

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0234.12
(607 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
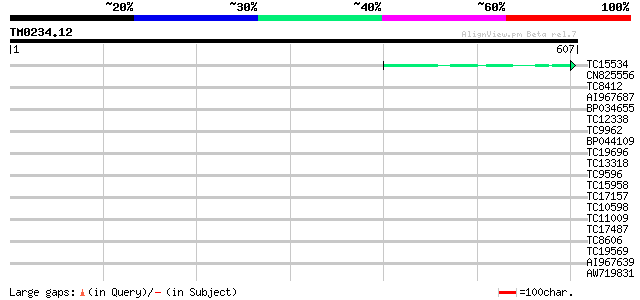
TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28... 42 4e-04
CN825556 35 0.027
TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromos... 35 0.045
AI967687 34 0.059
BP034655 34 0.078
TC12338 similar to GB|AAP13422.1|30023778|BT006314 At2g01660 {Ar... 33 0.13
TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial... 33 0.13
BP044109 32 0.23
TC19696 similar to UP|Q9ZQT8 (Q9ZQT8) WREBP-2 protein, partial (... 32 0.29
TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (... 32 0.29
TC9596 similar to UP|AAR24722 (AAR24722) At5g49400, partial (66%) 31 0.50
TC15958 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Ar... 31 0.66
TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly pr... 31 0.66
TC10598 weakly similar to GB|AAM78036.1|21928015|AY125526 At2g01... 31 0.66
TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Ar... 30 1.1
TC17487 UP|CAE45597 (CAE45597) SAR DNA-binding protein-like prot... 30 1.1
TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%) 30 1.1
TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor k... 30 1.1
AI967639 30 1.1
AW719831 30 1.5
>TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28040
{Arabidopsis thaliana;}, partial (8%)
Length = 937
Score = 41.6 bits (96), Expect = 4e-04
Identities = 47/205 (22%), Positives = 70/205 (33%)
Frame = -3
Query: 401 PSPERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDH 460
PS +P + + Q+Q S + SSSE E++D PL++ + EEEE ++
Sbjct: 923 PSAPKP*SQITMAQKQKRPSPLEDPPTASSSSESEEEDDQPLSQVKHAAAEEEELSSEE- 747
Query: 461 EIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRER 520
E S EE D P P Q + E S Q +
Sbjct: 746 -----------EGSSEEEEEDETTAPAPPAATTSHTKPPQPE-------PESDSATQSDS 621
Query: 521 RSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPI 580
S SD A +AS +P + P++T TQT
Sbjct: 620 ESDTDSDQAPSASAPTPNP-----------------------KVKPLATKPMDQTQTH-- 516
Query: 581 QPETQPQTQAEPQAPKSPIQTATTA 605
+P+ QP A K ++ A
Sbjct: 515 KPKAQPSPAPAKSAAKRAAESNANA 441
>CN825556
Length = 655
Score = 35.4 bits (80), Expect = 0.027
Identities = 18/43 (41%), Positives = 22/43 (50%)
Frame = +1
Query: 556 ALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSP 598
A V+ P QP+P SLP QP+ QPQ Q +PQ P
Sbjct: 415 APVLPPPQPVPPPPSLPQPQPQPQPQPQPQPQPQPQPQPQPQP 543
>TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromosome 3, P1
clone: MSJ11, partial (39%)
Length = 1403
Score = 34.7 bits (78), Expect = 0.045
