

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
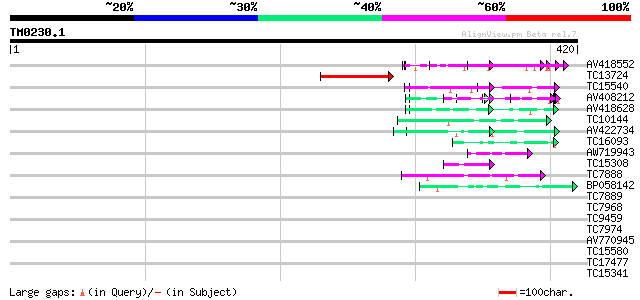
Query= TM0230.1
(420 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV418552 74 5e-14
TC13724 weakly similar to UP|Q9ZQ35 (Q9ZQ35) At2g24320 protein, ... 69 2e-12
TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fra... 65 2e-11
AV408212 53 8e-08
AV418628 44 4e-05
TC10144 similar to UP|Q93YH8 (Q93YH8) HMG I/Y like protein, part... 44 7e-05
AV422734 43 9e-05
TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transc... 42 1e-04
AW719943 41 4e-04
TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%) 41 4e-04
TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 41 4e-04
BP058142 40 7e-04
TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall prot... 40 0.001
TC7968 similar to UP|Q39360 (Q39360) (Clone PIS63-1), partial (15%) 38 0.003
TC9459 similar to AAQ23982 (AAQ23982) Transcription factor Rap21... 37 0.005
TC7974 similar to GB|AAA77675.1|607870|DSU11801 period protein {... 37 0.005
AV770945 37 0.006
TC15580 37 0.008
TC17477 weakly similar to UP|DHSS_ANACY (P16421) Soluble hydroge... 36 0.010
TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich pro... 36 0.014
>AV418552
Length = 386
Score = 73.9 bits (180), Expect = 5e-14
Identities = 43/119 (36%), Positives = 53/119 (44%), Gaps = 3/119 (2%)
Frame = -3
Query: 293 SCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCL 352
SC +C CC +C C +C C C C +CSC SC C CCC
Sbjct: 345 SC*NC------CCGKSC--CWSC------VC*C------CVSCSC--ASCCWSCASCCCW 232
Query: 353 PSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCC---MSCFCWRCCV 408
SC+ CC S C SCC C+ CC+ CCC+ + R SCC CW CC+
Sbjct: 231 -SCVSCCCWSSVS-CSCESCCWSCA-SCCCWSCCCSCK*GR-SCCWG*KYDSICWCCCI 67
Score = 73.6 bits (179), Expect = 6e-14
Identities = 39/101 (38%), Positives = 48/101 (46%), Gaps = 11/101 (10%)
Frame = -3
Query: 312 CSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSS 371
C NC C CC SC+ C +CSC +C S C CCC SC+ CC S C S
Sbjct: 342 C*NCCC-GKSCC-WSCVC*CCVSCSCASCCWS--CASCCCW-SCVSCCCWSSVS-CSCES 181
Query: 372 CCTKCSRPSCCFKCCCNLRCARPSC-----------CCMSC 401
CC C+ CC+ CCC+ + R C CC+SC
Sbjct: 180 CCWSCAS-CCCWSCCCSCK*GRSCCWG*KYDSICWCCCISC 61
Score = 65.5 bits (158), Expect = 2e-11
Identities = 32/81 (39%), Positives = 38/81 (46%), Gaps = 6/81 (7%)
Frame = -3
Query: 340 CSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSC--CFKCCC--NLRCARPS 395
C C C CCC SC C +C +SCC C+ C C CCC ++ C+ S
Sbjct: 360 C*C*GSC*NCCCGKSCCWSCVC*CCVSCSCASCCWSCASCCCWSCVSCCCWSSVSCSCES 181
Query: 396 CC--CMSCFCWRCCVGKHSGR 414
CC C SC CW CC GR
Sbjct: 180 CCWSCASCCCWSCCCSCK*GR 118
Score = 56.6 bits (135), Expect = 7e-09
Identities = 27/77 (35%), Positives = 33/77 (42%), Gaps = 11/77 (14%)
Frame = -3
Query: 294 CFSCSC--CRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCC 351
C SCSC C C S C C +C C + C+C C +C C +C CS K + CC
Sbjct: 288 CVSCSCASCCWSCASCCCWSCVSCCCWSSVSCSCESCCWSCASCCCWSCCCSCK*GRSCC 109
Query: 352 LP---------SCIKCC 359
CI CC
Sbjct: 108 WG*KYDSICWCCCISCC 58
Score = 55.1 bits (131), Expect = 2e-08
Identities = 36/111 (32%), Positives = 48/111 (42%), Gaps = 5/111 (4%)
Frame = -3
Query: 292 SSCFSCSC-CRMKC-CSNNCTGCSNCSCIN-PECCNCSCINPECFNC--SCINCSCSLKC 346
S C+SC C C + C C++ C C++C C + CC S ++ C +C SC +C C
Sbjct: 318 SCCWSCVC*CCVSCSCASCCWSCASCCCWSCVSCCCWSSVSCSCESCCWSCASCC----C 151
Query: 347 TQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCC 397
CCC S G SCC S C+ CC SCC
Sbjct: 150 WSCCC-------------SCK*GRSCCWG*KYDSICWCCCI-------SCC 58
Score = 30.0 bits (66), Expect = 0.75
Identities = 22/55 (40%), Positives = 26/55 (47%), Gaps = 1/55 (1%)
Frame = -3
Query: 290 NSSSCFSCSCCRMKCCSNNCTGCSNCSCI-NPECCNCSCINPECFNCSCINCSCS 343
+S SC SC C C S C C CSC CC + C+ C CI+C CS
Sbjct: 207 SSVSC-SCESCCWSCASCCCWSCC-CSCK*GRSCCWG*KYDSICW-CCCISC-CS 55
