

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.9
(223 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
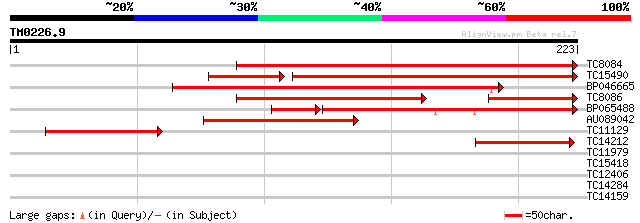
Score E
Sequences producing significant alignments: (bits) Value
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 176 3e-45
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 142 8e-42
BP046665 161 1e-40
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 92 2e-28
BP065488 86 1e-19
AU089042 66 5e-12
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 52 1e-07
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 42 1e-04
TC11979 similar to UP|Q9LVH4 (Q9LVH4) GTPase activating protein-... 28 1.3
TC15418 weakly similar to UP|Q8GT65 (Q8GT65) Serpin-like protein... 28 1.3
TC12406 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, p... 27 2.2
TC14284 similar to GB|BAB03165.1|11994775|AP002050 membrane impo... 26 6.3
TC14159 homologue to UP|ATP2_HEVBR (P29685) ATP synthase beta ch... 25 8.2
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 176 bits (446), Expect = 3e-45
Identities = 90/134 (67%), Positives = 106/134 (78%)
Frame = +1
Query: 90 IDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSKFPIKFRIRQF 149
I + +LCNGTRLIV +LG I ATVITGTN+ D + I R+D+V SDS +P KF R F
Sbjct: 43 IQKFENLCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 222
Query: 150 PIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGKQQ 209
PI++CFAMTINKSQGQ+LS V L+L RPVFTHGQLYVALSRVRSRKGLKL +LDE K
Sbjct: 223 PISLCFAMTINKSQGQSLSHVSLYLSRPVFTHGQLYVALSRVRSRKGLKLLVLDEEEKVT 402
Query: 210 TSTVNVVFKEVFEN 223
+T NVV++EVFEN
Sbjct: 403 NTTKNVVYREVFEN 444
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 142 bits (359), Expect(2) = 8e-42
Identities = 72/112 (64%), Positives = 87/112 (77%)
Frame = +2
Query: 112 IGATVITGTNVNDKVHILRMDLVLSDSKFPIKFRIRQFPIAICFAMTINKSQGQTLSQVG 171
I TVITGT++ D + I RMD+V SDS +P KF RQ PI++CFAMTINKSQG++LS VG
Sbjct: 137 IQVTVITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVG 316
Query: 172 LFLPRPVFTHGQLYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVFKEVFEN 223
L+L RPV THG LYVAL RVRSRK LKL +LDE K +T NVV++E+FEN
Sbjct: 317 LYLSRPVSTHG*LYVALPRVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFEN 472
Score = 43.5 bits (101), Expect(2) = 8e-42
Identities = 22/30 (73%), Positives = 24/30 (79%)
Frame = +1
Query: 79 KVGTPIMLLRNIDQAASLCNGTRLIVDELG 108
K G IMLLRNI QA+ LCNGTRLIV +LG
Sbjct: 43 KEGALIMLLRNIVQASGLCNGTRLIVVDLG 132
>BP046665
Length = 524
Score = 161 bits (407), Expect = 1e-40
Identities = 83/131 (63%), Positives = 99/131 (75%), Gaps = 1/131 (0%)
Frame = -1
Query: 65 DTKSSGMPNHQLLLKVGTPIMLLRNIDQAASLCNGTRLIVDELGERFIGATVITGTNVND 124
D G+PNH++ LK G PIMLLRNI QA CNGTRLIV +LG I ATVIT TN+ D
Sbjct: 524 DITCFGIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGD 345
Query: 125 KVHILRMDLVLSDSKFPIKFRIRQFPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQL 184
+ I RMD+V SDS +P KF RQFPI++C AMTINKSQGQ+LS VGL+L R VFTHGQL
Sbjct: 344 DIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHVGLYLSRHVFTHGQL 165
Query: 185 YVAL-SRVRSR 194
YVAL ++++ R
Sbjct: 164 YVALHAKIKKR 132
Score = 37.7 bits (86), Expect = 0.002
Identities = 19/40 (47%), Positives = 26/40 (64%)
Frame = -2
Query: 178 VFTHGQLYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVF 217
+++H LS +RSRKGLKL +LDE K +T NVV+
Sbjct: 187 MYSHMVSCTLLSMLRSRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 92.4 bits (228), Expect(2) = 2e-28
Identities = 46/75 (61%), Positives = 57/75 (75%)
Frame = +1
Query: 90 IDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSKFPIKFRIRQF 149
I + +LC+GTRLIV +LG I ATVITGTN+ D + I R+D+V SDS +P KF R F
Sbjct: 151 IQKFENLCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*F 330
Query: 150 PIAICFAMTINKSQG 164
PI++CFAMTINKSQG
Sbjct: 331 PISLCFAMTINKSQG 375
Score = 49.3 bits (116), Expect(2) = 2e-28
Identities = 24/35 (68%), Positives = 28/35 (79%)
Frame = +3
Query: 189 SRVRSRKGLKLCILDEAGKQQTSTVNVVFKEVFEN 223
SRVRSRKGLKL +LDE K +T NVV++EVFEN
Sbjct: 369 SRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFEN 473
>BP065488
Length = 439
Score = 85.5 bits (210), Expect(2) = 1e-19
