

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0234.6
(1379 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
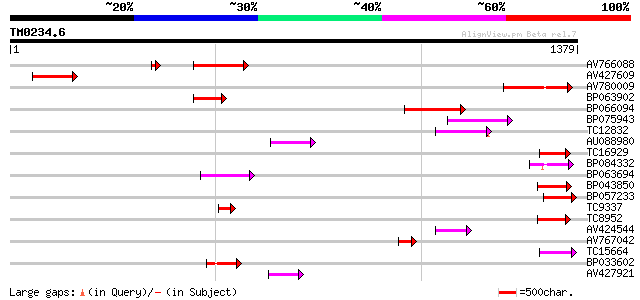
Sequences producing significant alignments: (bits) Value
AV766088 263 1e-70
AV427609 211 6e-55
AV780009 163 2e-40
BP063902 119 3e-27
BP066094 115 5e-26
BP075943 101 7e-22
TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-rela... 83 3e-16
AU088980 82 6e-16
TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%) 82 8e-16
BP084332 81 1e-15
BP063694 79 5e-15
BP043850 76 4e-14
BP057233 75 7e-14
TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B pre... 75 7e-14
TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%) 74 2e-13
AV424544 70 2e-12
AV767042 69 7e-12
TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, p... 68 9e-12
BP033602 68 1e-11
AV427921 65 6e-11
>AV766088
Length = 501
Score = 263 bits (673), Expect = 1e-70
Identities = 126/132 (95%), Positives = 128/132 (96%)
Frame = -3
Query: 448 QTGIPLMRCNSPGDLYPVTPSFPFAGLAQSLWHSRLGHPSSSALQSLRSNKFISYEHLNS 507
QTGIPLMRCNSPGDLYPVT SFPFAG+AQSLWHSRLGHPSSSAL+ LRSNKFISYE LN
Sbjct: 397 QTGIPLMRCNSPGDLYPVTLSFPFAGIAQSLWHSRLGHPSSSALRYLRSNKFISYELLNY 218
Query: 508 SPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFL 567
SPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFL
Sbjct: 217 SPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLSSAGHRFYVLFLDDFTDFL 38
Query: 568 WTFPLSNKSQVF 579
WTFPLSNKSQVF
Sbjct: 37 WTFPLSNKSQVF 2
Score = 46.2 bits (108), Expect = 4e-05
Identities = 21/24 (87%), Positives = 21/24 (87%)
Frame = -3
Query: 344 CPLILPLPCTP*RLLLQIAPGNLT 367
CPLILPLPCTP*RLLLQIAP T
Sbjct: 499 CPLILPLPCTP*RLLLQIAPSTWT 428
>AV427609
Length = 332
Score = 211 bits (537), Expect = 6e-55
Identities = 109/110 (99%), Positives = 110/110 (99%)
Frame = +2
Query: 55 AFMLVLTEFCITLSLLRTNHLHQ*LILPMSSGPLWMLRSFSGFIPPSLLT**LLSWSLAP 114
AFMLVLTEFCITLSLLRTNHLH+*LILPMSSGPLWMLRSFSGFIPPSLLT**LLSWSLAP
Sbjct: 2 AFMLVLTEFCITLSLLRTNHLHR*LILPMSSGPLWMLRSFSGFIPPSLLT**LLSWSLAP 181
Query: 115 LPSLLGRI*RICFRTIRTLVQSPSSRSSLLFAWRHSRPSLPTASV*RRFL 164
LPSLLGRI*RICFRTIRTLVQSPSSRSSLLFAWRHSRPSLPTASV*RRFL
Sbjct: 182 LPSLLGRI*RICFRTIRTLVQSPSSRSSLLFAWRHSRPSLPTASV*RRFL 331
>AV780009
Length = 529
Score = 163 bits (412), Expect = 2e-40
Identities = 84/171 (49%), Positives = 113/171 (65%), Gaps = 2/171 (1%)
Frame = -1
Query: 1201 ALKRILRYVQGTLQLGLHLYPSPIEKLISYTDADWGGCLDTRRSTSGYCVFLGDNLISWS 1260
A R+LRYV+G GL KL +Y+D+DW GC DTRRS +GY +FLG +LISW
Sbjct: 526 AATRVLRYVKGAPAQGLFFSADSPLKLQAYSDSDWAGCPDTRRSVTGYSIFLGTSLISWR 347