Identities = 39/163 (23%), Positives = 61/163 (36%), Gaps = 5/163 (3%)
Frame = +3
Query: 321 WSKKDNPACILEYVRMLRRQGDPITLTEFVNTLPESPPKLATRKSRRATSSKPSDPKGKG 380
W KK P ++ G P E V T P + ++ + + P P
Sbjct: 147 WQKKFFP----------KKSGTP-KKNEIVFTAPTGEEITSKKQLEQYLKAHPGGPAASE 293
Query: 381 I---LKEEPAKKKAASKTVIIREPSPER--PLKRTSAQQQQMSDSESSEDTWKDSSSEET 435
E P + S+ V + P+PE P KR D+ E+ EET
Sbjct: 294 FDWGTGETPRRSARISEKVKVT-PTPESDPPKKRGKRSSASKKDASGEEE------KEET 452
Query: 436 EDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEE 478
+D +M A K ++E +DE + N + EA+DA +
Sbjct: 453 KDVEMQEAGETKDAVKENQDENKAEDTDVNQDEKKAEATDANQ 581
>AI967687
Length = 460
Score = 34.3 bits (77), Expect = 0.059
Identities = 16/25 (64%), Positives = 20/25 (80%)
Frame = +2
Query: 1 MGKSRKPKANVSASTTQTAKDQEKQ 25
MGKSRK K NV+AS+ T ++QEKQ
Sbjct: 386 MGKSRKSKTNVTASSKLTDEEQEKQ 460
>BP034655
Length = 517
Score = 33.9 bits (76), Expect = 0.078
Identities = 18/72 (25%), Positives = 37/72 (51%)
Frame = +1
Query: 417 MSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDA 476
+SD + E+ ++ EE E+E+ + +EE+DE+D+ E V R + +
Sbjct: 130 LSDDDEEEEEEEEEEEEEEEEEEEEEEEEE----DEEDDEEDEDEDYIPVAPSHRVSPNV 297
Query: 477 EESTDSDAIPVV 488
++ SD++ V+
Sbjct: 298 HQNPTSDSVEVI 333
Score = 30.8 bits (68), Expect = 0.66
Identities = 19/67 (28%), Positives = 29/67 (42%)
Frame = +1
Query: 451 EEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQAS 510
EEEE+E+++ E + + + D EE D D IPV ++ + E
Sbjct: 157 EEEEEEEEEEEEEEEEEEEEEDEEDDEEDEDEDYIPVAPSHRVSPNVHQNPTSDSVEVIL 336
Query: 511 EPTSEVQ 517
P SE Q
Sbjct: 337 IPDSEQQ 357
>TC12338 similar to GB|AAP13422.1|30023778|BT006314 At2g01660 {Arabidopsis
thaliana;}, partial (26%)
Length = 903
Score = 33.1 bits (74), Expect = 0.13
Identities = 19/90 (21%), Positives = 37/90 (41%), Gaps = 3/90 (3%)
Frame = -2
Query: 520 RRSKRTSDPARAASLTRTSPIRSLSKV---ITESINSDSALVVIPEQPMPISTSLPTSTQ 576
+ +S P + +L+ +P + + S+++ P P+ST +
Sbjct: 839 KHGSSSSSPLHSHTLSHHNPFNFIINRHFHLRRSLSAQEHAGFRPRSTKPVSTRSSPPSS 660
Query: 577 TEPIQPETQPQTQAEPQAPKSPIQTATTAI 606
T+P P T P AP +P +T+A+
Sbjct: 659 TQPPSPTTTTSPSRPPPAPATPSTASTSAV 570
>TC9962 similar to UP|Q8L7Z6 (Q8L7Z6) AT3g54680/T5N23_40, partial (21%)
Length = 585
Score = 33.1 bits (74), Expect = 0.13
Identities = 21/81 (25%), Positives = 37/81 (44%), Gaps = 2/81 (2%)
Frame = +3
Query: 520 RRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEP 579
+ + T++P++ SL +S S S + N +++ +P P P + T P
Sbjct: 135 KNTTTTTNPSKNPSLAPSSSYASASYSAVSAPNKPHGRILVLTRPTP----KPVTHPTTP 302
Query: 580 IQPETQPQTQAEP--QAPKSP 598
P+ QP +Q P +AP P
Sbjct: 303 APPQPQPPSQPSPIQRAPDHP 365
>BP044109
Length = 497
Score = 32.3 bits (72), Expect = 0.23
