

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.8
(518 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
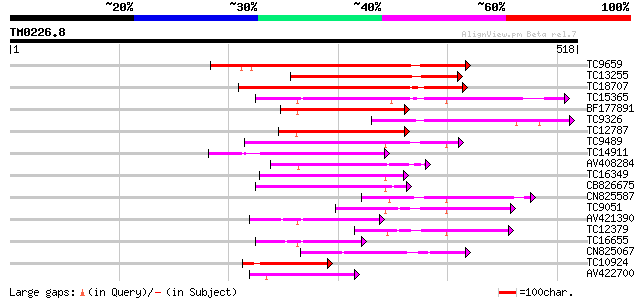
Sequences producing significant alignments: (bits) Value
TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATP... 236 9e-63
TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%) 203 5e-53
TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J... 184 4e-47
TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {... 161 3e-40
BF177891 130 7e-31
TC9326 homologue to UP|Q93WX4 (Q93WX4) Suppressor of K+ transpor... 127 3e-30
TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (T... 127 3e-30
TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase... 121 3e-28
TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), pa... 114 3e-26
AV408284 112 2e-25
TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulat... 105 2e-23
CB826675 102 2e-22
CN825587 91 3e-19
TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 ... 87 9e-18
AV421390 81 5e-16
TC12379 homologue to UP|O04327 (O04327) Cell division protein Ft... 79 1e-15
TC16655 homologue to UP|O99018 (O99018) Chloroplast protease pre... 77 7e-15
CN825067 67 9e-12
TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPas... 66 2e-11
AV422700 62 3e-10
>TC9659 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (74%)
Length = 1060
Score = 236 bits (601), Expect = 9e-63
Identities = 120/243 (49%), Positives = 168/243 (68%), Gaps = 5/243 (2%)
Frame = +1
Query: 184 GVSGKAGKTDSVISDAEESKSKKSQYE--GPDPELAEM---LERDVLETSPGVRWEDVAG 238
G G A D+ ++ ++K K G DPE A++ L ++ P ++W DVAG
Sbjct: 298 GGPGPASNGDAAVAARPKTKPKDGGEGDGGEDPEQAKLRAGLNSAIIREKPNIKWNDVAG 477
Query: 239 LTEAKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNV 298
L AK+ L+EAV+LP+ P++F G RRPW+ L++GPPGTGK+ LAKAVATE +TFF+V
Sbjct: 478 LESAKQSLQEAVILPVKFPQFFTGKRRPWRAFLLYGPPGTGKSYLAKAVATEADSTFFSV 657
Query: 299 SSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSEL 358
SS+ L SKW GESE++V LF +AR APS IF+DEIDSLC RG E E+SRR+K+EL
Sbjct: 658 SSSDLGSKWMGESEKLVSNLFQMARESAPSIIFVDEIDSLCGQRGEGNESEASRRIKTEL 837
Query: 359 LVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESRKELIR 418
LVQ+ GV N + + V+VLAATN P+ +D+A+RRR +KRIYIPLP+ ++R+ + +
Sbjct: 838 LVQMQGVGN-------NDQKVLVLAATNTPYALDQAIRRRFDKRIYIPLPDVKARQHMFK 996
Query: 419 INL 421
++L
Sbjct: 997 VHL 1005
>TC13255 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (41%)
Length = 532
Score = 203 bits (517), Expect = 5e-53
Identities = 96/158 (60%), Positives = 129/158 (80%), Gaps = 1/158 (0%)
Frame = +1
Query: 257 PEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVR 316
P+YF G+ PWKG+L+FGPPGTGKT+LAKAVATEC TTFFN+S++++ SKWRG+SE++V+
Sbjct: 4 PKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVK 183