>TC13724 weakly similar to UP|Q9ZQ35 (Q9ZQ35) At2g24320 protein, partial
(23%)
Length = 639
Score = 68.6 bits (166), Expect = 2e-12
Identities = 28/55 (50%), Positives = 41/55 (73%), Gaps = 1/55 (1%)
Frame = +2
Query: 231 LWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESIT-TEKRLKKLRQS 284
LW++++E YDPLW+K GGHCNLE +P+Y+++L KFI ME + T K+L Q+
Sbjct: 8 LWELSKEKYDPLWVKXGGHCNLETFPEYLKYLRKFINAMEKLAITRPTNKQLTQN 172
>TC15540 similar to UP|Q9NGS5 (Q9NGS5) Prespore protein MF12 (Fragment),
partial (6%)
Length = 806
Score = 65.1 bits (157), Expect = 2e-11
Identities = 40/112 (35%), Positives = 44/112 (38%), Gaps = 1/112 (0%)
Frame = -3
Query: 297 CSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCL-PSC 355
C CC C NC C +C C CSC +C C + CCC+ SC
Sbjct: 240 CCCC*GSC*CCNCCNCGSCCC----------------GCSCESCCC*S*GSCCCCIWESC 109
Query: 356 IKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCC 407
CC TC GS CC C C CCC SCCC C C CC
Sbjct: 108 --CCW-----TC-GSCCCCCCWSWVSCGGCCC-------SCCC-CCSCGSCC 1
Score = 57.4 bits (137), Expect = 4e-09
Identities = 29/75 (38%), Positives = 34/75 (44%), Gaps = 8/75 (10%)
Frame = -3
Query: 293 SCFSCSCCRMKCCSNNCTGCSNCSCIN-PECCNC---SCINPECFNCSCINC----SCSL 344
SC C+CC C C+ C +C C + CC C SC C +C C C SC
Sbjct: 222 SC*CCNCCNCGSCCCGCS-CESCCC*S*GSCCCCIWESCCCWTCGSCCCCCCWSWVSCGG 46
Query: 345 KCTQCCCLPSCIKCC 359
C CCC SC CC
Sbjct: 45 CCCSCCCCCSCGSCC 1
Score = 51.6 bits (122), Expect = 2e-07
Identities = 26/65 (40%), Positives = 28/65 (43%), Gaps = 4/65 (6%)
Frame = -3
Query: 347 TQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCF----KCCCNLRCARPSCCCMSCF 402
T CCC CC + C SCC CS SCC CCC C SCCC +C
Sbjct: 246 TICCCC*GSC*CC-----NCCNCGSCCCGCSCESCCC*S*GSCCC---CIWESCCCWTCG 91
Query: 403 CWRCC 407
CC
Sbjct: 90 SCCCC 76
Score = 38.9 bits (89), Expect = 0.002
Identities = 23/67 (34%), Positives = 26/67 (38%), Gaps = 12/67 (17%)
Frame = -3
Query: 352 LPSCIKCCHLPKFSTCFGSS-CCTKCSRPSCCFKCCCNLRCARPSCCC-----------M 399
+P+ CC C GS CC C+ SCC C C SCCC
Sbjct: 258 IPAPTICC------CC*GSC*CCNCCNCGSCCCGCSCE------SCCC*S*GSCCCCIWE 115
Query: 400 SCFCWRC 406
SC CW C
Sbjct: 114 SCCCWTC 94
Score = 28.1 bits (61), Expect = 2.8
Identities = 13/31 (41%), Positives = 14/31 (44%)
Frame = -3
Query: 379 PSCCFKCCCNLRCARPSCCCMSCFCWRCCVG 409
P+ C CCC C CC C C CC G
Sbjct: 249 PTIC--CCC*GSC*----CCNCCNCGSCCCG 175
>AV408212
Length = 362
Score = 53.1 bits (126), Expect = 8e-08
Identities = 28/83 (33%), Positives = 34/83 (40%)
Frame = -2
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSC 381
CC+C C C +C C C C CCC C CC C+ ++C C+R C
Sbjct: 172 CCDCGCC---CCDCCCCCCCC------CCC---CCCCC-------CWRAACL--CNRICC 56
Query: 382 CFKCCCNLRCARPSCCCMSCFCW 404
CCC CCC CW
Sbjct: 55 ICMCCC-------CCCC*CGDCW 8
Score = 52.8 bits (125), Expect = 1e-07
Identities = 26/76 (34%), Positives = 29/76 (37%), Gaps = 1/76 (1%)
Frame = -2
Query: 332 CFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSC-CFKCCCNLR 390
C +C C C C C CCC CC CC C R +C C + CC
Sbjct: 172 CCDCGCCCCDCCCCCCCCCC------CC------------CCCCCWRAACLCNRICCICM 47
Query: 391 CARPSCCCMSCFCWRC 406
C CCC C C C
Sbjct: 46 C----CCCCCC*CGDC 11
Score = 48.1 bits (113), Expect = 3e-06
Identities = 23/62 (37%), Positives = 24/62 (38%)
Frame = -2
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLP 353
C C CC CC C C C C CC C C C C+ I C C C CC
Sbjct: 172 CCDCGCCCCDCC---CCCCCCCCC----CCCCCCWRAACL-CNRICCICMCCCCCCC*CG 17
Query: 354 SC 355
C
Sbjct: 16 DC 11
Score = 45.4 bits (106), Expect = 2e-05
Identities = 20/63 (31%), Positives = 24/63 (37%)
Frame = -2
Query: 297 CSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCI 356
C+ + CC C C C C CC C C C C C +C C C+ C
Sbjct: 193 CALSLLCCCDCGCCCCDCCCCC---CCCCCC----CCCCCCWRAACLCNRICCICMCCCC 35
Query: 357 KCC 359
CC
Sbjct: 34 CCC 26
Score = 44.7 bits (104), Expect = 3e-05
Identities = 21/59 (35%), Positives = 24/59 (40%), Gaps = 2/59 (3%)