Query: 1261 SKRQPTLSRSSAEAEYRGVANVVSESCWLRNL--LLELHFPLS*ATLVYCDNVSAIYLSG 1318
+K+Q T+SRSS+EAEYR +A V E WL L L+L+ PL ++CDN SA++++
Sbjct: 346 TKKQTTVSRSSSEAEYRALAATVCEVQWLSYLFQFLKLNVPL--PVPLFCDNQSALHIAH 173
Query: 1319 NPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLF 1369
NP H+RTKHIE+D H VR K+ G +L + + HQ+ADIFTK P + F
Sbjct: 172 NPTFHERTKHIELDCHVVRAKLQAGLIHLLPISTHHQLADIFTKPCPHLCF 20
>BP063902
Length = 494
Score = 119 bits (298), Expect = 3e-27
Identities = 57/80 (71%), Positives = 64/80 (79%)
Frame = -3
Query: 448 QTGIPLMRCNSPGDLYPVTPSFPFAGLAQSLWHSRLGHPSSSALQSLRSNKFISYEHLNS 507
+TG+PL+RCNS GDLYPVT S PFAGLA S+WH RLGHP+SSAL LR+NK I E S
Sbjct: 273 KTGMPLLRCNSLGDLYPVTRSSPFAGLASSVWHHRLGHPASSALNHLRNNKLIFCEPSRS 94
Query: 508 SPVCESCVFGKHVRLPFVSS 527
S VC+SCV GKHVRLPF SS
Sbjct: 93 SSVCDSCVLGKHVRLPFSSS 34
Score = 32.3 bits (72), Expect = 0.53
Identities = 27/62 (43%), Positives = 31/62 (49%), Gaps = 3/62 (4%)
Frame = -3
Query: 309 HAHILL---LSGLVLLLLLGNLAFWARVLRLMLLRPPPCPLILPLPCTP*RLLLQIAPGN 365
HA+++L LS V + N AF RVLR L PLI LPCTP LLQ G
Sbjct: 489 HAYLVLTPHLSXRVPWVFRSNPAF*VRVLRPTPLLLLQHPLISLLPCTPCP*LLQTPRGT 310
Query: 366 LT 367
T
Sbjct: 309 WT 304
>BP066094
Length = 532
Score = 115 bits (288), Expect = 5e-26
Identities = 57/147 (38%), Positives = 92/147 (61%)
Frame = +2
Query: 961 KPATIRTVLTIALSRSWPIHQLDVQNAFLHGDLHETVYMHQPLGFRDPNHPDYVCRLRKS 1020
K IR +++ +++ + +HQ++V++AFL+G + E VY+HQP G D + D++ +L+KS
Sbjct: 89 KTEAIRLLISFSVNHNIILHQMNVKSAFLNGYISEEVYVHQPPGXEDEKNSDHIFKLKKS 268
Query: 1021 LYGLKQAPRAWYQCFADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHD 1080
LYGLKQAPRAWY+ + ++ D +LF D+ + +YVDDII S++
Sbjct: 269 LYGLKQAPRAWYERLSSFLLENEXVRGKVDTTLFCKTYKDDILIVQIYVDDIIFGSANPS 448
Query: 1081 LRKSIMALLASEFAMKDLGPLSYFLGI 1107
L K ++ +EF M+ +G L YFLGI
Sbjct: 449 LCKEFSEMMQAEFEMRMMGELKYFLGI 529
>BP075943
Length = 547
Score = 101 bits (252), Expect = 7e-22
Identities = 61/159 (38%), Positives = 86/159 (53%), Gaps = 1/159 (0%)
Frame = -3
Query: 1065 LLLYVDDIILISSSHDLRKSIMALLASEFAMKDLGPLSYFLGIAVTRHAGGLFLSQSTYA 1124
LLLYVDD++L + + + + L +F +KDLG YFLG+ + R G+ L+Q YA
Sbjct: 521 LLLYVDDVLLAGNDMHEIQLVKSSLHDQFRIKDLGEAKYFLGLEIARSTSGIVLNQRKYA 342
Query: 1125 RDIIARAGMASCHPSATPVDTK-QKLSTSAGTPCDDPTLYRSLVGALQYLTFTRPDISYA 1183
+I+ +G P T Q L T+ GTP D YR +VG L YL TRPDI++A
Sbjct: 341 LQLISDSGHFGFQPRFYSHGTNSQTLGTNTGTPLTDIGSYRRIVGRLLYLNTTRPDITFA 162
Query: 1184 VQQVCLHMHAPRTEHMLALKRILRYVQGTLQLGLHLYPS 1222
V Q+ + AP H L L+Y+ G+ GL YP+