Identities = 30/110 (27%), Positives = 43/110 (38%), Gaps = 2/110 (1%)
Frame = +2
Query: 498 PLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITE--SINSDS 555
P + +T AS TS+ R T+ P S + P +K T S S
Sbjct: 128 PPNKNKTTPSHASNATSK-STPRTQTPTTPPPSPTSSPKPPPSTFATKPSTSPPETPSSS 304
Query: 556 ALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPKSPIQTATTA 605
+ P P P S+S PTST + P P + P+A +P + A
Sbjct: 305 SHGPAPTLPSPPSSSTPTSTPSPP-SPPNGSTLPSPPRATPTPAPSLPAA 451
>TC19696 similar to UP|Q9ZQT8 (Q9ZQT8) WREBP-2 protein, partial (12%)
Length = 496
Score = 32.0 bits (71), Expect = 0.29
Identities = 22/80 (27%), Positives = 35/80 (43%), Gaps = 10/80 (12%)
Frame = +3
Query: 370 SSKPSDPKGKGILKEEPAKKKAASKTV-----IIREPSPERPLKRTSAQQQQMSDSESSE 424
S K S PK L +K ++ T +I++P P + ++ D ++ E
Sbjct: 18 SQKQSKPKPNLKLPPSKMARKGSAPTPSAPLNVIKKPWPVKQETYDEDSEETEEDRDNVE 197
Query: 425 DTWK-----DSSSEETEDED 439
D W+ + EETEDED
Sbjct: 198 DGWRYAGNNEDDDEETEDED 257
>TC13318 similar to UP|CAA20314 (CAA20314) SPBC30B4.01c protein (SPBC3D6.14C
protein) (Fragment), partial (15%)
Length = 543
Score = 32.0 bits (71), Expect = 0.29
Identities = 17/67 (25%), Positives = 30/67 (44%), Gaps = 2/67 (2%)
Frame = +3
Query: 397 IIREPS--PERPLKRTSAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEE 454
++ +P P+ P K ++ D + ED + ++ +D+D EEEE
Sbjct: 192 LVHDPEIYPDGPPKPVQTDNEEEEDDDEEEDDEDEDDDDDDDDDDDG---------EEEE 344
Query: 455 DEQDDHE 461
D+ DD E
Sbjct: 345 DDDDDDE 365
>TC9596 similar to UP|AAR24722 (AAR24722) At5g49400, partial (66%)
Length = 1196
Score = 31.2 bits (69), Expect = 0.50
Identities = 30/115 (26%), Positives = 53/115 (46%), Gaps = 5/115 (4%)
Frame = +2
Query: 383 KEEPAKKKAASKTVIIREPSP-----ERPLKRTSAQQQQMSDSESSEDTWKDSSSEETED 437
KEE AK + R S E + T + ++ S+ S + DSSS ++E+
Sbjct: 590 KEEKAKSSSKKAKRKHRSDSDSGSDSEDSVFETESGSSSVTGSDYSSGSSSDSSSSDSEE 769
Query: 438 EDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVKRRK 492
E +RR+ +++ + H+ + E+SD+E ++DSD+ RRK
Sbjct: 770 E----RRRRRKKKTQKKKKGRRHKKYSSS----SESSDSESASDSDSDDKSSRRK 910
>TC15958 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Arabidopsis
thaliana;}, partial (52%)
Length = 800
Score = 30.8 bits (68), Expect = 0.66
Identities = 32/106 (30%), Positives = 40/106 (37%), Gaps = 15/106 (14%)
Frame = +3
Query: 509 ASEPTSEVQRERRSKRTSDPARAASLTRTSP--IRSLSKVITESINSDSALVVIPEQPMP 566
++ P S R P AA + SP + IT+S S SA IP P P
Sbjct: 261 SASPASSATASPRFSLFPTPPAAAGKSTASPAALDPPGTPITQSTTS-SATSPIPTTPKP 437
Query: 567 ISTSLPTS-----------TQTEPIQPETQP--QTQAEPQAPKSPI 599
S S PTS T I T P +T + P +P PI
Sbjct: 438 SSPSSPTSPTSSTSPGPTATPNRRIARSTAPCSETSSAPSSPTRPI 575