Frame = -2
Query: 351 CLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCC--MSCFCWRCC 407
C S + CC G CC C CC CCC C R +C C + C C CC
Sbjct: 193 CALSLLCCCDC-------GCCCCDCCCCCCCCCCCCCCCCCWRAACLCNRICCICMCCC 38
Score = 41.2 bits (95), Expect = 3e-04
Identities = 18/45 (40%), Positives = 19/45 (42%), Gaps = 8/45 (17%)
Frame = -2
Query: 372 CCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWR--------CCV 408
CC C CC CCC C CCC C CWR CC+
Sbjct: 175 CCCDCGC-CCCDCCCC---CCCCCCCCCCCCCWRAACLCNRICCI 53
Score = 35.4 bits (80), Expect = 0.018
Identities = 14/40 (35%), Positives = 17/40 (42%)
Frame = -2
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECF 333
C C CC CC C+ CI CC C C +C+
Sbjct: 127 CCCCCCCCCCCCWRAACLCNRICCICMCCCCCCC*CGDCW 8
>AV418628
Length = 309
Score = 44.3 bits (103), Expect = 4e-05
Identities = 35/116 (30%), Positives = 38/116 (32%), Gaps = 6/116 (5%)
Frame = -2
Query: 297 CSCCRMKCCSNNCTGCSNCSCINPECCNCSCINP-ECFNCSCINCSCSLKCTQCCCLPSC 355
C C + S C C CC S + +C C C C Q CC C
Sbjct: 305 CCCPSSRPSSRQCGRRQGGRCRYRCCCPSSRPSSRQCGRRQGGRCRCRCCCCQSCCPSRC 126
Query: 356 IKCCHLPKFSTCFGSSCCTKCSRPSCCFK-----CCCNLRCARPSCCCMSCFCWRC 406
C SSCC RPS C C RC CCC SC RC
Sbjct: 125 SSC-----------SSCC----RPSSCRSGRRQGGRCRCRC----CCCQSCCPSRC 15
Score = 41.6 bits (96), Expect = 2e-04
Identities = 21/65 (32%), Positives = 24/65 (36%)
Frame = -2
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLP 353
C CC CC + C+ CS SC P C + C C C C Q CC
Sbjct: 173 CRCRCCCCQSCCPSRCSSCS--SCCRPSSCRSG--RRQGGRCRCRCC-----CCQSCCPS 21
Query: 354 SCIKC 358
C C
Sbjct: 20 RCSSC 6
Score = 37.4 bits (85), Expect = 0.005
Identities = 26/88 (29%), Positives = 31/88 (34%), Gaps = 20/88 (22%)
Frame = -2
Query: 345 KCTQCCCLPSCIKCCHLPKFSTCF---GSSCCTKCSRPSCCFKCCCNLRCA------RPS 395
+C CC PS P C G C +C CC + CC RC+ RPS
Sbjct: 248 RCRYRCCCPSS-----RPSSRQCGRRQGGRCRCRC----CCCQSCCPSRCSSCSSCCRPS 96
Query: 396 CC-----------CMSCFCWRCCVGKHS 412
C C C C CC + S
Sbjct: 95 SCRSGRRQGGRCRCRCCCCQSCCPSRCS 12
Score = 28.5 bits (62), Expect = 2.2
Identities = 25/83 (30%), Positives = 31/83 (37%), Gaps = 10/83 (12%)
Frame = -2
Query: 348 QCCCLPS--CIKCCHLPKFSTCFGSSCCTKCSRPSC--CFK-----CCCNLRCARPSCCC 398
+CCC S + C + C CC SRPS C + C C C + SCC
Sbjct: 308 RCCCPSSRPSSRQCGRRQGGRC-RYRCCCPSSRPSSRQCGRRQGGRCRCRCCCCQ-SCCP 135
Query: 399 MSC-FCWRCCVGKHSGRDEKQSG 420
C C CC +Q G
Sbjct: 134 SRCSSCSSCCRPSSCRSGRRQGG 66
Score = 26.6 bits (57), Expect = 8.2
Identities = 8/20 (40%), Positives = 10/20 (50%)
Frame = -2
Query: 294 CFSCSCCRMKCCSNNCTGCS 313
C CC CC + C+ CS
Sbjct: 62 CRCRCCCCQSCCPSRCSSCS 3
>TC10144 similar to UP|Q93YH8 (Q93YH8) HMG I/Y like protein, partial (8%)
Length = 600
Score = 43.5 bits (101), Expect = 7e-05
Identities = 31/127 (24%), Positives = 48/127 (37%), Gaps = 13/127 (10%)
Frame = -3
Query: 288 QSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECC-------------NCSCINPECFN 334
+++S++ +S C C+NCS +N C N S + P+
Sbjct: 454 KTSSATSYSLYKSVTSCALLT*NTCTNCS*LN*RPCSSFLAS*SN*GPLNSSYLGPDS*G 275
Query: 335 CSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARP 394
C C C CC+ + CC ++ C G +CC+ CC CCC
Sbjct: 274 C*NGCCCCRW---WYCCI*NGGYCC*---YAGCCGGTCCS--GGTCCCGTCCCGCASCGS 119
Query: 395 SCCCMSC 401
+C C C
Sbjct: 118 TCGCCCC 98
Score = 33.9 bits (76), Expect = 0.052
Identities = 23/75 (30%), Positives = 26/75 (34%), Gaps = 1/75 (1%)
Frame = -3
Query: 297 CSCCRM-KCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSC 355
C CCR CC N C C CC +C CS C C C C S
Sbjct: 262 CCCCRWWYCCI*NGGYC----C*YAGCCGGTC-------CSGGTCCCGTCCCGCASCGST 116
Query: 356 IKCCHLPKFSTCFGS 370
CC S+ G+
Sbjct: 115 CGCCCCGSVSSLKGA 71
>AV422734
Length = 448
Score = 43.1 bits (100), Expect = 9e-05
Identities = 22/75 (29%), Positives = 30/75 (39%)
Frame = -3
Query: 285 QGPQSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSL 344
+G + +++C C CC CC++ C C C + CC C C C C C