Sbjct: 161 VNQLSQFLSAPTDIHEQQLTGFLKYL*GSPGSGL-FYPA 48
>TC12832 weakly similar to UP|POLX_TOBAC (P10978) Retrovirus-related Pol
polyprotein from transposon TNT 1-94 [Contains: Protease
; Reverse transcriptase ; Endonuclease] , partial (9%)
Length = 747
Score = 83.2 bits (204), Expect = 3e-16
Identities = 49/147 (33%), Positives = 79/147 (53%), Gaps = 9/147 (6%)
Frame = +2
Query: 1035 FADYVSTIGFQHSTSDHSLFIYR-RGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEF 1093
F ++ ++G+ +SDH + R +D LLLYVDD++++ + D + + A LA EF
Sbjct: 2 FDSFIMSLGYNRLSSDHCTYHKRFDDNDFIILLLYVDDMLVVGPNKDRVQELKAQLAREF 181
Query: 1094 AMKDLGPLSYFLGIAVTRHAGG--LFLSQSTYARDIIARAGMASCHPSATPVDTKQKLST 1151
MKDLGP + LG+ + R ++LSQ Y + ++ R M C+P +TP+ KLS+
Sbjct: 182 DMKDLGPANKILGMQIHRDRKDRRIWLSQKNYLQKVLRRFNMQDCNPISTPLPVNYKLSS 361
Query: 1152 SAGTPCDDPTL------YRSLVGALQY 1172
S + + Y S VG+L Y
Sbjct: 362 SMIPSSEAERMEMSRVPYASAVGSLMY 442
>AU088980
Length = 360
Score = 82.0 bits (201), Expect = 6e-16
Identities = 43/110 (39%), Positives = 57/110 (51%)
Frame = +2
Query: 635 NGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQLLYRR 694
N ERK R I N+ R L+ H+ +P + W A++ A YL+N +P L P Q L
Sbjct: 2 NSIVERKHRHILNVTRALMFHSYLPKNLWTFAVKHAAYLINXLPSPLLKGQCPYQFLNND 181
Query: 695 DPSYTHLRVFGCLCYPLVPSSTINKLQPRSTPCVFLGYPLHHRGYKCFDL 744
P+ L+VFG LCY + KL PR+ CVFLG+ GY DL
Sbjct: 182 SPTLLDLKVFGTLCYSSTLTHNXQKLDPRARKCVFLGFKTGTXGYIVMDL 331
>TC16929 weakly similar to UP|Q9LFY6 (Q9LFY6) T7N9.5, partial (4%)
Length = 553
Score = 81.6 bits (200), Expect = 8e-16
Identities = 38/77 (49%), Positives = 51/77 (65%)
Frame = +1
Query: 1288 WLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARV 1347
WL LL +L P LVYCDN SA +++ NPV H+RTKHIE+D H VRE++ +G +
Sbjct: 4 WLTYLLQDLKVPFEQPALVYCDNNSARHIAANPVFHERTKHIEIDCHIVRERIQKGLIHL 183
Query: 1348 LHVPSRHQIADIFTKGL 1364
L + S +ADI+TK L
Sbjct: 184 LPISSSEPLADIYTKAL 234
>BP084332
Length = 368
Score = 81.3 bits (199), Expect = 1e-15
Identities = 46/112 (41%), Positives = 65/112 (57%), Gaps = 5/112 (4%)
Frame = +2
Query: 1264 QPTLSRSSAEAEYRGVANVVSESCWLRNLL-----LELHFPLS*ATLVYCDNVSAIYLSG 1318
Q T++ S+AEAEY A ++ W+++ L LE + P +YCDN +AI LS
Sbjct: 32 QSTIALSTAEAEYISAAICSTQMLWMKHQLEDYQILESNIP------IYCDNTAAISLSK 193
Query: 1319 NPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIADIFTKGLPRVLFD 1370
NP+ H R KHIE+ HF+R+ V +G + V + HQ ADIFTK L F+
Sbjct: 194 NPILHSRAKHIEVKYHFIRDYVQKGVLLLKFVDTDHQWADIFTKPLAEDRFN 349
>BP063694
Length = 511
Score = 79.0 bits (193), Expect = 5e-15
Identities = 43/132 (32%), Positives = 65/132 (48%), Gaps = 2/132 (1%)