Query: 317 CLFDLARAYAPSTIFIDEIDSLCNSRG-ASGEHESSRRVKSELLVQVDGVSNTATNEDGS 375
LF LAR +APSTIF+DEID++ + RG A EHE+SRR+K+ELL+Q+DG++ T
Sbjct: 184 VLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRT------- 342
Query: 376 RKIVMVLAATNFPWDIDEALRRRLEKRIYIPLPNFESR 413
++V VLAATN PW++D A+ RRLEKRI +PLP E+R
Sbjct: 343 DELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEAR 456
>TC18707 homologue to PIR|T02901|T02901 MSP1 protein homolog T13J8.110 -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(29%)
Length = 771
Score = 184 bits (466), Expect = 4e-47
Identities = 95/211 (45%), Positives = 141/211 (66%), Gaps = 2/211 (0%)
Frame = +3
Query: 210 EGPDPELAEMLERDVLETSP-GVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQG-IRRPW 267
E PD E + + +V+ + GV + D+ + E K L+E V+LPL P+ F+G + +P
Sbjct: 126 EVPDNEFEKRIRPEVIPANEIGVTFGDIGAMDEIKESLQELVMLPLRRPDLFKGGLLKPC 305
Query: 268 KGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAP 327
+G+L+FGPPGTGKT+LA A+A E G +F NVS +T+ SKW GE E+ VR LF LA AP
Sbjct: 306 RGILLFGPPGTGKTMLAXAIANEAGASFINVSMSTITSKWFGEDEKNVRALFSLAAKVAP 485
Query: 328 STIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNF 387
+ IF+DE+DS+ R GEHE+ R++K+E + DG+ TA E ++VLAATN
Sbjct: 486 TIIFVDEVDSMLGQRTRVGEHEAMRKIKNEFMTHWDGLL-TAPGEQ-----ILVLAATNR 647
Query: 388 PWDIDEALRRRLEKRIYIPLPNFESRKELIR 418
P+D+DEA+ RR E+RI + LP+ E+R+ +++
Sbjct: 648 PFDLDEAIIRRFERRIMVGLPSVENREMILK 740
>TC15365 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;}, partial (87%)
Length = 1385
Score = 161 bits (407), Expect = 3e-40
Identities = 104/294 (35%), Positives = 168/294 (56%), Gaps = 7/294 (2%)
Frame = +3
Query: 225 LETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTL 282
+E P ++ + GL + + ++E + LP+ PE F+ GI +P KGVL++GPPGTGKTL
Sbjct: 300 VEKVPDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQP-KGVLLYGPPGTGKTL 476
Query: 283 LAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR 342
LA+AVA TF VS + L K+ GE RMVR LF +AR +APS IF+DEIDS+ ++R
Sbjct: 477 LARAVAHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSAR 656
Query: 343 GASGE---HESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR-- 397
SG +R ELL Q+DG A+N+ + VL ATN +D+AL R
Sbjct: 657 MESGSGXGDSEVQRTMLELLNQLDGFE--ASNK------IKVLMATNRIDILDQALLRPG 812
Query: 398 RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLN 457
R++++I P PN ESR+++++I+ + + + +++ ++A + +G SG +L VC +A +
Sbjct: 813 RIDRKIEFPNPNEESRQDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMF 992
Query: 458 GMRRKIAGKTRDEIKNMSKDDISKDPVAMCDFEEALVKVQRSVSQADIERHEKW 511
+R + T++ DFE A+ KV + ++ ++ + W
Sbjct: 993 ALRERRVHVTQE------------------DFEMAVAKVMKKETEKNMSLRKLW 1100
>BF177891
Length = 390
Score = 130 bits (326), Expect = 7e-31
