

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
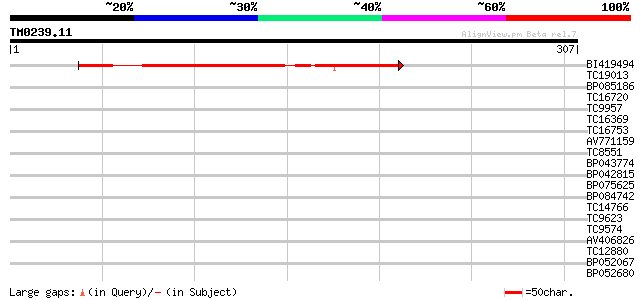
Query= TM0239.11
(307 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI419494 141 1e-34
TC19013 34 0.036
BP085186 30 0.51
TC16720 homologue to UP|Q9XH46 (Q9XH46) THA4 protein precursor, ... 30 0.67
TC9957 similar to UP|Q9FXS1 (Q9FXS1) WRKY transcription factor N... 30 0.67
TC16369 29 0.88
TC16753 similar to UP|Q9SIA7 (Q9SIA7) Legumin-like protein, part... 29 1.1
AV771159 28 2.0
TC8551 similar to PIR|T52138|T52138 eukaryotic cap-binding prote... 28 2.5
BP043774 27 3.3
BP042815 27 3.3
BP075625 27 3.3
BP084742 27 4.3
TC14766 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associ... 27 4.3
TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxid... 27 4.3
TC9574 27 4.3
AV406826 27 4.3
TC12880 UP|Q84L98 (Q84L98) Asparagine synthetase 3 , complete 27 4.3
BP052067 27 4.3
BP052680 27 4.3
>BI419494
Length = 478
Score = 141 bits (356), Expect = 1e-34
Identities = 89/180 (49%), Positives = 115/180 (63%), Gaps = 4/180 (2%)
Frame = -2
Query: 38 PLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPC-ILYVLVSIVSIA 96
PLSF LL++LSAA+ YLQ S LF L + PC ILY+LV ++++A
Sbjct: 477 PLSFLLLSNLSAAQFYLQR---------------SNLFPCFLSLAPCNILYILVFLLTLA 343
Query: 97 TLINGLMGKITLLNDSSTSPVLQPS-LYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSS 155
+LI+ L GK+T+ N S +S S L AWI+LC FQ CVGLG E SI AG++ S
Sbjct: 342 SLIHCLTGKLTVFNVSPSSTSSSTSRLCTAWIILCTFQVCVGLGTEGSIVAGVY-----S 178
Query: 156 SSSSSSFGESAAERSLLSK--VFFLLGLHETTQVWYRVVVRPVVDDTVFGGAKKEKWIER 213
S SSFG ++SLL + + FLLGLHETT+ W +VVVRPVVDDTVFG A++E+W R
Sbjct: 177 DDSISSFG--VEKKSLLRRGVLGFLLGLHETTRFWSKVVVRPVVDDTVFGVAREERWPTR 4
>TC19013
Length = 378
Score = 33.9 bits (76), Expect = 0.036
Identities = 27/99 (27%), Positives = 49/99 (49%), Gaps = 6/99 (6%)
Frame = +3
Query: 30 ITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINPCILYVL 89
++L+ L + F++ + HY F L HS ++++P F+FT+ I Y+
Sbjct: 18 LSLIKLCVSF*FYIFS------HYRSNF*LHTHSI*RRKKPSCFVFTVFFSIRIRFSYIS 179
Query: 90 VSIVSIATLI-----NGLMGKITL-LNDSSTSPVLQPSL 122
S+ SI T++ G MG + N ++SP+ +P L
Sbjct: 180 FSLTSILTVMANRWWTGSMGLGGVDDNPDNSSPMGKPDL 296
>BP085186
Length = 388
Score = 30.0 bits (66), Expect = 0.51
Identities = 18/52 (34%), Positives = 27/52 (51%)
Frame = +3
Query: 125 AWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVF 176
+W+L + F G +S ++ +SSSSSSSSF +A L+S F
Sbjct: 150 SWVLTFSSSFFSSSGSASSSSSSSSSSSSSSSSSSSSFPSPSALSGLISLPF 305
>TC16720 homologue to UP|Q9XH46 (Q9XH46) THA4 protein precursor, partial
(42%)
Length = 567
Score = 29.6 bits (65), Expect = 0.67
Identities = 19/51 (37%), Positives = 26/51 (50%), Gaps = 3/51 (5%)
Frame = -2
Query: 57 FTLFLHSPQQQQQPFS---FLFTLALQINPCILYVLVSIVSIATLINGLMG 104
F+LF HS + + P + FLF L L+ C+L L S AT L+G
Sbjct: 422 FSLFAHSKRFLRLPSNGIRFLFKLGLKFLCCLLEALESFADAATDFRQLLG 270
>TC9957 similar to UP|Q9FXS1 (Q9FXS1) WRKY transcription factor Nt-SubD48,
partial (16%)
Length = 653
