

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0228a.2
(510 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
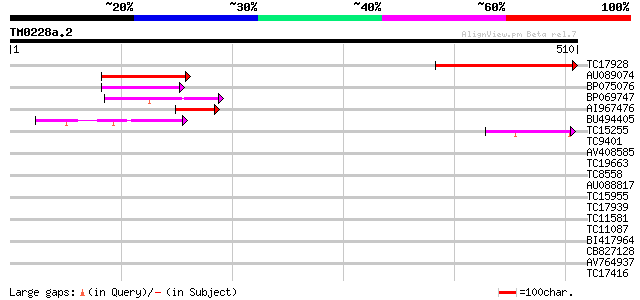
Sequences producing significant alignments: (bits) Value
TC17928 240 3e-64
AU089074 103 9e-23
BP075076 72 2e-13
BP069747 60 1e-09
AI967476 49 2e-06
BU494405 45 2e-05
TC15255 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (10%) 42 2e-04
TC9401 similar to UP|Q84LW7 (Q84LW7) Peptidylprolyl cis-trans is... 33 0.11
AV408585 33 0.11
TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation fac... 32 0.32
TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200... 32 0.32
AU088817 31 0.42
TC15955 31 0.55
TC17939 weakly similar to UP|Q945P2 (Q945P2) AT5g49210/K21P3_8, ... 31 0.55
TC11581 weakly similar to UP|Q99736 (Q99736) HsGCN1 (Fragment), ... 30 1.2
TC11087 weakly similar to UP|Q8GS92 (Q8GS92) OJ1513_F02.5 protei... 29 1.6
BI417964 29 2.1
CB827128 28 2.7
AV764937 28 2.7
TC17416 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate kin... 28 3.6
>TC17928
Length = 555
Score = 240 bits (613), Expect = 3e-64
Identities = 126/127 (99%), Positives = 126/127 (99%)
Frame = +3
Query: 384 RIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIV 443
RIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIV
Sbjct: 3 RIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIV 182
Query: 444 KELETLSGCTLSYKALGGRNDYKVALETLDKLQFENPLEQLGLFIKDELADEDEEVINGT 503
KEL TLSGCTLSYKALGGRNDYKVALETLDKLQFENPLEQLGLFIKDELADEDEEVINGT
Sbjct: 183 KELGTLSGCTLSYKALGGRNDYKVALETLDKLQFENPLEQLGLFIKDELADEDEEVINGT 362
Query: 504 VEEGEIL 510
VEEGEIL
Sbjct: 363 VEEGEIL 383
>AU089074
Length = 244
Score = 103 bits (256), Expect = 9e-23
Identities = 45/80 (56%), Positives = 60/80 (74%)
Frame = +1
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C +L+ LM + + W+FN PVD V + + DY+ II KPMDLGT+ + L KNVY+ EEFA
Sbjct: 4 CVQVLQKLMKHKHGWIFNVPVDAVGMGLHDYYEIIKKPMDLGTVKANLSKNVYATPEEFA 183
Query: 143 ADVRLTFSNAMTYNPPHNDV 162
+DVRLTF+NA+TYNP +DV
Sbjct: 184 SDVRLTFNNALTYNPKGHDV 243
>BP075076
Length = 527
Score = 72.0 bits (175), Expect = 2e-13
Identities = 34/75 (45%), Positives = 44/75 (58%)
Frame = -1
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C+++L+ LM + WVFN PVD L + DYF II PMDLGT+ ++L KN+
Sbjct: 242 CSSLLEKLMKHSLGWVFNTPVDVEGLGLHDYFTIITHPMDLGTVKTRLNKNLVHIT*GIC 63
Query: 143 ADVRLTFSNAMTYNP 157
FS AMTYNP
Sbjct: 62 RGCETHFSYAMTYNP 18
>BP069747
Length = 436
Score = 59.7 bits (143), Expect = 1e-09
Identities = 39/109 (35%), Positives = 54/109 (48%), Gaps = 2/109 (1%)
Frame = -1
Query: 86 ILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLG--TISSKLKKNVYSCVEEFAA 143
ILK +M W F P+DP K LG I SKLK +YS E+F
Sbjct: 433 ILKRMMIGRDGWAFKHPLDPFDSESLLMEKNKKKKPILGFVDIESKLKNWLYSEPEQFVD 254
Query: 144 DVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELGK 192
D+R FS+A+ Y P ++VH++ + L+ F+ W + KKW ED K