Identities = 61/120 (50%), Positives = 86/120 (70%), Gaps = 2/120 (1%)
Frame = +1
Query: 248 EAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLAS 305
E V+LP+ PE F + RP KG+L+FGPPGTGKTL+AKA+ATE G F +VS +TL S
Sbjct: 16 ELVILPMRRPELFTHGNLLRPVKGILLFGPPGTGKTLMAKALATEAGANFISVSGSTLTS 195
Query: 306 KWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGV 365
KW G++E++ + LF A AP IF+DE+DSL +RG + EHE++RR+++E + DG+
Sbjct: 196 KWFGDAEKLTKALFSFASKLAPVIIFVDEVDSLLGARGGAFEHEATRRMRNEFMAAWDGL 375
>TC9326 homologue to UP|Q93WX4 (Q93WX4) Suppressor of K+ transport growth
defect-like protein (Fragment), partial (73%)
Length = 894
Score = 127 bits (320), Expect = 3e-30
Identities = 73/211 (34%), Positives = 120/211 (56%), Gaps = 25/211 (11%)
Frame = +2
Query: 331 FIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWD 390
F+DEIDSLC RG E E+SRR+K+ELLVQ+ GV N V+VLAATN P+
Sbjct: 20 FVDEIDSLCGQRGEGNESEASRRIKTELLVQMQGVGNNDQK-------VLVLAATNTPYA 178
Query: 391 IDEALRRRLEKRIYIPLPNFESRKELIRINL-KTVEVAPDVNIDEVARRTDGYSGDDLTN 449
+D+A+RRR +KRIYIPLP+ ++R+ + +++L T + + + +A RT+G+SG D++
Sbjct: 179 LDQAIRRRFDKRIYIPLPDLKARQHMFKVHLGDTPHNLNESDFEYLASRTEGFSGSDISV 358
Query: 450 VCRDASLNGMRR--------------------KIAGKTRDEIKNMSKDDISKD----PVA 485
+D +R+ K G + +++++ ++ P+
Sbjct: 359 CVKDVLFEPVRKTQDAMFFFKNPEGMWIPCGPKQQGAVQITMQDLAAKGLASQILPPPIT 538
Query: 486 MCDFEEALVKVQRSVSQADIERHEKWFHEFG 516
DFE+ L + + +VS+ D+E HE++ EFG
Sbjct: 539 RTDFEKVLARQRPTVSKKDLEVHERFTKEFG 631
>TC12787 weakly similar to UP|MSP1_YEAST (P28737) MSP1 protein (TAT-binding
homolog 4), partial (20%)
Length = 389
Score = 127 bits (320), Expect = 3e-30
Identities = 61/122 (50%), Positives = 85/122 (69%), Gaps = 2/122 (1%)
Frame = +3
Query: 246 LEEAVVLPLWMPEYF--QGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATL 303
L+E V+LPL PE F + +P KG+L+FGPPGTGKT+LAKAVATE G F N+S +++
Sbjct: 12 LKELVMLPLKRPELFCKGQLTKPCKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSI 191
Query: 304 ASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVD 363
SKW GE E+ V+ +F LA +PS IF+DE+D + R GEHE+ R++ +E +V D
Sbjct: 192 TSKWFGEGEKYVKAVFSLASKISPSVIFVDEVDXMLXRRENPGEHEAMRKMXNEFMVHWD 371
Query: 364 GV 365
G+
Sbjct: 372 GL 377
>TC9489 homologue to UP|Q9SEI5 (Q9SEI5) 26S proteasome AAA-ATPase subunit
RPT2a, partial (69%)
Length = 916
Score = 121 bits (303), Expect = 3e-28
Identities = 80/205 (39%), Positives = 117/205 (57%), Gaps = 5/205 (2%)
Frame = +1
Query: 215 ELAEMLERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMF 273
E+ M+ +E +P + D+ GL + ++EAV LPL PE ++ I +P KGV+++
Sbjct: 325 EVDPMVSVMKVEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILY 504
Query: 274 GPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFID 333
G PGTGKTLLAKAVA TF V + L K+ G+ ++VR LF +A +PS +FID