Score = 29.6 bits (65), Expect = 0.67
Identities = 21/58 (36%), Positives = 28/58 (48%), Gaps = 12/58 (20%)
Frame = -3
Query: 188 WYRVVVRPVVDDTVFGGAK---------KEKWIERVAVAA---SLGVLWWWRLREEVE 233
W V V PV ++ GA+ ++ ++ RVAV A S G LWWW R VE
Sbjct: 630 WCLVSVSPVWEEMKELGAEV*DEKLRVLEDSFLSRVAVVAAPISGGSLWWWWWRVAVE 457
>TC16369
Length = 751
Score = 29.3 bits (64), Expect = 0.88
Identities = 17/45 (37%), Positives = 26/45 (57%)
Frame = -2
Query: 4 WQQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLS 48
W++ K+ F + P+ KA + ITLL L P FFLL+++S
Sbjct: 684 WRKGMKQNFFSFLSRPSAKALTTI--ITLLFLFTPFFFFLLSTIS 556
>TC16753 similar to UP|Q9SIA7 (Q9SIA7) Legumin-like protein, partial (42%)
Length = 534
Score = 28.9 bits (63), Expect = 1.1
Identities = 13/36 (36%), Positives = 23/36 (63%), Gaps = 1/36 (2%)
Frame = +1
Query: 215 AVAASLGVLWWWRLREEVESLVV-MAEAKKSEQFGE 249
A+A LGV+ WW +E+ E +V+ + + K+ + GE
Sbjct: 343 ALALPLGVVTWWYNKEDTELVVLCLGDTSKAHKAGE 450
>AV771159
Length = 486
Score = 28.1 bits (61), Expect = 2.0
Identities = 17/40 (42%), Positives = 22/40 (54%), Gaps = 6/40 (15%)
Frame = -1
Query: 11 FFSNHGGAPT------PKATFHLLTITLLSLLLPLSFFLL 44
FFS G T P +TFHLL L+SLL+ L +L+
Sbjct: 168 FFSLKGAPSTMVLLQLPLSTFHLLVSPLVSLLIQLISYLI 49
>TC8551 similar to PIR|T52138|T52138 eukaryotic cap-binding protein
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (71%)
Length = 1167
Score = 27.7 bits (60), Expect = 2.5
Identities = 14/36 (38%), Positives = 22/36 (60%)
Frame = +2
Query: 88 VLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLY 123
VLV++ + TL+ G+ G + LLN S T+ + LY
Sbjct: 767 VLVNVPAAGTLMLGVHGLVCLLNSSRTATLTGSFLY 874
>BP043774
Length = 439
Score = 27.3 bits (59), Expect = 3.3
Identities = 15/26 (57%), Positives = 17/26 (64%)
Frame = +1
Query: 145 AAGIFEFDNSSSSSSSSFGESAAERS 170
A GI E SSSSSSSS AAE++
Sbjct: 331 AVGIHESPPSSSSSSSSLATGAAEQA 408
>BP042815
Length = 526
Score = 27.3 bits (59), Expect = 3.3
Identities = 23/68 (33%), Positives = 33/68 (47%), Gaps = 5/68 (7%)
Frame = -2
Query: 21 PKATFHLLT-ITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQ----QQQQPFSFLF 75
P + F++ T I L+LL+P S L +L H + F+H+PQ Q Q FLF
Sbjct: 372 PISDFNITTQIIHLNLLIPDST-LNITLQVTNHKINLIKKFIHAPQFLKGQFQILLLFLF 196
Query: 76 TLALQINP 83
L +P
Sbjct: 195 PLLTSTHP 172
>BP075625
Length = 488
Score = 27.3 bits (59), Expect = 3.3
Identities = 22/86 (25%), Positives = 38/86 (43%), Gaps = 17/86 (19%)
Frame = -1
Query: 1 MGRWQQKAKEFFSNHGGAPTPKATFHLLTITLL---------------SLLLPLSFFLLA 45
+GRW + KE F G + + L + LL L + SFF+L
Sbjct: 263 LGRWYEFYKETFFRLGNS-VARHQMQLWKMVLLHFYRVRCFDGVI*HPKLHISCSFFVLH 87
Query: 46 SL--SAAKHYLQTFTLFLHSPQQQQQ 69
++ S +H+L TLF+H ++ ++
Sbjct: 86 NIT*SGCRHHLNRATLFVHCEEKXKK 9
>BP084742
Length = 459
Score = 26.9 bits (58), Expect = 4.3
Identities = 16/37 (43%), Positives = 21/37 (56%)
Frame = +3
Query: 39 LSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLF 75
L+F ASL ++H+LQ FL Q + QPF LF
Sbjct: 354 LNFDFSASLYLSQHFLQLHPSFL--IQLKHQPFELLF 458