Frame = -2
Query: 465 VTPSFPFAGLAQSLWHSRLGHPSSSALQSLRSNKFISYEHLNSSPVC-ESCVFGKHVRLP 523
+T S P + LWHSRLGH + ++ L + + + P+C SC K RL
Sbjct: 399 LTTSSPLPRASFELWHSRLGHVNFDIIKQLHKHGCLDVSSILPKPICCTSCQMAKSKRLV 220
Query: 524 FVSSNNVTVMPFDILHSDL-WTSPVLSSAGHRFYVLFLDDFTDFLWTFPLSNKSQVFEMF 582
F +N D++H DL SPV S G ++V+F+DDF+ F W +PL KS ++
Sbjct: 219 FHDNNKRASAVLDLIHCDLRGPSPVASIDGFSYFVIFVDDFSRFTWFYPLKRKSDFSDVL 40
Query: 583 ISLSNQIRTHFS 594
+ + FS
Sbjct: 39 LRFKVFMENRFS 4
>BP043850
Length = 515
Score = 75.9 bits (185), Expect = 4e-14
Identities = 42/83 (50%), Positives = 51/83 (60%)
Frame = -1
Query: 1284 SESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARG 1343
SE WLR LL EL F S T ++ DN SAI ++ NPV H+ T+HIE+D H VRE R
Sbjct: 515 SEIIWLRGLLSELGFLQSQPTPLHADNTSAIQIAANPVYHEWTRHIEVDCHSVREAYDRR 336
Query: 1344 QARVLHVPSRHQIADIFTKGLPR 1366
+ HV + QIADI TK L R
Sbjct: 335 VITLPHVSTSVQIADILTKSLTR 267
>BP057233
Length = 473
Score = 75.1 bits (183), Expect = 7e-14
Identities = 34/80 (42%), Positives = 51/80 (63%)
Frame = -2
Query: 1299 PLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARVLHVPSRHQIAD 1358
P+ +++CDN+SA L+ NPV H R+KHIE+D+H++R++V + + V +VP+ QIAD
Sbjct: 466 PMPRKPILWCDNLSAKALASNPVLHARSKHIEIDVHYIRDQVLQNEVVVAYVPTTDQIAD 287
Query: 1359 IFTKGLPRVLFDDFRSSLSV 1378
TK L F R L V
Sbjct: 286 CLTKPLSHTRFSQLRDKLGV 227
>TC9337 similar to UP|PME_PRUPE (Q43062) Pectinesterase PPE8B precursor
(Pectin methylesterase) (PE) , partial (54%)
Length = 1145
Score = 75.1 bits (183), Expect = 7e-14
Identities = 33/43 (76%), Positives = 37/43 (85%)
Frame = +3
Query: 507 SSPVCESCVFGKHVRLPFVSSNNVTVMPFDILHSDLWTSPVLS 549
S+ VC+SCV GKHVRLPF SS +T+ PFDILHSDLWTSPVLS
Sbjct: 1017 SNSVCDSCVLGKHVRLPFSSSETITLRPFDILHSDLWTSPVLS 1145
>TC8952 similar to UP|O82607 (O82607) T2L5.9 protein, partial (3%)
Length = 550
Score = 73.6 bits (179), Expect = 2e-13
Identities = 36/78 (46%), Positives = 50/78 (63%)
Frame = +3
Query: 1285 ESCWLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQ 1344
E+ WL L +L ++YCDN SA++L+ N V H+RT++IE+D H V KV G
Sbjct: 21 EALWLTYALADLRIASLLLVVIYCDNRSALHLAANSVFHKRTENIEIDCHIV*VKVLFGI 200
Query: 1345 ARVLHVPSRHQIADIFTK 1362
+LHVPS Q+AD+FTK
Sbjct: 201 LHLLHVPSSDQVADVFTK 254
>AV424544
Length = 276
Score = 70.1 bits (170), Expect = 2e-12
Identities = 36/88 (40%), Positives = 52/88 (58%)
Frame = +3
Query: 1036 ADYVSTIGFQHSTSDHSLFIYRRGSDMAYLLLYVDDIILISSSHDLRKSIMALLASEFAM 1095
+ Y+ +G+ S DHSLF R + +L+YVDD+IL + + + + L +F +
Sbjct: 12 SSYLHILGYIQSAHDHSLFTKFRDASFTVILVYVDDLILAGNDLNEIQCVKNKLDIQFRI 191