>TC17157 similar to UP|CAD27462 (CAD27462) Nucleosome assembly protein
1-like protein 3, partial (81%)
Length = 1298
Score = 30.8 bits (68), Expect = 0.66
Identities = 17/48 (35%), Positives = 24/48 (49%)
Frame = +2
Query: 414 QQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHE 461
+ + SD E E+ +D +E ED+D EEEED+ DD E
Sbjct: 743 EAEQSDFEDIEEDDEDGDEDEDEDDDDD---------EEEEDDDDDDE 859
>TC10598 weakly similar to GB|AAM78036.1|21928015|AY125526
At2g01100/F23H14.7 {Arabidopsis thaliana;}, partial
(33%)
Length = 902
Score = 30.8 bits (68), Expect = 0.66
Identities = 45/189 (23%), Positives = 76/189 (39%), Gaps = 19/189 (10%)
Frame = +3
Query: 375 DPKGKGILKEEPA------KKKAASKTVIIREPSPERPLKRTSAQQQQMSDSESSEDTWK 428
+ K K I++ + A K +AA+K E + ER LK +++ SDSES D+
Sbjct: 261 EEKKKKIMERKEAPLKWEQKLEAAAKAKADAE-ARERKLKAAKHKKRADSDSESDYDS-- 431
Query: 429 DSSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVV 488
+DE ++R ++ DHE + + + + ES+D
Sbjct: 432 -------DDESKRTSRRSHRKHKKRSHYDSDHEKRKEKLSKRKTKKWSSESSDFSGDESE 590
Query: 489 KRRKLPLRGPLQQKETAAEQASEPTSE---------VQRERRSKR----TSDPARAASLT 535
R + R +QK+ +Q S S +R+R+ KR S + +S
Sbjct: 591 SRSEEERRRKRKQKKKLRDQVSRSDSSDSDSSADEVSKRKRQHKRHRVTKSSGSDFSSDE 770
Query: 536 RTSPIRSLS 544
SP+R S
Sbjct: 771 ENSPVRKRS 797
>TC11009 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;}, partial (44%)
Length = 700
Score = 30.0 bits (66), Expect = 1.1
Identities = 24/81 (29%), Positives = 35/81 (42%), Gaps = 15/81 (18%)
Frame = +2
Query: 419 DSESSEDTWKDSSSEETEDEDM--PLAKRRKII----------LEEEEDEQDDHEIIQNV 466
D + +D D EE E+ED+ RR + E EEDE DD++
Sbjct: 170 DPDDDDDDDDDDDEEEEEEEDLGTEYLVRRTVAAAEDEEASSDFEPEEDEGDDNDNDDGE 349
Query: 467 IQGI---REASDAEESTDSDA 484
G+ R+ SD + S D D+
Sbjct: 350 KAGVPSKRKRSDKDGSGDDDS 412
>TC17487 UP|CAE45597 (CAE45597) SAR DNA-binding protein-like protein, complete
Length = 1794
Score = 30.0 bits (66), Expect = 1.1
Identities = 28/118 (23%), Positives = 55/118 (45%), Gaps = 6/118 (5%)
Frame = +1
Query: 416 QMSDSESSEDTWKDSSSEETEDEDMPLAKRRK----IILEEEEDEQDDHEIIQNVIQGIR 471
Q SDS EDT + S+ ++ E ++ K + + L +ED ++ E+++ + R
Sbjct: 1471 QKSDSAMDEDTHEPSADKKKEKKEKKKKKEKNEEKDVPLLADEDGDEEKEVVKKEKKKKR 1650
Query: 472 EAS--DAEESTDSDAIPVVKRRKLPLRGPLQQKETAAEQASEPTSEVQRERRSKRTSD 527
+ S + E DA K+RK K E+++E S +++ + K++ D
Sbjct: 1651 KESTENVELQNGDDAGEKKKKRK---------KHAEPEESAEMPS--KKKGKKKKSED 1791
>TC8606 similar to UP|AAQ82688 (AAQ82688) Epa4p, partial (7%)
Length = 1008
Score = 30.0 bits (66), Expect = 1.1
Identities = 13/51 (25%), Positives = 25/51 (48%)
Frame = -2
Query: 411 SAQQQQMSDSESSEDTWKDSSSEETEDEDMPLAKRRKIILEEEEDEQDDHE 461