Sbjct: 505 GEPGTGKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFID 684
Query: 334 EIDSLCNSR--GASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDI 391
EID++ R SG +R ELL Q+DG SR V V+ ATN +
Sbjct: 685 EIDAVGTKRYDAHSGGEREIQRTMLELLNQLDGFD--------SRGDVKVILATNRIESL 840
Query: 392 DEALRR--RLEKRIYIPLPNFESRK 414
D AL R R++++I PLP+ ++R+
Sbjct: 841 DPALLRPGRIDRKIEFPLPDTKTRR 915
>TC14911 homologue to UP|Q93X55 (Q93X55) Peroxin 6 (Fragment), partial (32%)
Length = 1418
Score = 114 bits (286), Expect = 3e-26
Identities = 68/166 (40%), Positives = 92/166 (54%)
Frame = +2
Query: 182 KSGVSGKAGKTDSVISDAEESKSKKSQYEGPDPELAEMLERDVLETSPGVRWEDVAGLTE 241
KS VS + + V++ E SK + + G P++ P V+WEDV GL +
Sbjct: 107 KSKVSPRIPGKEDVMNALERSKKRNASALGT-PKV------------PNVKWEDVGGLED 247
Query: 242 AKRLLEEAVVLPLWMPEYFQGIRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSA 301
K+ + + V LPL + F R GVL++GPPGTGKTLLAKAVATEC F +V
Sbjct: 248 VKKSILDTVQLPLLHKDLFASGLRKRSGVLLYGPPGTGKTLLAKAVATECSLNFLSVKGP 427
Query: 302 TLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGE 347
L + + GESE+ VR +F AR+ P IF DE+DSL + G E
Sbjct: 428 ELINMYIGESEKNVRDIFQKARSARPCVIFFDELDSLAPASGMLSE 565
>AV408284
Length = 425
Score = 112 bits (279), Expect = 2e-25
Identities = 65/148 (43%), Positives = 89/148 (59%), Gaps = 2/148 (1%)
Frame = +1
Query: 239 LTEAKRLLEEAVVLPLWMPEYFQG--IRRPWKGVLMFGPPGTGKTLLAKAVATECGTTFF 296
L K L E V+LPL P+ F + P KGVL++GPPGTGKT+LAKA+A E G F
Sbjct: 1 LESIKEALFELVILPLKRPDLFSHGKLLGPQKGVLLYGPPGTGKTMLAKAIAKESGAVFI 180
Query: 297 NVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSRGASGEHESSRRVKS 356
NV + L SKW G+++++V +F LA P+ IFIDE+DS R + +HE+ +K+
Sbjct: 181 NVRISNLMSKWFGDAQKLVAAVFSLAHKLQPAIIFIDEVDSFLGQRRTT-DHEALLNMKT 357
Query: 357 ELLVQVDGVSNTATNEDGSRKIVMVLAA 384
E + DG T + +R VMVLAA
Sbjct: 358 EFMALWDGF----TTDQNAR--VMVLAA 423
>TC16349 homologue to UP|PRS6_SOLTU (P54778) 26S protease regulatory subunit
6B homolog, partial (65%)
Length = 901
Score = 105 bits (262), Expect = 2e-23
Identities = 60/139 (43%), Positives = 83/139 (59%), Gaps = 3/139 (2%)
Frame = +3
Query: 229 PGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGKTLLAKAV 287
P V + D+ G K+ + EAV LPL E ++ I P +GVL++GPPGTGKT+LAKAV
Sbjct: 483 PDVTYNDIGGCDIQKQEIREAVELPLTHHELYKQIGIDPPRGVLLYGPPGTGKTMLAKAV 662
Query: 288 ATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR--GAS 345
A F V + K+ GE RMVR +F LA+ AP+ IFIDE+D++ +R +
Sbjct: 663 ANHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLAKENAPAIIFIDEVDAIATARFDAQT 842
Query: 346 GEHESSRRVKSELLVQVDG 364
G +R+ ELL Q+DG
Sbjct: 843 GADREVQRILMELLNQMDG 899
>CB826675
Length = 551
Score = 102 bits (253), Expect = 2e-22
Identities = 61/147 (41%), Positives = 88/147 (59%), Gaps = 4/147 (2%)
Frame = +1