Sbjct: 251 EGKREIAAACCCCCCCFCNCCNSRKCCCCCCCC*SISCCWC---------C*C--CCC*C 105
Query: 345 KCTQCCCLPSCIKCC 359
C C S I C
Sbjct: 104 *CCWCNAFASSIFVC 60
Score = 40.4 bits (93), Expect = 6e-04
Identities = 35/118 (29%), Positives = 40/118 (33%), Gaps = 5/118 (4%)
Frame = -3
Query: 295 FSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCIN--PECFNCSCINCSCSLKCTQCCCL 352
F C CC C GC CC C C F C L C L
Sbjct: 416 FCCCCC--------CWGC---------CCCCCC*RWLELGFWCKLRMGLLGLGLGFCWRL 288
Query: 353 PSCI--KCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLR-CARPSCCCMSCFCWRCC 407
+ K + + ++CC C CCF CCN R C CCC S C CC
Sbjct: 287 MGLMGRK*ASMLEGKREIAAACCCCC----CCFCNCCNSRKCCCCCCCC*SISCCWCC 126
Score = 38.5 bits (88), Expect = 0.002
Identities = 26/77 (33%), Positives = 27/77 (34%)
Frame = -3
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSC 381
CC C C CF C+C N S KC CCC I CC C
Sbjct: 224 CCCCCC----CF-CNCCN---SRKCCCCCCCC*SISCCW--------------------C 129
Query: 382 CFKCCCNLRCARPSCCC 398
C CCC C C C
Sbjct: 128 C*CCCC*C*C----CWC 90
Score = 35.4 bits (80), Expect = 0.018
Identities = 16/38 (42%), Positives = 17/38 (44%), Gaps = 2/38 (5%)
Frame = -3
Query: 372 CCTKCSRPSCCF--KCCCNLRCARPSCCCMSCFCWRCC 407
CC C +CC KCCC C CC C C CC
Sbjct: 221 CCCCCCFCNCCNSRKCCCCCCCC*SISCCWCC*C--CC 114
Score = 35.0 bits (79), Expect = 0.023
Identities = 21/60 (35%), Positives = 22/60 (36%), Gaps = 4/60 (6%)
Frame = -3
Query: 349 CCCLPSCIKCCHLPKFSTCFGSS----CCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCW 404
CCC CC F C S CC C SCC+ C C CCC CW
Sbjct: 218 CCC------CC----FCNCCNSRKCCCCCCCC*SISCCWCC*C--------CCC*C*CCW 93
Score = 33.9 bits (76), Expect = 0.052
Identities = 18/58 (31%), Positives = 22/58 (37%)
Frame = -3
Query: 290 NSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCT 347
N + C CC C S +C C C C CC C+ F C I S + T
Sbjct: 197 NCCNSRKCCCCCCCC*SISCCWCC*CCCC*C*CCWCNAFASSIFVCQEIEQSDETETT 24
Score = 33.1 bits (74), Expect = 0.088
Identities = 19/61 (31%), Positives = 20/61 (32%)
Frame = -3
Query: 346 CTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWR 405
C CCC +C C CC C CC C C CCC C C
Sbjct: 221 CCCCCCFCNC-----------CNSRKCCCCCC---CC*SISCCWCC---*CCCC*C*CCW 93
Query: 406 C 406
C
Sbjct: 92 C 90
>TC16093 weakly similar to UP|PF21_ARATH (Q04088) Possible transcription
factor PosF21 (AtbZIP59), partial (17%)
Length = 570
Score = 42.4 bits (98), Expect = 1e-04
Identities = 27/81 (33%), Positives = 30/81 (36%), Gaps = 3/81 (3%)
Frame = -2
Query: 329 NPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCN 388
+P+C CSC CCC CI CC CC C CC CCC
Sbjct: 248 SPDC*CCSC-----------CCC---CICCC-----------GCCIGC----CCCCCCC* 156
Query: 389 LRCARPSCCCMSCF---CWRC 406
+ C CC CF W C
Sbjct: 155 IGC*N*CWCCW-CF*G*IWSC 96
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/44 (43%), Positives = 21/44 (47%)
Frame = -2
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSC 337
C SC CC + CC C GC C C CC C N C+ C C
Sbjct: 233 CCSCCCCCI-CCCGCCIGCCCCCC----CC*IGC*N-*CWCCWC 120
Score = 35.8 bits (81), Expect = 0.014
Identities = 17/37 (45%), Positives = 17/37 (45%)
Frame = -2
Query: 291 SSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSC 327
S C CSCC CC C GC CI CC C C
Sbjct: 248 SPDC*CCSCC---CCCICCCGC----CIGCCCCCCCC 159
Score = 33.9 bits (76), Expect = 0.052
Identities = 18/51 (35%), Positives = 18/51 (35%), Gaps = 7/51 (13%)
Frame = -2
Query: 363 KFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCF-------CWRC 406
K C SCC CC CCC C CCC C CW C
Sbjct: 251 KSPDC*CCSCC-------CCCICCCG--CCIGCCCCCCCC*IGC*N*CWCC 126
Score = 33.1 bits (74), Expect = 0.088
Identities = 12/27 (44%), Positives = 13/27 (47%)
Frame = -2
Query: 381 CCFKCCCNLRCARPSCCCMSCFCWRCC 407
CC CCC + C CC C C CC
Sbjct: 233 CCSCCCCCICCC--GCCIGCCCCCCCC 159
Score = 32.3 bits (72), Expect = 0.15
Identities = 15/38 (39%), Positives = 16/38 (41%)