+ + ++ D E ED D ++ ED+ + + E+ EDE++D E
Sbjct: 437 NGEDEEDEDGEDQEDDDDDDEDDDEEDDGGEDEEEEGVEEEDNEDEEEDEE 285
>TC19569 similar to UP|AAR11301 (AAR11301) Lectin-like receptor kinase 1;1,
partial (16%)
Length = 486
Score = 30.0 bits (66), Expect = 1.1
Identities = 31/121 (25%), Positives = 48/121 (39%), Gaps = 14/121 (11%)
Frame = +1
Query: 498 PLQQKETAAEQASEPTSEVQRERRSKRTSDPARAASLTRTSPIRSLSKVITESINSDSAL 557
P Q+ T + S ++ +K T +P A S + SP S S+ + N ++
Sbjct: 82 PPSQQPTHSPSTSPTSTTTPPPSPTKVTENPPTAPSTSTKSPTSSASEEPSPP-NPSTST 258
Query: 558 VVIPEQ----------PMPISTSLPT---STQTEPIQ-PETQPQTQAEPQAPKSPIQTAT 603
P+ P P ST PT S T P+ ++ P A P A +P T T
Sbjct: 259 TQQPKHSPTSPPASPSPSPPSTKPPTAMASPSTSPLSATKSHPTPPAAPSASSTPPPTTT 438
Query: 604 T 604
+
Sbjct: 439 S 441
Score = 28.9 bits (63), Expect = 2.5
Identities = 21/69 (30%), Positives = 32/69 (45%)
Frame = +1
Query: 537 TSPIRSLSKVITESINSDSALVVIPEQPMPISTSLPTSTQTEPIQPETQPQTQAEPQAPK 596
TSP L + S +S S +QP ++ PTST T P P T+ P AP
Sbjct: 16 TSPCGLLLHTSSSSSSSYSPPPPPSQQPTHSPSTSPTSTTTPP--PSPTKVTENPPTAPS 189
Query: 597 SPIQTATTA 605
+ ++ T++
Sbjct: 190 TSTKSPTSS 216
>AI967639
Length = 335
Score = 30.0 bits (66), Expect = 1.1
Identities = 26/89 (29%), Positives = 41/89 (45%)
Frame = +1
Query: 430 SSSEETEDEDMPLAKRRKIILEEEEDEQDDHEIIQNVIQGIREASDAEESTDSDAIPVVK 489
SSS E ++ED + + LE+EE++ DD + D S DSD+ K
Sbjct: 85 SSSSEGDEEDEEEDQGDEYNLEDEEEDDDDD-----------DDDDLSISEDSDSDKPRK 231
Query: 490 RRKLPLRGPLQQKETAAEQASEPTSEVQR 518
++LP R ++ET E S ++R
Sbjct: 232 VKQLPGR---TRRETKRRSVDEIQSGLRR 309
>AW719831
Length = 436
Score = 29.6 bits (65), Expect = 1.5
Identities = 20/77 (25%), Positives = 30/77 (37%), Gaps = 3/77 (3%)
Frame = +1
Query: 525 TSDPARAASLTRTSPIRSLSKVIT---ESINSDSALVVIPEQPMPISTSLPTSTQTEPIQ 581
T P + T T P + ++ + +S A P P +T+ PTS P
Sbjct: 166 TPPPTMTPTTTTTPPSTNPHLLLPPPPHARSSSCAATAAKSSPAPTTTTSPTS----PAT 333
Query: 582 PETQPQTQAEPQAPKSP 598
P + P T + P SP
Sbjct: 334 PRSSPSTASSSSPPSSP 384
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.312 0.129 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,725,262
Number of Sequences: 28460
Number of extensions: 113023
Number of successful extensions: 862
Number of sequences better than 10.0: 84
Number of HSP's better than 10.0 without gapping: 805
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 836
length of query: 607
length of database: 4,897,600
effective HSP length: 96
effective length of query: 511
effective length of database: 2,165,440
effective search space: 1106539840
effective search space used: 1106539840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0234.12


