+R+KHIE+ H+IRD V + +++ +V T Q AD TK L+
Sbjct: 445 LWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIADCLTKPLS 266
Query: 1612 EDRFNFILKNLNM 1624
RF+ + L +
Sbjct: 265 HTRFSQLRDKLGV 227
>BP063694
Length = 511
Score = 67.0 bits (162), Expect = 2e-11
Identities = 44/122 (36%), Positives = 60/122 (49%), Gaps = 3/122 (2%)
Frame = -2
Query: 740 VWHRRLGHASMRKISQLSK---LNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVS 796
+WH RLGH + I QL K L++ LP C +CQ K ++ F N
Sbjct: 360 LWHSRLGHVNFDIIKQLHKHGCLDVSSILPK-----PICCTSCQMAKSKRLVFHDNNK-R 199
Query: 797 TSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDESHAVFSTFIAQV 856
S L+L+H DL GP SI G Y ++ VDD+SR+TW L RK + V F +
Sbjct: 198 ASAVLDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVLLRFKVFM 19
Query: 857 QN 858
+N
Sbjct: 18 EN 13
>TC10186 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein, partial
(4%)
Length = 528
Score = 50.4 bits (119), Expect(2) = 5e-11
Identities = 26/51 (50%), Positives = 32/51 (61%)
Frame = +3
Query: 1574 HIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+IE KYHF+RD V KG + LK TD Q ADI TK L +RF + LN+
Sbjct: 69 NIETKYHFLRDQVTKGKISLKHCGTDLQVADIMTKGLKTERFRNMRAMLNV 221
Score = 35.4 bits (80), Expect(2) = 5e-11
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = +2
Query: 1555 DNTAAISLSKNPILHSRAKHIE 1576
DN +AI L+KNP+ H R+KHIE
Sbjct: 5 DNKSAIDLAKNPVSHGRSKHIE 70
>AV427921
Length = 284
Score = 65.5 bits (158), Expect = 7e-11
Identities = 32/79 (40%), Positives = 44/79 (55%)
Frame = +1
Query: 898 TPQQNGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYEL 957
TPQQNGV ERKNRT+ M R++L +G+ K F E V + +I NR + N P E
Sbjct: 22 TPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEEA 201
Query: 958 WKNIKPNISYFHPFGCVCY 976
W +P + +F C+ Y
Sbjct: 202 WSGRQPAVDHFRISRCLAY 258
>AV766665
Length = 601
Score = 62.4 bits (150), Expect = 6e-10
Identities = 32/76 (42%), Positives = 44/76 (57%)
Frame = +3
Query: 1447 TAVKRILRYLKGTTNLGLMYKKTSEYKLSGYCDADYAGDRTERKSTSGNCQFLGSNLVSW 1506
T RI YLK + G +++K + + G+ ADY G +R ST G FL NLV+W
Sbjct: 360 TFTTRIF*YLKANSRRGPLFQKEGKSSMDGFTYADYLGSIVDRLSTMGYYMFLSGNLVTW 539
Query: 1507 ASKRQSTIALSTAEAE 1522
SK+Q+ IA S+ EAE
Sbjct: 540 RSKQQNIIARSSGEAE 587
>AU088980
Length = 360
Score = 61.2 bits (147), Expect = 1e-09
Identities = 28/105 (26%), Positives = 50/105 (46%)
Frame = +2
Query: 902 NGVVERKNRTLQEMARTMLQETGMAKHFWAETVNTACYIQNRISVRPILNKTPYELWKNI 961
N +VERK+R + + R ++ + + K+ W V A Y+ N + + + PY+ N
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 962 KPNISYFHPFGCVCYVLNTKDRLHKFDAKSSKCLLLGYSDRSKGF 1006
P + FG +CY K D ++ KC+ LG+ + G+
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGY 316
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 59.3 bits (142), Expect = 5e-09
Identities = 37/145 (25%), Positives = 71/145 (48%), Gaps = 9/145 (6%)
Frame = +2
Query: 1284 SFLLENEFVRGKVDTTLFCKTYKD-DILIVQIYVDDIIFGSANQSLCKEFSEMMQAEFEM 1342
SF++ + R D + K + D D +I+ +YVDD++ N+ +E + EF+M
Sbjct: 8 SFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREFDM 187
Query: 1343 SMMGELKYFLGIQV--DQTPEGTYIHQSKYTKELLKKFNMLESTVAKTP------MHPTC 1394
+G LG+Q+ D+ ++ Q Y +++L++FNM + TP + +
Sbjct: 188 KDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSSSM 367
Query: 1395 ILEKEDKSGKVCQKLYRGMIGSLLY 1419
I E + ++ + Y +GSL+Y
Sbjct: 368 IPSSEAERMEMSRVPYASAVGSLMY 442
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 58.2 bits (139), Expect = 1e-08
Identities = 29/82 (35%), Positives = 49/82 (59%), Gaps = 1/82 (1%)
Frame = +3