Sbjct: 253 DMRNVFSHALMY-PSRSEVHIIGRRLSDNFEHNWTTLKKKWASEDRKQK 110
>AI967476
Length = 397
Score = 48.9 bits (115), Expect = 2e-06
Identities = 21/39 (53%), Positives = 29/39 (73%)
Frame = +3
Query: 150 SNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCED 188
SNAMTYNP NDVH+MA L++ F+ +WK ++KK +D
Sbjct: 33 SNAMTYNPRGNDVHIMADTLSKYFEVRWKSIEKKLPRKD 149
>BU494405
Length = 563
Score = 45.4 bits (106), Expect = 2e-05
Identities = 40/149 (26%), Positives = 61/149 (40%), Gaps = 12/149 (8%)
Frame = +1
Query: 24 VDPGQKCEFGQRGLQTDENGRCNSAE------KLFKPDSIKRGPPEGFEGQKEKRQKIDR 77
+D + + +R L+ D +G+ SAE + P +RG
Sbjct: 49 LDDPRSRRYDKRMLERDRSGKRRSAELGNIGVDIMPPTKRRRGGG--------------- 183
Query: 78 KGSTQCAAILKCLMS------YPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLK 131
G A IL+ ++ + S++F KPV PDYF II +PMDL I +++
Sbjct: 184 -GEVGLANILEGIVDVLVKDRHDLSYLFQKPVSKK--EAPDYFDIIERPMDLSRIRERVR 354
Query: 132 KNVYSCVEEFAADVRLTFSNAMTYNPPHN 160
Y E+F D+ NA YN N
Sbjct: 355 NMEYKSREDFRHDIWQITYNAHKYNNGRN 441
>TC15255 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (10%)
Length = 600
Score = 42.0 bits (97), Expect = 2e-04
Identities = 30/91 (32%), Positives = 48/91 (51%), Gaps = 10/91 (10%)
Frame = +1
Query: 429 EMKRTVEIEHNLEIVKELETLSGCT----LSYKALGGRNDYKVALETLDKLQFENPLEQL 484
+M++TV+I + +++LE LSG +S+ + + L + + NPLEQL
Sbjct: 1 KMEKTVDINETSQFLEDLEMLSGVQDEPLISFAEESSPDRPQNGLGSFKRQGSFNPLEQL 180
Query: 485 GLFIKDELADEDEEVING------TVEEGEI 509
GL++K + DE+EE VEEGEI
Sbjct: 181 GLYMKVDDEDEEEEPPKNEAGPTDDVEEGEI 273
>TC9401 similar to UP|Q84LW7 (Q84LW7) Peptidylprolyl cis-trans isomerase ,
partial (97%)
Length = 918
Score = 33.1 bits (74), Expect = 0.11
Identities = 43/157 (27%), Positives = 60/157 (37%), Gaps = 22/157 (14%)
Frame = +1
Query: 290 RKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPD-----FPVQLSPKKALRA 344
RKS PD+ L V+ T A +GEV+ T F +L ++A
Sbjct: 217 RKSKPDAVAPSDDLPLVDVH------YEGTLADTGEVFDTTHEDNTIFSFELGKGSVIKA 378
Query: 345 --AMLKSRFADTILKAQQKTLLEHGDKGDP------LKMQLEKERLERIQR--------- 387
+K+ I K K +G G P + E E L R
Sbjct: 379 WDVAVKTMKVGEIAKITCKPEYAYGSAGSPPDIPPDATLVFEVELLACNPRKGSSLGSVT 558
Query: 388 EERARIEAQIKTAEAAARARAEEELRQQREKEREAAR 424
EERAR+E K E AA + EE+ +++ K AAR
Sbjct: 559 EERARLEELKKQREIAAAVKEEEKKKREEAKAAAAAR 669
>AV408585
Length = 430
Score = 33.1 bits (74), Expect = 0.11
Identities = 35/131 (26%), Positives = 60/131 (45%), Gaps = 31/131 (23%)
Frame = +2
Query: 335 QLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIE 394
+LS K+A + + ++ L+AQ + E KG K+Q+E+ ++E I+R++ A +
Sbjct: 50 ELSKKQAAQESTIRK------LRAQIRDF-EEEKKGLTTKLQIEENKVESIKRDKTATEK 208
Query: 395 ------------------------AQIKTAEAAARARAEEELRQQRE------KEREAAR 424
A K AEA A ARA E R + E +ERE+
Sbjct: 209 LLQETIEKHQNELAAQKEYYTNALAAAKEAEALAEARANNEARTELESRLREAEERESML 388
Query: 425 A-AIEEMKRTV 434
+EE+++T+
Sbjct: 389 VQTLEELRQTL 421
>TC19663 similar to UP|Q8H6S8 (Q8H6S8) Translation initiation factor,