>TC14766 similar to UP|Q7XXR8 (Q7XXR8) Nascent polypeptide associated
complex alpha chain, partial (58%)
Length = 600
Score = 26.9 bits (58), Expect = 4.3
Identities = 19/57 (33%), Positives = 26/57 (45%)
Frame = -1
Query: 120 PSLYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVF 176
PSL IA L + C L + + +SSSSSSSS S++ S +F
Sbjct: 279 PSLSIALRLFFSLLVCFDLPDASPSCPSMLSSSSSSSSSSSSSSSSSSTTGSSSWIF 109
>TC9623 homologue to UP|Q8H1K8 (Q8H1K8) Stromal ascorbate peroxidase,
partial (42%)
Length = 640
Score = 26.9 bits (58), Expect = 4.3
Identities = 18/58 (31%), Positives = 26/58 (44%), Gaps = 2/58 (3%)
Frame = +2
Query: 3 RWQQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSF--FLLASLSAAKHYLQTFT 58
RW++++ + F H P LL LL LLL L FLL L+ + F+
Sbjct: 65 RWRRRSHDSFHRHQSHPLTLFLLILLPFLLLILLLLLLLFPFLLPMLALLSSHFSPFS 238
>TC9574
Length = 980
Score = 26.9 bits (58), Expect = 4.3
Identities = 22/60 (36%), Positives = 29/60 (47%)
Frame = -1
Query: 114 TSPVLQPSLYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLS 173
T P+ +P I WIL + C + S + I +SSSSS S +AAE LLS
Sbjct: 554 TLPIDEP---ITWILFTSKLTC---SLYISFKSTISARSSSSSSSRSPINSNAAECLLLS 393
>AV406826
Length = 425
Score = 26.9 bits (58), Expect = 4.3
Identities = 16/39 (41%), Positives = 18/39 (46%)
Frame = -3
Query: 214 VAVAASLGVLWWWRLREEVESLVVMAEAKKSEQFGELEL 252
VAVA L WWWRL S V EA + G + L
Sbjct: 366 VAVALDLEA-WWWRLESS*PSAVAEGEAPPLHERGVVGL 253
>TC12880 UP|Q84L98 (Q84L98) Asparagine synthetase 3 , complete
Length = 2392
Score = 26.9 bits (58), Expect = 4.3
Identities = 17/51 (33%), Positives = 24/51 (46%), Gaps = 9/51 (17%)
Frame = +2
Query: 204 GAKKEKWIERVAVAASLGVLWWWRLREE---------VESLVVMAEAKKSE 245
G + EKWI R A L WR +E+ ++SL A+AK S+
Sbjct: 1415 GGRIEKWIVRKAFEGYLPPAILWRQKEQFSDGVGYSWIDSLKAFADAKVSD 1567
>BP052067
Length = 501
Score = 26.9 bits (58), Expect = 4.3
Identities = 18/53 (33%), Positives = 21/53 (38%)
Frame = -2
Query: 207 KEKWIERVAVAASLGVLWWWRLREEVESLVVMAEAKKSEQFGELELKDFIGWW 259
KE W E A+ G WWW E E + + E FG E F WW
Sbjct: 212 KEWWWESEAMRRRCGW*WWWW*E*E*E***QLRRHHRCEAFG--EDASFWAWW 60
>BP052680
Length = 463
Score = 26.9 bits (58), Expect = 4.3
Identities = 16/60 (26%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Frame = +2
Query: 75 FTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSS-TSPVLQPSLYIAWILLCAFQ 133
++L +N C + + + SI + LMG +T ND++ T+ L+ +L + +C Q
Sbjct: 8 YSLTR*MNGCCKSIEIQVTSILNANSNLMGTVTGTNDNTITNK*LKKTLKVKHAKICTEQ 187
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,446,896
Number of Sequences: 28460
Number of extensions: 110647
Number of successful extensions: 1445
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 1289
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1381
length of query: 307
length of database: 4,897,600
effective HSP length: 90
effective length of query: 217
effective length of database: 2,336,200
effective search space: 506955400
effective search space used: 506955400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0239.11