Query: 1096 KDLGPLSYFLGIAVTRHAGGLFLSQSTY 1123
KDLG L YFLG+ V R + GLFLSQ Y
Sbjct: 192 KDLGTLKYFLGLEVARSSCGLFLSQRKY 275
>AV767042
Length = 444
Score = 68.6 bits (166), Expect = 7e-12
Identities = 33/44 (75%), Positives = 38/44 (86%)
Frame = -3
Query: 946 QIAGVDCDETFSHVVKPATIRTVLTIALSRSWPIHQLDVQNAFL 989
Q AGVDC++TFS VVKPA IRTV TIALSRSWPI QL+VQN ++
Sbjct: 349 QTAGVDCNKTFSPVVKPAPIRTVFTIALSRSWPIPQLNVQNLWI 218
>TC15664 weakly similar to UP|O81617 (O81617) F8M12.17 protein, partial (4%)
Length = 670
Score = 68.2 bits (165), Expect = 9e-12
Identities = 37/91 (40%), Positives = 52/91 (56%)
Frame = +1
Query: 1288 WLRNLLLELHFPLS*ATLVYCDNVSAIYLSGNPVQHQRTKHIEMDIHFVREKVARGQARV 1347
WL +L +L P +L+YCD+ SA +++ N V H+RTKH+++D H VREK+ +
Sbjct: 13 WLTYILQDLRVPFISPSLLYCDSQSARHIATNAVFHERTKHLDIDCHVVREKLQAKLFHL 192
Query: 1348 LHVPSRHQIADIFTKGLPRVLFDDFRSSLSV 1378
L + S Q ADI TK L F S L V
Sbjct: 193 LPISSVDQTADILTKPLESGPFSHLVSKLGV 285
>BP033602
Length = 533
Score = 67.8 bits (164), Expect = 1e-11
Identities = 36/89 (40%), Positives = 54/89 (60%), Gaps = 2/89 (2%)
Frame = +3
Query: 478 LWHSRLGHPSSSALQSLRSNKFISYEHLNSSPV-CESCVFGKHVRLPFVSSNNVTVMPFD 536
LWH+RLGHP+ L+ L + F +++N S + CESCV K+ R P+ S PF
Sbjct: 276 LWHNRLGHPNFQYLRHLFPDLF---KNVNCSSLECESCVLAKNQRAPYYSQPYHASRPFY 446
Query: 537 ILHSDLW-TSPVLSSAGHRFYVLFLDDFT 564
++HSD+W S + + G R++V F+DD T
Sbjct: 447 LIHSDVWGPSKITTQFGKRWFVTFIDDHT 533
>AV427921
Length = 284
Score = 65.5 bits (158), Expect = 6e-11
Identities = 33/84 (39%), Positives = 48/84 (56%)
Frame = +1
Query: 630 HTSSQNGKAERKIRTINNMIRTLLAHASVPPSFWHHALQMATYLLNIIPRKNLSNLSPTQ 689
+T QNG +ERK RTI NM+R+LL + +P SF A+ + ++LN P + N+ P +
Sbjct: 19 YTPQQNGVSERKNRTILNMVRSLLTMSGLPKSFLPEAVMWSLHILNRSPTLVVQNMMPEE 198
Query: 690 LLYRRDPSYTHLRVFGCLCYPLVP 713
R P+ H R+ CL Y VP
Sbjct: 199 AWSGRQPAVDHFRISRCLAYAHVP 270
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.329 0.141 0.453
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,846,220
Number of Sequences: 28460
Number of extensions: 597363
Number of successful extensions: 13103
Number of sequences better than 10.0: 973
Number of HSP's better than 10.0 without gapping: 6722
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9949
length of query: 1379
length of database: 4,897,600
effective HSP length: 101
effective length of query: 1278
effective length of database: 2,023,140
effective search space: 2585572920
effective search space used: 2585572920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0234.6