partial (10%)
Length = 485
Score = 31.6 bits (70), Expect = 0.32
Identities = 31/110 (28%), Positives = 53/110 (48%)
Frame = +3
Query: 342 LRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAE 401
++A +++S+ ++ KA K L +H + + KE ER +REE ++
Sbjct: 126 VKAEVVESKKNESKTKAADKKLPKHVREMQEA-LARRKEAEERKKREEEEKLR------- 281
Query: 402 AAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSG 451
+ EEE R+Q E ER+A A K+ E E L+ +E + L+G
Sbjct: 282 -----KEEEERRRQEELERQAEEA--RRRKKEREKEKLLKKKQEGKLLTG 410
>TC8558 weakly similar to UP|Q9NDI0 (Q9NDI0) 200 kDa antigen p200
(Fragment), partial (4%)
Length = 585
Score = 31.6 bits (70), Expect = 0.32
Identities = 20/70 (28%), Positives = 39/70 (55%)
Frame = +1
Query: 363 LLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKEREA 422
L E D ++ ++ R ER +REE+ R+ + AEA + + EEE R++ +E
Sbjct: 67 LREEQDAAYRAALEADQAR-ERQRREEQERLARE--AAEAERKRKEEEEAREREAREAAE 237
Query: 423 ARAAIEEMKR 432
+AA+ ++++
Sbjct: 238 KQAALAKIRQ 267
>AU088817
Length = 191
Score = 31.2 bits (69), Expect = 0.42
Identities = 18/48 (37%), Positives = 25/48 (51%)
Frame = +2
Query: 409 EEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSY 456
E E ++RE+ERE R E +R E E E V L++ S TL +
Sbjct: 32 EREREREREREREREREREREREREXEREREREXVLSLDSPSPPTLCF 175
>TC15955
Length = 1042
Score = 30.8 bits (68), Expect = 0.55
Identities = 36/173 (20%), Positives = 75/173 (42%)
Frame = +1
Query: 114 FAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIF 173
+ + KP L + + ++KKN +EE + + + NA N + M ++++
Sbjct: 241 YKVTLKPA-LNSFADEIKKNSMGKLEESISLRQKSNENAARLEGKRNQLAAMQSYIDEM- 414
Query: 174 DSKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKIS 233
+++ M K+ + TG A+K + D + +V+E VLK S
Sbjct: 415 EAQMNIMKKETQ-------DYTGRCSAEAQKMLEDVQLADHDLDNMEREVAE---VLKAS 564
Query: 234 SLAARDAKVEVPKLSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTS 286
L ++A + + QVH + + K + S+ ++ + G + KR + T+
Sbjct: 565 ELKLQEATKQSEEEIQVHARKLFKLVDSVSQYQEHV--GTKVSEMKRDLSETA 717
>TC17939 weakly similar to UP|Q945P2 (Q945P2) AT5g49210/K21P3_8, partial
(61%)
Length = 999
Score = 30.8 bits (68), Expect = 0.55
Identities = 28/101 (27%), Positives = 45/101 (43%), Gaps = 7/101 (6%)
Frame = +2
Query: 327 WSTPDFPVQLSPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLE--- 383
W P P + K A L+ +A + K +K +E + M+LEK+R +
Sbjct: 269 WEAPKDPKEAEAK----LAQLRRDYAKQV-KELRKEYIEEMEL-----MRLEKQRKDEAR 418
Query: 384 ----RIQREERARIEAQIKTAEAAARARAEEELRQQREKER 420
R+ EER R++AQ A R +++ R+ KER
Sbjct: 419 REALRVATEERKRLKAQEAEIRAQQRNIEQQQFRETLLKER 541
>TC11581 weakly similar to UP|Q99736 (Q99736) HsGCN1 (Fragment), partial
(9%)
Length = 755
Score = 29.6 bits (65), Expect = 1.2
Identities = 25/95 (26%), Positives = 45/95 (47%), Gaps = 3/95 (3%)
Frame = +2
Query: 419 EREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSYKALGGRNDYKVALETLDK---L 475
ER A + E+ + I ++ ++ + C S++ R+ Y + L + +