Query: 225 LETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGVLMFGPPGTGKTLL 283
++ P + DV GL + + L EA+VLP+ E FQ + RP KGVL++GPPGTGKTL+
Sbjct: 109 VDEKPTEDYNDVGGLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLM 288
Query: 284 AKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR- 342
A+A A + TF ++ L + G+ ++VR F LA+ +P IFIDEID++ R
Sbjct: 289 ARACAAQTNATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRF 468
Query: 343 --GASGEHESSRRVKSELLVQVDGVSN 367
SG+ E +R ELL Q+DG S+
Sbjct: 469 DSEVSGDRE-VQRTMLELLNQLDGFSS 546
>CN825587
Length = 663
Score = 91.3 bits (225), Expect = 3e-19
Identities = 55/163 (33%), Positives = 97/163 (58%), Gaps = 4/163 (2%)
Frame = +2
Query: 322 ARAYAPSTIFIDEIDSLCNSRGAS--GEHESSRRVKSELLVQVDGVSNTATNEDGSRKIV 379
A++ AP +FIDEID++ RGA G ++ + ++LL ++DG S + V
Sbjct: 2 AKSKAPCIVFIDEIDAVGRQRGAGLGGGNDEREQTINQLLTEMDGFSGNSG--------V 157
Query: 380 MVLAATNFPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDVNIDEVAR 437
+VLAATN P +D AL R R ++++ + P+ R ++++++ + +A DV+ D++AR
Sbjct: 158 IVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRVKILQVHSRGKALAKDVDFDKIAR 337
Query: 438 RTDGYSGDDLTNVCRDASLNGMRRKIAGKTRDEIKNMSKDDIS 480
RT G++G DL N+ +A++ RR ++K +SKD+IS
Sbjct: 338 RTPGFTGADLQNLMNEAAILAARR--------DLKEISKDEIS 442
>TC9051 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (46%)
Length = 867
Score = 86.7 bits (213), Expect = 9e-18
Identities = 54/170 (31%), Positives = 94/170 (54%), Gaps = 5/170 (2%)
Frame = +2
Query: 298 VSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDSLCNSR---GASGEHESSRRV 354
V + L K+ GE RMVR LF +AR+ +F DE+D++ +R G G++E R +
Sbjct: 8 VIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAIGGARFDDGVGGDNEVQRTM 187
Query: 355 KSELLVQVDGVSNTATNEDGSRKIVMVLAATNFPWDIDEALRR--RLEKRIYIPLPNFES 412
E++ Q+DG +R + VL ATN P +D AL R RL++++ LP+ ES
Sbjct: 188 L-EIVNQLDGFD--------ARGNIKVLMATNRPDTLDPALLRPGRLDRKVEFGLPDLES 340
Query: 413 RKELIRINLKTVEVAPDVNIDEVARRTDGYSGDDLTNVCRDASLNGMRRK 462
R ++ +I+ +T+ D+ + ++R +G D+ +VC +A + +R +
Sbjct: 341 RTQIFKIHTRTMNCERDIRFELLSRLCPNSTGADIRSVCTEAGMYAIRAR 490
>AV421390
Length = 409
Score = 80.9 bits (198), Expect = 5e-16
Identities = 54/125 (43%), Positives = 73/125 (58%), Gaps = 2/125 (1%)
Frame = +2
Query: 220 LERDVLETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPG 277
L ++V+ ++DV G +AK+ LEE VV L P F G + P KG+L+ G PG
Sbjct: 29 LNKEVMPEKNVKTFKDVKGCDDAKQELEE-VVEYLRNPAKFTRLGGKLP-KGILLTGAPG 202
Query: 278 TGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYAPSTIFIDEIDS 337
TGKTLLAKA+A E G FF + + + G R VR LF A+ AP IFIDEID+
Sbjct: 203 TGKTLLAKAIAGEAGVPFFYRAGSEFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDA 382
Query: 338 LCNSR 342
+ ++R
Sbjct: 383 VGSTR 397
>TC12379 homologue to UP|O04327 (O04327) Cell division protein FtsH isolog,