Frame = -2
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCC 359
CC+C C C C CI C C CCC C C
Sbjct: 233 CCSCCCCCICCCGC-CIGCCCCC----CCC*IGC*N*C 135
Score = 32.0 bits (71), Expect = 0.20
Identities = 13/30 (43%), Positives = 14/30 (46%), Gaps = 1/30 (3%)
Frame = -2
Query: 379 PSC-CFKCCCNLRCARPSCCCMSCFCWRCC 407
P C C CCC C CC+ C C CC
Sbjct: 245 PDC*CCSCCCCCICC--CGCCIGCCCCCCC 162
Score = 26.9 bits (58), Expect = 6.3
Identities = 8/15 (53%), Positives = 9/15 (59%)
Frame = -2
Query: 395 SCCCMSCFCWRCCVG 409
SCCC C CC+G
Sbjct: 227 SCCCCCICCCGCCIG 183
>AW719943
Length = 508
Score = 40.8 bits (94), Expect = 4e-04
Identities = 18/48 (37%), Positives = 22/48 (45%)
Frame = -2
Query: 340 CSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC 387
C C C CCC +C CC L + C+ CC + CC CCC
Sbjct: 219 CCC---CCCCCCC*NCC-CCELNRLF-CWNGDCCCG*NNCCCCCGCCC 91
Score = 37.7 bits (86), Expect = 0.004
Identities = 20/64 (31%), Positives = 24/64 (37%)
Frame = -2
Query: 288 QSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCT 347
++N C C CC CC C + C N +CC C NC C C
Sbjct: 231 KNN*CCCCCCCCCC*NCC---CCELNRLFCWNGDCC-----------CG*NNCCC---CC 103
Query: 348 QCCC 351
CCC
Sbjct: 102 GCCC 91
Score = 34.7 bits (78), Expect = 0.030
Identities = 21/65 (32%), Positives = 21/65 (32%), Gaps = 6/65 (9%)
Frame = -2
Query: 349 CCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLR---CARPSCCC---MSCF 402
CCC C CC CC C CCC L C CCC C
Sbjct: 219 CCC---CCCCC------------CC*NC--------CCCELNRLFCWNGDCCCG*NNCCC 109
Query: 403 CWRCC 407
C CC
Sbjct: 108 CCGCC 94
Score = 33.9 bits (76), Expect = 0.052
Identities = 15/34 (44%), Positives = 16/34 (46%), Gaps = 5/34 (14%)
Frame = -2
Query: 381 CCFKCCCNLRCARPSCCCMS---CFCWR--CCVG 409
CC CCC C +CCC FCW CC G
Sbjct: 219 CCCCCCC---CCC*NCCCCELNRLFCWNGDCCCG 127
Score = 33.1 bits (74), Expect = 0.088
Identities = 18/53 (33%), Positives = 19/53 (34%)
Frame = -2
Query: 306 SNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKC 358
+N C C C C C NC C E C N C CCC C C
Sbjct: 228 NN*CCCCCCCCC----C*NCCCC--ELNRLFCWNGDCCCG*NNCCCCCGCCCC 88
>TC15308 similar to UP|Q41725 (Q41725) TED3, partial (16%)
Length = 1103
Score = 40.8 bits (94), Expect = 4e-04
Identities = 15/38 (39%), Positives = 18/38 (46%)
Frame = -3
Query: 322 CCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCC 359
CC C C C C +CSC+L+C C C CC
Sbjct: 555 CCYCCCSKVPCH---CCHCSCTLQCCSFLCTLHCCSCC 451
Score = 36.6 bits (83), Expect = 0.008
Identities = 19/62 (30%), Positives = 25/62 (39%)
Frame = -3
Query: 289 SNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQ 348
S+S S + C C CCS + CC+CSC C + C+L C
Sbjct: 588 SSSHSHYYCHNCCYCCCSK----------VPCHCCHCSC------TLQCCSFLCTLHCCS 457
Query: 349 CC 350
CC
Sbjct: 456 CC 451
Score = 32.7 bits (73), Expect = 0.11
Identities = 18/54 (33%), Positives = 21/54 (38%), Gaps = 4/54 (7%)
Frame = -3
Query: 355 CIKCCHLPKFSTCFGSSCCTKCSRPSC-CFKCCCNLRCARPSC---CCMSCFCW 404
C CC+ CC CS+ C C C C L+C C CC C W
Sbjct: 564 CHNCCY-----------CC--CSKVPCHCCHCSCTLQCCSFLCTLHCCSCCGNW 442
Score = 32.0 bits (71), Expect = 0.20
Identities = 22/80 (27%), Positives = 26/80 (32%), Gaps = 5/80 (6%)
Frame = -3
Query: 323 CNCSCINPECFNCSCINCSCSL-----KCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCS 377
CN +C +P S + S C CCC CCH SC +C
Sbjct: 639 CNGTCHSP*TLCHSFLPSSSHSHYYCHNCCYCCCSKVPCHCCH---------CSCTLQCC 487
Query: 378 RPSCCFKCCCNLRCARPSCC 397
C CC SCC
Sbjct: 486 SFLCTLHCC--------SCC 451
Score = 28.1 bits (61), Expect = 2.8
Identities = 13/41 (31%), Positives = 18/41 (43%), Gaps = 7/41 (17%)
Frame = -3
Query: 380 SCCFKCCCNLRCARPSC-CCMSCFCWRC------CVGKHSG 413
+CC+ CC + C C C + C + C C G SG
Sbjct: 558 NCCYCCCSKVPCHCCHCSCTLQCCSFLCTLHCCSCCGNWSG 436
>TC7888 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (60%)
Length = 913
Score = 40.8 bits (94), Expect = 4e-04
Identities = 30/113 (26%), Positives = 46/113 (40%), Gaps = 6/113 (5%)