Sbjct: 74 ERSGAAQGLSEVLAALGIGFFEHVLPDI--IRNC--SHQKASVRDGYLTLFKFLPRSLGV 241
Query: 476 QFENPLEQLGLFIKDELADEDEEVINGTVEEGEIL 510
QF+N L Q+ I D LADE+E V + + G +L
Sbjct: 242 QFQNYLSQVLPAILDGLADENESVRDAALGAGHVL 346
>TC11087 weakly similar to UP|Q8GS92 (Q8GS92) OJ1513_F02.5 protein
(OJ1118_G09.13 protein), partial (6%)
Length = 689
Score = 29.3 bits (64), Expect = 1.6
Identities = 18/55 (32%), Positives = 26/55 (46%), Gaps = 1/55 (1%)
Frame = -3
Query: 277 KQKRQIKSTSPLERK-SDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTP 330
KQ+ Q T P +D + ++SS S+H Y +TP+T SS TP
Sbjct: 603 KQRHQ*PHTEPAPHSGNDHPTCSSISSSASQHSYQAHAASTPSTQPSSSPDHRTP 439
>BI417964
Length = 433
Score = 28.9 bits (63), Expect = 2.1
Identities = 20/50 (40%), Positives = 24/50 (48%), Gaps = 2/50 (4%)
Frame = +2
Query: 379 KERLERIQREERARIEAQIKTAEAAARARAEEELRQQ--REKEREAARAA 426
+ER ER ERAR A T A RAR E + + R+KE R A
Sbjct: 2 EERKERTLERERARGAAAQATTTAV*RAREENDKGESFARKKEERGTRTA 151
>CB827128
Length = 556
Score = 28.5 bits (62), Expect = 2.7
Identities = 35/153 (22%), Positives = 64/153 (40%)
Frame = +1
Query: 347 LKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARA 406
L+SR ++T+ + + L + L+ EKE + E+ +E QIK E AR
Sbjct: 97 LQSRSSETLSANEARILEVESQLQEALQRHAEKEAEAKDLNEKLNALEGQIKLHEEQAR- 273
Query: 407 RAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSYKALGGRNDYK 466
+ E ++ +EE ++++H +V EL+ + K G N+
Sbjct: 274 --------EAVAVSETHKSGLEE--SLLKLKHLETVVDELQ--NKLLHHEKETAGLNEEN 417
Query: 467 VALETLDKLQFENPLEQLGLFIKDELADEDEEV 499
L + +E+ L L + L ++DE V
Sbjct: 418 TKLNQ-EIAIYESKLSDLQSKLSAALVEKDETV 513
>AV764937
Length = 392
Score = 28.5 bits (62), Expect = 2.7
Identities = 15/49 (30%), Positives = 28/49 (56%)
Frame = +1
Query: 398 KTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKEL 446
K AEA R A++ + ++ ++AR +++ + TV EH+L + EL
Sbjct: 238 KEAEAKQRLGADDFILSTNQQHLQSARRSLDFVSDTVSAEHSLLPLLEL 384
>TC17416 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate kinase 2,
chloroplast precursor (ATP-AMP transphosphorylase) ,
partial (36%)
Length = 537
Score = 28.1 bits (61), Expect = 3.6
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +3
Query: 22 IEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRG 60
++ D QK + L DEN E+L KPDSI +G
Sbjct: 372 VKTDSVQKTTMEKGQLVPDENVVMMGKERLLKPDSIHKG 488
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.130 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,229,826
Number of Sequences: 28460
Number of extensions: 86343
Number of successful extensions: 400
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 395
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 398
length of query: 510
length of database: 4,897,600
effective HSP length: 94
effective length of query: 416
effective length of database: 2,222,360
effective search space: 924501760
effective search space used: 924501760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0228a.2


