

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0227.11
(425 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
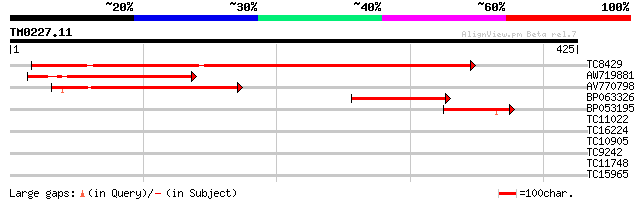
Score E
Sequences producing significant alignments: (bits) Value
TC8429 weakly similar to UP|BAD03306 (BAD03306) MAP3K-like prote... 396 e-111
AW719881 155 9e-39
AV770798 149 9e-37
BP063326 94 6e-20
BP053195 64 6e-11
TC11022 similar to UP|O81702 (O81702) Gr1-protein, partial (9%) 30 0.99
TC16224 similar to UP|O49575 (O49575) Receptor kinase - like pro... 30 0.99
TC10905 similar to UP|Q94BV2 (Q94BV2) AT5g58570/mzn1_20, partial... 28 2.2
TC9242 similar to PIR|B86320|B86320 3-phosphoserine phosphatase ... 27 4.9
TC11748 weakly similar to UP|O82076 (O82076) Lipase homolog (Fra... 27 6.4
TC15965 homologue to UP|Q8S902 (Q8S902) Syringolide-induced prot... 27 8.4
>TC8429 weakly similar to UP|BAD03306 (BAD03306) MAP3K-like protein kinase,
partial (55%)
Length = 1298
Score = 396 bits (1018), Expect = e-111
Identities = 196/336 (58%), Positives = 252/336 (74%), Gaps = 3/336 (0%)
Frame = +3
Query: 17 ILVLLDSSLALK-FKPLQQEGQICVANKN-CNSGLHCETCVTNGNVRPRCTRTQPINPTS 74
I L+ +SL + L Q + C + N C+ GL C C + RCTR + I+PTS
Sbjct: 102 IATLVSASLVFGCYYILVQIAETCSRDINDCDLGLQCLECHSQN----RCTRIRTISPTS 269
Query: 75 KVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDF 134
KV LPFN YSWLTTHNSFA G TGS +L+ TNQ+D+ITDQL NGVRGLMLD++D+
Sbjct: 270 KVMELPFNEYSWLTTHNSFAAKGVNWSTGSPVLAFTNQEDSITDQLKNGVRGLMLDMWDY 449
Query: 135 ENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPKGLTKVF 194
E+ +WLC G C YT FQPA+NVLKE++VFL +P+EI+TI I+D+VTS G+ KVF
Sbjct: 450 EDTIWLCR---GPCTKYTTFQPALNVLKEVRVFLVTHPTEIITIFIDDHVTSGNGVNKVF 620
Query: 195 DAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVEN 254
D A LRK+WFPVS+MPKNG DWP V M++KN RL+VFTS AS+EASEGIAYEW Y+VE+
Sbjct: 621 DKARLRKFWFPVSKMPKNGSDWPTVKTMIRKNYRLIVFTSNASREASEGIAYEWNYVVES 800
Query: 255 QYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPD-VAQSCKDNSAPLLSMVSTCN 313
Q+GN G+K GSC NR ES MN ++SLVL+N+FR++ + ++C+DNS+PL++M+ C
Sbjct: 801 QFGNVGIKGGSCQNRPESLPMNNATKSLVLMNYFRNVRNHDDEACRDNSSPLIAMMHVCF 980
Query: 314 QAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLV 349
+AAG RWPNFIAVDFYKR DGGGAPEA+D+AN +L+
Sbjct: 981 RAAGNRWPNFIAVDFYKRGDGGGAPEALDLANRNLI 1088
>AW719881
Length = 495
Score = 155 bits (393), Expect = 9e-39
Identities = 74/127 (58%), Positives = 89/127 (69%)
Frame = +1
Query: 14 LFTILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPT 73
L IL L S + K G+ C +C+ GL C+TC NGN RPRC+R QP NPT
Sbjct: 142 LIAILCLFTCSSSSKI------GETC---GSCDGGLTCQTCPANGNTRPRCSRIQPSNPT 294
Query: 74 SKVKGLPFNRYSWLTTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYD 133
+KVKGLPFNRYSWLT+HNSFA G +S TGS I++P NQ DT+ QL NGVRG MLD+YD
Sbjct: 295 TKVKGLPFNRYSWLTSHNSFAQAGARSATGSFIIAPMNQDDTVAQQLKNGVRGFMLDMYD 474
Query: 134 FENDVWL 140
F+ND+WL
Sbjct: 475 FQNDIWL 495
>AV770798
Length = 503
Score = 149 bits (376), Expect = 9e-37
Identities = 73/147 (49%), Positives = 102/147 (68%), Gaps = 4/147 (2%)
Frame = +1
Query: 32 LQQEGQI---CVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKG-LPFNRYSWL 87
LQ + Q+ C ++ +C SGL+C +C + R RC R+ + + LPFN+Y++L
Sbjct: 64 LQWKSQLLEECSSDDDCGSGLYCFSCGAKFS-RSRCVRSSITDQFKLINNSLPFNKYAFL 240
Query: 88 TTHNSFALLGQKSMTGSVILSPTNQQDTITDQLNNGVRGLMLDLYDFENDVWLCHSFGGQ 147
TTHN+FA+ G+ S TG L+ NQ+D++T QL NG RGLMLD YD++ DVWLCHSF G+
Sbjct: 241 TTHNAFAMDGEPSHTGVPRLTIINQEDSVTQQLKNGGRGLMLDTYDYDGDVWLCHSFQGK 420
Query: 148 CYNYTAFQPAINVLKEIQVFLEANPSE 174
C++YTAF+PAI+ LKE+ FL ANP E
Sbjct: 421 CHDYTAFEPAIDTLKEVAAFLSANPKE 501
>BP063326
Length = 492
Score = 93.6 bits (231), Expect = 6e-20
Identities = 43/75 (57%), Positives = 59/75 (78%), Gaps = 1/75 (1%)
Frame = -3