Query: 1532 QMLWMKHQLEDYQILESN-IPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGV 1590
+ LW+ + L D +I + IYCDN +A+ L+ N + H R ++IE+ H + V G+
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 1591 LLLKFVDTDHQWADIFTKSLAE 1612
L L V + Q AD+FTK++++
Sbjct: 201 LHLLHVPSSDQVADVFTKTISQ 266
>AV766088
Length = 501
Score = 53.9 bits (128), Expect = 2e-07
Identities = 36/108 (33%), Positives = 58/108 (53%), Gaps = 1/108 (0%)
Frame = -3
Query: 738 QWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAKNVVST 797
Q +WH RLGH S + L + N L ++ +CE+C GK ++PF + N V T
Sbjct: 313 QSLWHSRLGHPSSSALRYL-RSNKFISYELLNYSP--VCESCVFGKHVRLPFVSSNNV-T 146
Query: 798 SRPLELLHIDLF-GPVKTESIGGKRYGMVIVDDYSRWTWVKFLTRKDE 844
P ++LH DL+ PV S G R+ ++ +DD++ + W L+ K +
Sbjct: 145 VMPFDILHSDLWTSPVL--SSAGHRFYVLFLDDFTDFLWTFPLSNKSQ 8
>TC19389 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (6%)
Length = 498
Score = 53.9 bits (128), Expect = 2e-07
Identities = 36/114 (31%), Positives = 60/114 (52%), Gaps = 3/114 (2%)
Frame = +3
Query: 1504 VSWASKRQSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIPIYC---DNTAAI 1560
V+W S+ Q +ALSTAEAE+I+A ++LWMK+ L++ + PI C A
Sbjct: 27 VAWPSRLQKCVALSTAEAEFIAATEACHELLWMKNFLQNAWF--HSHPILCCIVITKALF 200
Query: 1561 SLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKSLAEDR 1614
+L++ + + +RD + +L L+ + TD AD+ TKSL ++
Sbjct: 201 TLARILLFIQDPSTLMFVIIGLRDVLNSKLLELEKIHTDDDGADMMTKSLPREK 362
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 53.5 bits (127), Expect = 3e-07
Identities = 26/91 (28%), Positives = 50/91 (54%), Gaps = 1/91 (1%)
Frame = +1
Query: 1535 WMKHQLEDYQI-LESNIPIYCDNTAAISLSKNPILHSRAKHIEVKYHFIRDYVQKGVLLL 1593
W+ + L+D ++ S +YCD+ +A ++ N + H R KH+++ H +R+ +Q + L
Sbjct: 13 WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKLFHL 192
Query: 1594 KFVDTDHQWADIFTKSLAEDRFNFILKNLNM 1624
+ + Q ADI TK L F+ ++ L +
Sbjct: 193 LPISSVDQTADILTKPLESGPFSHLVSKLGV 285
>TC17418 similar to UP|Q94AX2 (Q94AX2) AT5g39790/MKM21_80, partial (20%)
Length = 739
Score = 53.1 bits (126), Expect = 3e-07
Identities = 36/99 (36%), Positives = 49/99 (49%), Gaps = 1/99 (1%)
Frame = -1
Query: 1367 QSKYTKELLKKFNMLESTVAKTPMHPTCILEKEDKSGK-VCQKLYRGMIGSLLYLTASRP 1425
Q Y ++LKKF M S T + I GK V Y+ +IGS+ YL R
Sbjct: 739 QKIYVDDILKKFKMTNSKYISTTIGGKEIEAGRRNGGKRVDSTYYKSLIGSVRYLNTVRS 560
Query: 1426 DILFSVHLCARFQSDPRETHLTAVKRILRYLKGTTNLGL 1464
DI+ V L +RF +P + H +R LRY+KGT G+
Sbjct: 559 DIVCGVGLRSRFM-EP*DCH*QGAQRSLRYIKGTLKDGI 446
>BP033602
Length = 533
Score = 52.4 bits (124), Expect = 6e-07
Identities = 30/99 (30%), Positives = 53/99 (53%)
Frame = +3
Query: 733 SVNEEQWVWHRRLGHASMRKISQLSKLNLVRGLPNLKFASDALCEACQKGKFTKVPFKAK 792
SV ++ +WH RLGH + + + L +L + + S CE+C K + P+ ++
Sbjct: 255 SVRDQIMLWHNRLGHPNFQYLRHLFP-DLFKNVN----CSSLECESCVLAKNQRAPYYSQ 419
Query: 793 NVVSTSRPLELLHIDLFGPVKTESIGGKRYGMVIVDDYS 831
SRP L+H D++GP K + GKR+ + +DD++
Sbjct: 420 PY-HASRPFYLIHSDVWGPSKITTQFGKRWFVTFIDDHT 533
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.332 0.142 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,579,087
Number of Sequences: 28460
Number of extensions: 362321
Number of successful extensions: 2282
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 2238
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2269
length of query: 1626
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1523
effective length of database: 1,966,220
effective search space: 2994553060
effective search space used: 2994553060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0243.12