Frame = +2
Query: 291 SSSCFSCSCCRMKCCSNN---CTGCSNCSCINPECCNCS-CINPECFNCSCINCSCSLKC 346
++SCFS C + CC++ C + C+ C + C C + CS + C
Sbjct: 545 ATSCFSTLCAKTTCCTSYTAICA*ATGCTGDTTLCAKTTHCETTHCLPSTL--CSKTSYC 718
Query: 347 TQCCCLPSCIKCCHLPKFST--CFGSSCCTKCSRPSCCFKCCCNLRCARPSCC 397
CLPS + C + T C S+ C+ + PSC C P+ C
Sbjct: 719 ETTNCLPSTL-CSNTSYCETTNCLPSTLCS--TTPSCTITTYCYTTYTNPTDC 868
Score = 37.4 bits (85), Expect = 0.005
Identities = 43/178 (24%), Positives = 61/178 (34%), Gaps = 46/178 (25%)
Frame = +2
Query: 273 TTEKRLKKLRQSQGPQSNSSSCFSCSCCRMKCCS--NNC--------TGCS---NC---- 315
TT+ K SQ ++SC + C CS NC T CS NC
Sbjct: 320 TTQAITKTTSLSQASSG*TTSCAKTTSCETTPCSKTTNCETTLCAKTTPCSQTTNCETSL 499
Query: 316 -----SCINPECCN--CSCINPECFNCSCIN------------------CSCSLKCTQCC 350
SC C+ SC + C +C C+ + C
Sbjct: 500 CA*TTSCETTTLCS*ATSCFSTLCAKTTCCTSYTAICA*ATGCTGDTTLCAKTTHCETTH 679
Query: 351 CLPSCIKCCHLPKFSTCFGSSC--CTKCSRPSCC--FKCCCNLRCARPSCCCMSCFCW 404
CLPS + K S C ++C T CS S C C + C+ C ++ +C+
Sbjct: 680 CLPSTL----CSKTSYCETTNCLPSTLCSNTSYCETTNCLPSTLCSTTPSCTITTYCY 841
>BP058142
Length = 571
Score = 40.0 bits (92), Expect = 7e-04
Identities = 35/126 (27%), Positives = 45/126 (34%), Gaps = 9/126 (7%)
Frame = -2
Query: 304 CCSNNCTGCSNC---------SCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPS 354
CCS+ TG + C SC CC+ S P C+CS+ C+ CC +
Sbjct: 531 CCSSIFTGSAACKSASIFTILSCTARICCSLSAFIPAV-------CACSI-CSSCCA--A 382
Query: 355 CIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCCVGKHSGR 414
C P C +S +R RC S CC S FC R C G +
Sbjct: 381 ATSCRSAP----CVAASLWFSVAR-----------RCI--STCCDSSFCSRTCGGGRRSK 253
Query: 415 DEKQSG 420
G
Sbjct: 252 GPPDRG 235
Score = 28.9 bits (63), Expect = 1.7
Identities = 21/70 (30%), Positives = 28/70 (40%), Gaps = 3/70 (4%)
Frame = -2
Query: 293 SCFSCSCCRMKCCSNNCTGCSNCSCINPECCNC--SCINPECFNCSCINCSCSLKCTQCC 350
SC + CC + CS CS CC SC + C S + S + +C C
Sbjct: 468 SCTARICCSLSAFIPAVCACSICS----SCCAAATSCRSAPCVAAS-LWFSVARRCISTC 304
Query: 351 CLPS-CIKCC 359
C S C + C
Sbjct: 303 CDSSFCSRTC 274
>TC7889 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (66%)
Length = 1158
Score = 39.7 bits (91), Expect = 0.001
Identities = 32/112 (28%), Positives = 46/112 (40%), Gaps = 14/112 (12%)
Frame = +3
Query: 291 SSSCFSCSCCRMKCCSNN------CTGCSNCS--CINPECCNCSCINPECFNCSCINCSC 342
++SCFS C + CC++ TGC+ + C C S + + C NC
Sbjct: 114 ATSCFSTLCAKTTCCTSYTAICA*ATGCTGDTTLCAKTTHCLPSTLCSKTSYCETTNCLP 293
Query: 343 SLKC--TQCC----CLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCN 388
S C T C CLPS + C P S + C T + P+ C+ N
Sbjct: 294 STLCSNTSYCETTNCLPSTL-CSTTP--SCTITTYCYTTYTNPTDCYTTYTN 440
Score = 36.2 bits (82), Expect = 0.010
Identities = 28/126 (22%), Positives = 43/126 (33%), Gaps = 5/126 (3%)
Frame = +3
Query: 284 SQGPQSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCI-NCSC 342
SQ +S C + C + T C + C CC + C C
Sbjct: 39 SQTTNCETSLCA*TTSCETTTLCS*ATSCFSTLCAKTTCCT-------SYTAICA*ATGC 197
Query: 343 SLKCTQCCCLPSCIKCCHLPKFSTCFGSSC--CTKCSRPSCC--FKCCCNLRCARPSCCC 398
+ T C C+ K S C ++C T CS S C C + C+ C
Sbjct: 198 TGDTTLCAKTTHCLPSTLCSKTSYCETTNCLPSTLCSNTSYCETTNCLPSTLCSTTPSCT 377
Query: 399 MSCFCW 404
++ +C+
Sbjct: 378 ITTYCY 395
>TC7968 similar to UP|Q39360 (Q39360) (Clone PIS63-1), partial (15%)
Length = 1193
Score = 38.1 bits (87), Expect = 0.003
Identities = 29/104 (27%), Positives = 38/104 (35%), Gaps = 21/104 (20%)
Frame = -2
Query: 294 CFSCSCC-------RMKCCSNNCTGCSN-----CSC---INPECCNCSCINPECFNCSCI 338
C+ CC + CC +GCS+ C+C + CC+ P +C C
Sbjct: 655 CWQHYCC*SCNFH*QQHCCCRKTSGCSSDFPWGCTCFLLL*TLCCSIHFGYPHKHSCCC- 479
Query: 339 NCSCSLKCTQCCCLPSC--IKCCHLPKFSTC----FGSSCCTKC 376
S C CCC S I C + F C SCC C
Sbjct: 478 ----SSNCCCCCCCESFL*ISCH*VDHFGWC*RNHPSGSCCNCC 359
Score = 32.3 bits (72), Expect = 0.15
Identities = 25/82 (30%), Positives = 31/82 (37%), Gaps = 11/82 (13%)
Frame = -2