Identities = 52/102 (50%), Positives = 66/102 (63%), Gaps = 2/102 (1%)
Frame = -1
Query: 124 DKVHILRMDLVLSDSKFPIKFRIRQFPIAICFAMTINKSQGQT-LSQVGLFLPRPVFTH- 181
D + I RM++V S S P+KF QFPI++CFAMTINKSQ V L ++H
Sbjct: 376 DDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQXSVHYPTVALISFLACYSHM 197
Query: 182 GQLYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVFKEVFEN 223
+LYVALS VRSRKGLKL +L E K +T N V++EVF+N
Sbjct: 196 DKLYVALSGVRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKN 71
Score = 26.6 bits (57), Expect(2) = 1e-19
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = -3
Query: 104 VDELGERFIGATVITGTNV 122
V +LG I ATVITGTN+
Sbjct: 437 VPDLGTNVIKATVITGTNI 381
>AU089042
Length = 191
Score = 65.9 bits (159), Expect = 5e-12
Identities = 31/61 (50%), Positives = 43/61 (69%)
Frame = +3
Query: 77 LLKVGTPIMLLRNIDQAASLCNGTRLIVDELGERFIGATVITGTNVNDKVHILRMDLVLS 136
+LK G P+ML+ N+ + LCNGTRLIVD LG IGAT+++GT++ V+I M+L S
Sbjct: 6 VLKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPS 185
Query: 137 D 137
D
Sbjct: 186 D 188
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 51.6 bits (122), Expect = 1e-07
Identities = 23/46 (50%), Positives = 33/46 (71%)
Frame = +2
Query: 15 PTLEAVEMLNTYMLGMIAAPEHEYLSSDSALRSDEDSEIKVEWFTT 60
PTLE+VE +N +ML ++ EYLSSD+ + DED+E++ WFTT
Sbjct: 458 PTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDEDTELQS*WFTT 595
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 41.6 bits (96), Expect = 1e-04
Identities = 19/39 (48%), Positives = 28/39 (71%)
Frame = +2
Query: 184 LYVALSRVRSRKGLKLCILDEAGKQQTSTVNVVFKEVFE 222
LYVA+SRV+S+ GLK+ I + +T N+V+KEVF+
Sbjct: 2 LYVAVSRVKSKDGLKILISSDGTSTPGATKNIVYKEVFQ 118
>TC11979 similar to UP|Q9LVH4 (Q9LVH4) GTPase activating protein-like,
partial (94%)
Length = 711
Score = 28.1 bits (61), Expect = 1.3
Identities = 16/66 (24%), Positives = 32/66 (48%)
Frame = +1
Query: 138 SKFPIKFRIRQFPIAICFAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGL 197
S+ P+ R R FP +C+ N + + ++ P+ + + + L RVR ++G+
Sbjct: 85 SQVPLPHRDRHFPYWLCYYYNHNIKREERKKEMEDSTPKSLMEN---LLGLLRVRVKRGV 255
Query: 198 KLCILD 203
L + D
Sbjct: 256 NLAVRD 273
>TC15418 weakly similar to UP|Q8GT65 (Q8GT65) Serpin-like protein
(Fragment), partial (44%)
Length = 602
Score = 28.1 bits (61), Expect = 1.3
Identities = 21/65 (32%), Positives = 30/65 (45%), Gaps = 6/65 (9%)
Frame = +1
Query: 162 SQGQTLSQVGLFLPRPVFTHGQLY------VALSRVRSRKGLKLCILDEAGKQQTSTVNV 215
S+G TL Q+ FL H + V LS S G +LC D +Q+ T+N
Sbjct: 151 SEGPTLHQLLTFLRSNSTDHLNSFASQLVSVVLSDASSVGGPRLCFADGVWVEQSLTLNP 330
Query: 216 VFKEV 220
FK++
Sbjct: 331 SFKQL 345
>TC12406 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial (6%)
Length = 450
Score = 27.3 bits (59), Expect = 2.2
Identities = 15/38 (39%), Positives = 19/38 (49%)
Frame = -3
Query: 118 TGTNVNDKVHILRMDLVLSDSKFPIKFRIRQFPIAICF 155
T T V + ILR++L LS FP FP+ CF
Sbjct: 325 TTTTVGSRHRILRLNLNLSHQNFP------NFPLFFCF 230
>TC14284 similar to GB|BAB03165.1|11994775|AP002050 membrane import
protein-like {Arabidopsis thaliana;} , partial (36%)
Length = 702
Score = 25.8 bits (55), Expect = 6.3
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +2
Query: 86 LLRNIDQAASLCNGTRLIVDELGERF 111
L + I + ASLC T+ I+ EL E+F
Sbjct: 413 LKKEITEEASLCRETQFIMHELLEKF 490
>TC14159 homologue to UP|ATP2_HEVBR (P29685) ATP synthase beta chain,
mitochondrial precursor , partial (93%)
Length = 2056
Score = 25.4 bits (54), Expect = 8.2
Identities = 13/31 (41%), Positives = 16/31 (50%), Gaps = 9/31 (29%)
Frame = +2
Query: 134 VLSDSKFP---------IKFRIRQFPIAICF 155
++SDS FP +FRI FPI CF
Sbjct: 65 IVSDSIFPPSISIQTLHFRFRIADFPIPTCF 157
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,201,540
Number of Sequences: 28460
Number of extensions: 37239
Number of successful extensions: 144
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 143
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 143
length of query: 223
length of database: 4,897,600
effective HSP length: 87
effective length of query: 136
effective length of database: 2,421,580
effective search space: 329334880
effective search space used: 329334880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0226.9