Query: 257 GNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVA-QSCKDNSAPLLSMVSTCNQA 315
GN G+K GSC NR ES MN ++SLVL+N+FR++ + ++C+DNS+PL++M+ C +A
Sbjct: 394 GNVGIKGGSCQNRPESLPMNNATKSLVLMNYFRNVRNHDDEACRDNSSPLIAMMHVCFRA 215
Query: 316 AGKRWPNFIAVDFYK 330
AG RWPNFIAVDFYK
Sbjct: 214 AGNRWPNFIAVDFYK 170
>BP053195
Length = 417
Score = 63.5 bits (153), Expect = 6e-11
Identities = 32/60 (53%), Positives = 39/60 (64%), Gaps = 7/60 (11%)
Frame = -2
Query: 326 VDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCKVRLT-------PITFIPPHTHLNLHV 378
VD+Y RSDGGGAPEAVDVA+GHL CGC ++A C+ T PI+ PP + HV
Sbjct: 416 VDYYPRSDGGGAPEAVDVASGHLACGCDHLAYCQGNGTFGSCDVPPISPPPPASEATPHV 237
>TC11022 similar to UP|O81702 (O81702) Gr1-protein, partial (9%)
Length = 514
Score = 29.6 bits (65), Expect = 0.99
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = -3
Query: 366 TFIPPHTHLNLHVLNFDLCRKTWDLEHVNYLKLKQLLNM 404
TF+ H H +L +N + W L +L LKQ +NM
Sbjct: 419 TFLNKHGHSSLATMNLPFYMRKW*LLFPPFLNLKQQINM 303
>TC16224 similar to UP|O49575 (O49575) Receptor kinase - like protein
(Receptor kinase-like protein), partial (3%)
Length = 446
Score = 29.6 bits (65), Expect = 0.99
Identities = 15/50 (30%), Positives = 26/50 (52%)
Frame = +1
Query: 210 PKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEWRYLVENQYGNG 259
PK+GG WP++ + Q L + ++ EG + WR+ + N + NG
Sbjct: 202 PKHGGGWPELRQGTGEIQLLT--DLQPPEQVLEGETWFWRFCLWNGWENG 345
>TC10905 similar to UP|Q94BV2 (Q94BV2) AT5g58570/mzn1_20, partial (76%)
Length = 955
Score = 28.5 bits (62), Expect = 2.2
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 1/56 (1%)
Frame = +3
Query: 50 HCETCVTNGN-VRPRCTRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSMTGS 104
H E C+ G R + TR+ +G+P +RYS + H+S ++ +S GS
Sbjct: 402 HLEKCMGKGRKARLKVTRSSTATQKRYSRGIPASRYSSYSNHSSNSI--NRSANGS 563
>TC9242 similar to PIR|B86320|B86320 3-phosphoserine phosphatase [imported]
- Arabidopsis thaliana {Arabidopsis
thaliana;}, partial (35%)
Length = 516
Score = 27.3 bits (59), Expect = 4.9
Identities = 14/40 (35%), Positives = 18/40 (45%), Gaps = 2/40 (5%)
Frame = +2
Query: 347 HLVCGCGN--IATCKVRLTPITFIPPHTHLNLHVLNFDLC 384
HL+CGC CK L F PH + + V F +C
Sbjct: 221 HLLCGCSTS*SCCCKS*LASFRF*RPHKFVGVTVFFFFIC 340
>TC11748 weakly similar to UP|O82076 (O82076) Lipase homolog (Fragment),
partial (48%)
Length = 679
Score = 26.9 bits (58), Expect = 6.4
Identities = 14/36 (38%), Positives = 17/36 (46%), Gaps = 4/36 (11%)
Frame = +3
Query: 132 YDFENDVWLCHSFGGQCYNY----TAFQPAINVLKE 163
Y F N + +C FGG YN+ T QP V E
Sbjct: 288 YGFSNPLTVCCGFGGPPYNFDLRVTCGQPGYQVCDE 395
>TC15965 homologue to UP|Q8S902 (Q8S902) Syringolide-induced protein 19-1-5,
partial (86%)
Length = 1021
Score = 26.6 bits (57), Expect = 8.4
Identities = 13/40 (32%), Positives = 19/40 (47%)
Frame = -2
Query: 47 SGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSW 86
S HC C++ + P+ T +NPT V + R SW
Sbjct: 882 SWYHCNNCISCNSSAPKATCLDLLNPT-LVTATHWMRQSW 766
Score = 26.6 bits (57), Expect = 8.4
Identities = 19/59 (32%), Positives = 29/59 (48%), Gaps = 13/59 (22%)
Frame = +1
Query: 8 LAIATTLFTIL---------VLLDSSLALKFKPL----QQEGQICVANKNCNSGLHCET 53
LA +T + T+L ++ SL++ KPL E IC+A C+S L+ ET
Sbjct: 127 LATSTRISTLLGEMVVPRYSTMVTFSLSILTKPLGLASSPEMNICLARLTCSSSLYLET 303
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,126,667
Number of Sequences: 28460
Number of extensions: 119089
Number of successful extensions: 584
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 579
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 580
length of query: 425
length of database: 4,897,600
effective HSP length: 93
effective length of query: 332
effective length of database: 2,250,820
effective search space: 747272240
effective search space used: 747272240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0227.11