Query: 337 CINCSCSLKCTQCCCLPSCIKCC--HLPKFSTCFGSSCCTKCSRPSC-CF----KCCCNL 389
C + L CC C C H + C +S C+ C CF CC++
Sbjct: 694 CHSSLAPLHPFHCCWQHYCC*SCNFH*QQHCCCRKTSGCSSDFPWGCTCFLLL*TLCCSI 515
Query: 390 RCARP---SCCCMS-CFCWRCC 407
P SCCC S C C CC
Sbjct: 514 HFGYPHKHSCCCSSNCCCCCCC 449
Score = 28.5 bits (62), Expect = 2.2
Identities = 22/83 (26%), Positives = 28/83 (33%), Gaps = 18/83 (21%)
Frame = -2
Query: 323 CNCSCINPECFNCSCINCSCSLKCT-----QCCCLPSCIKCCHLPKFSTCFG---SSCCT 374
C+ S F+C C C C CCC + P TCF + CC+
Sbjct: 694 CHSSLAPLHPFHC-CWQHYCC*SCNFH*QQHCCCRKTSGCSSDFPWGCTCFLLL*TLCCS 518
Query: 375 K----------CSRPSCCFKCCC 387
C +CC CCC
Sbjct: 517 IHFGYPHKHSCCCSSNCCCCCCC 449
>TC9459 similar to AAQ23982 (AAQ23982) Transcription factor Rap212, partial
(12%)
Length = 434
Score = 37.4 bits (85), Expect = 0.005
Identities = 21/60 (35%), Positives = 28/60 (46%)
Frame = -1
Query: 305 CSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKF 364
C ++ + SC CC C C C C C C CSL + C+PS ++CC KF
Sbjct: 350 CISSLPSSAASSC----CCYCYC----CCCCCCCCCYCSLLPGR-ICIPSILQCCIPCKF 198
Score = 33.9 bits (76), Expect = 0.052
Identities = 19/44 (43%), Positives = 20/44 (45%), Gaps = 8/44 (18%)
Frame = -1
Query: 370 SSCCTKCSRPSCCFKCCCNL-----RCARPSC--CCMSC-FCWR 405
SSCC C CC CCC R PS CC+ C F WR
Sbjct: 320 SSCCCYCYCCCCCCCCCCYCSLLPGRICIPSILQCCIPCKFPWR 189
Score = 30.0 bits (66), Expect = 0.75
Identities = 12/34 (35%), Positives = 12/34 (35%)
Frame = -1
Query: 373 CTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRC 406
C S CCC C CCC C C C
Sbjct: 350 CISSLPSSAASSCCCYCYC----CCCCCCCCCYC 261
Score = 28.5 bits (62), Expect = 2.2
Identities = 10/28 (35%), Positives = 13/28 (45%)
Frame = -1
Query: 380 SCCFKCCCNLRCARPSCCCMSCFCWRCC 407
S C +L + S CC C+C CC
Sbjct: 365 SASLPCISSLPSSAASSCCCYCYCCCCC 282
>TC7974 similar to GB|AAA77675.1|607870|DSU11801 period protein {Drosophila
simulans;} , partial (46%)
Length = 689
Score = 37.4 bits (85), Expect = 0.005
Identities = 17/47 (36%), Positives = 23/47 (48%)
Frame = -1
Query: 296 SCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSC 342
+CSC C CT C+C + C CSC + +CSC C+C
Sbjct: 131 ACSCANASTCG*TCTSAYACACSS--ACACSCAS----SCSC-GCAC 12
Score = 33.5 bits (75), Expect = 0.067
Identities = 17/48 (35%), Positives = 25/48 (51%), Gaps = 3/48 (6%)
Frame = -1
Query: 311 GCSNCSCINPECCNCSCINPECFNCSCIN---CSCSLKCTQCCCLPSC 355
GC+ CSC N C +C + + C+C + CSC+ C+ C C C
Sbjct: 137 GCA-CSCANASTCG*TCTS--AYACACSSACACSCASSCS-CGCACGC 6
Score = 27.7 bits (60), Expect = 3.7
Identities = 18/57 (31%), Positives = 21/57 (36%)
Frame = -1
Query: 347 TQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFC 403
T C C SC STC CT +C C C+ CA C +C C
Sbjct: 146 TPCGCACSCANA------STC--G*TCTSAYACACSSACACS--CASSCSCGCACGC 6
Score = 27.3 bits (59), Expect = 4.8
Identities = 13/43 (30%), Positives = 16/43 (36%), Gaps = 7/43 (16%)
Frame = -1
Query: 372 CCTKCSRPSCC-------FKCCCNLRCARPSCCCMSCFCWRCC 407
C C+ S C + C C+ CA C SC C C
Sbjct: 134 CACSCANASTCG*TCTSAYACACSSACA--CSCASSCSCGCAC 12
>AV770945
Length = 422
Score = 37.0 bits (84), Expect = 0.006
Identities = 21/72 (29%), Positives = 30/72 (41%), Gaps = 3/72 (4%)
Frame = -2
Query: 309 CTGCSNCSCINPECCNCSCINPECFNCSCI---NCSCSLKCTQCCCLPSCIKCCHLPKFS 365
C C + SC++ C + I+ C C+C+ C C C C CL + H+
Sbjct: 337 CVICVSHSCVSHVC---AYIDSHCL-CACVCVYMCLCVRMCVLCVCLYTHTDISHIYHMC 170
Query: 366 TCFGSSCCTKCS 377
CCT CS
Sbjct: 169 VSQCPHCCTLCS 134
Score = 30.4 bits (67), Expect = 0.57
Identities = 15/47 (31%), Positives = 16/47 (33%), Gaps = 2/47 (4%)
Frame = -2
Query: 364 FSTCFGSSCCTKCSRPSCCFKCCCNL--RCARPSCCCMSCFCWRCCV 408
F C S C C SC C + C C C C R CV
Sbjct: 364 FHDCVSSLVCVICVSHSCVSHVCAYIDSHCLCACVCVYMCLCVRMCV 224
Score = 26.6 bits (57), Expect = 8.2
Identities = 17/75 (22%), Positives = 28/75 (36%), Gaps = 12/75 (16%)
Frame = -2
Query: 323 CNCSCINPECFNCSCINCSCSLKCTQCCCLPSCI---KCCHLPKFSTCFGS--------- 370
C S + C + SC++ C+ + C C C+ C + C +
Sbjct: 355 CVSSLVCVICVSHSCVSHVCAYIDSHCLCACVCVYMCLCVRMCVLCVCLYTHTDISHIYH 176
Query: 371 SCCTKCSRPSCCFKC 385
C ++C P CC C
Sbjct: 175 MCVSQC--PHCCTLC 137
>TC15580
Length = 719
Score = 36.6 bits (83), Expect = 0.008
Identities = 18/50 (36%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Frame = +3
Query: 313 SNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSC---IKCC 359
SNC + C C C +CS ++ C++ CTQ C +C IKCC
Sbjct: 309 SNCCGRLADTCCCMCFLQ---SCSLVSDRCTIVCTQLCAALACFELIKCC 449
>TC17477 weakly similar to UP|DHSS_ANACY (P16421) Soluble hydrogenase 42 kDa
subunit (Tritium exchange subunit) , partial (6%)
Length = 539
Score = 36.2 bits (82), Expect = 0.010
Identities = 22/83 (26%), Positives = 27/83 (32%)
Frame = -2
Query: 305 CSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKF 364
C+ + CSC N C N + + CFN SC N
Sbjct: 442 CAIESSSTKLCSCFNRTCFNRTESSSLCFNRSCFN--------------------RTESS 323
Query: 365 STCFGSSCCTKCSRPSCCFKCCC 387
S CF SC + S CF C
Sbjct: 322 SLCFNRSCFNRTESSSLCFNRTC 254
Score = 31.6 bits (70), Expect = 0.26
Identities = 22/61 (36%), Positives = 25/61 (40%), Gaps = 12/61 (19%)
Frame = -2
Query: 291 SSSCFSCSCCRMKCCSNNCTG--CSNCSCINPE-----CCNCSCINPE-----CFNCSCI 338
SSS CSC C + + C N SC N C N SC N CFN +C
Sbjct: 430 SSSTKLCSCFNRTCFNRTESSSLCFNRSCFNRTESSSLCFNRSCFNRTESSSLCFNRTCF 251
Query: 339 N 339
N
Sbjct: 250 N 248
Score = 30.0 bits (66), Expect = 0.75
Identities = 16/45 (35%), Positives = 22/45 (48%)
Frame = -2
Query: 290 NSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFN 334
+SS CF+ SC + + + C N SC N + C N CFN
Sbjct: 373 SSSLCFNRSCFNR---TESSSLCFNRSCFNRTESSSLCFNRTCFN 248
Score = 29.3 bits (64), Expect = 1.3
Identities = 21/79 (26%), Positives = 23/79 (28%), Gaps = 1/79 (1%)
Frame = -2
Query: 325 CSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFK 384
CSC N CFN S CF SC + S CF
Sbjct: 412 CSCFNRTCFN-------------------------RTESSSLCFNRSCFNRTESSSLCFN 308
Query: 385 CCCNLRCARPSCCC-MSCF 402
C R S C +CF
Sbjct: 307 RSCFNRTESSSLCFNRTCF 251
Score = 27.7 bits (60), Expect = 3.7
Identities = 14/41 (34%), Positives = 17/41 (41%), Gaps = 1/41 (2%)
Frame = -2
Query: 363 KFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCC-MSCF 402
K +CF +C + S CF C R S C SCF
Sbjct: 418 KLCSCFNRTCFNRTESSSLCFNRSCFNRTESSSLCFNRSCF 296
>TC15341 similar to UP|Q9M0H8 (Q9M0H8) Predicted proline-rich protein,
partial (33%)
Length = 1002
Score = 35.8 bits (81), Expect = 0.014
Identities = 16/44 (36%), Positives = 18/44 (40%), Gaps = 1/44 (2%)
Frame = -2
Query: 367 CFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSC-FCWRCCVG 409
C G +C C CC CCC CCC C + CC G
Sbjct: 938 CVGCTC*GNCWGHCCCCCCCC--------CCCCGC*Y*GYCCTG 831
Score = 32.0 bits (71), Expect = 0.20
Identities = 11/30 (36%), Positives = 14/30 (46%), Gaps = 1/30 (3%)
Frame = -2
Query: 331 ECFNCSCINCSCSLKC-TQCCCLPSCIKCC 359
+C C+ C+C C CCC C CC
Sbjct: 956 DCIEGGCVGCTC*GNCWGHCCCCCCCCCCC 867
Score = 28.1 bits (61), Expect = 2.8
Identities = 11/30 (36%), Positives = 13/30 (42%), Gaps = 5/30 (16%)
Frame = -2
Query: 327 CINPECFNCSCI-----NCSCSLKCTQCCC 351
CI C C+C +C C C CCC
Sbjct: 953 CIEGGCVGCTC*GNCWGHCCCCCCCCCCCC 864
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.139 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,503,730
Number of Sequences: 28460
Number of extensions: 339496
Number of successful extensions: 6213
Number of sequences better than 10.0: 277
Number of HSP's better than 10.0 without gapping: 4005
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5385
length of query: 420
length of database: 4,897,600
effective HSP length: 93
effective length of query: 327
effective length of database: 2,250,820
effective search space: 736018140
effective search space used: 736018140
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0230.1