partial (15%)
Length = 505
Score = 79.3 bits (194), Expect = 1e-15
Identities = 53/150 (35%), Positives = 83/150 (55%), Gaps = 5/150 (3%)
Frame = +1
Query: 316 RCLFDLARAYAPSTIFIDEIDSLCNSRG---ASGEHESSRRVKSELLVQVDGVSNTATNE 372
R L+ A+ APS +FIDE+D++ RG SG E + ++LLV +DG
Sbjct: 61 RALYQEAKENAPSVVFIDELDAVGRERGLIKGSGGQERDATL-NQLLVCLDGFEG----- 222
Query: 373 DGSRKIVMVLAATNFPWDIDEALRR--RLEKRIYIPLPNFESRKELIRINLKTVEVAPDV 430
R V+ +A+TN P +D AL R R +++I+IP P R E+++++ + +A DV
Sbjct: 223 ---RGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGRIEILKVHARKKPIAGDV 393
Query: 431 NIDEVARRTDGYSGDDLTNVCRDASLNGMR 460
+ VA TDG G +L N+ A++N MR
Sbjct: 394 DYMAVASMTDGMVGAELANIVEVAAINMMR 483
>TC16655 homologue to UP|O99018 (O99018) Chloroplast protease precursor,
partial (39%)
Length = 934
Score = 77.0 bits (188), Expect = 7e-15
Identities = 49/104 (47%), Positives = 62/104 (59%), Gaps = 2/104 (1%)
Frame = +3
Query: 225 LETSPGVRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQ--GIRRPWKGVLMFGPPGTGKTL 282
+E + GV ++DVAG+ EAK+ E V L PE F G R P KGVL+ GPPGTGKTL
Sbjct: 627 MEPNTGVTFDDVAGVDEAKQDFMEVVEF-LKKPERFTAVGARIP-KGVLLIGPPGTGKTL 800
Query: 283 LAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLARAYA 326
LAKA+A E G FF++S + + G VR LF A+ A
Sbjct: 801 LAKAIAGEAGVPFFSISGSEFVEMFVGVGASRVRDLFKKAKENA 932
>CN825067
Length = 687
Score = 66.6 bits (161), Expect = 9e-12
Identities = 46/157 (29%), Positives = 82/157 (51%), Gaps = 1/157 (0%)
Frame = +1
Query: 266 PWKGVLMFGPPGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLFDLA-RA 324
P++ +L +GPPGTGKT+ A+ +A + G + ++ +A ++ + LFD + ++
Sbjct: 187 PFRNMLFYGPPGTGKTMAARELARKSGLDYALMTGGDVA-PLGSQAVTKIHELFDWSKKS 363
Query: 325 YAPSTIFIDEIDSLCNSRGASGEHESSRRVKSELLVQVDGVSNTATNEDGSRKIVMVLAA 384
+FIDE D+ R + E+ R + LL + D S+ IV+ L A
Sbjct: 364 KRGLLLFIDEADAFLCERNKTYMSEAQRSALNALLFRTG---------DQSKDIVLAL-A 513
Query: 385 TNFPWDIDEALRRRLEKRIYIPLPNFESRKELIRINL 421
TN P D+D A+ R+++ + PLP R +L+++ L
Sbjct: 514 TNRPGDLDSAVADRIDEVLEFPLPGEGERYKLLKLYL 624
>TC10924 homologue to UP|Q8H9D4 (Q8H9D4) 26S proteasome AAA-ATPase subunit
RPT4a, partial (49%)
Length = 648
Score = 65.9 bits (159), Expect = 2e-11
Identities = 39/85 (45%), Positives = 52/85 (60%), Gaps = 2/85 (2%)
Frame = +2
Query: 213 DPELAEMLERDVLETSPG-VRWEDVAGLTEAKRLLEEAVVLPLWMPEYFQGIR-RPWKGV 270
DP + ML D PG V + V GL++ R L E++ LPL PE F + +P KGV
Sbjct: 407 DPVVYNMLHED-----PGNVSYSAVGGLSDQIRELRESIELPLMNPELFLRVGIKPPKGV 571
Query: 271 LMFGPPGTGKTLLAKAVATECGTTF 295
L++GPPGTGKTLLA+A+A+ F
Sbjct: 572 LLYGPPGTGKTLLARAIASNIDANF 646
>AV422700
Length = 472
Score = 61.6 bits (148), Expect = 3e-10
Identities = 35/104 (33%), Positives = 55/104 (52%), Gaps = 4/104 (3%)
Frame = +2
Query: 220 LERDVLETSPGVRW---EDVAGLTEAKRLLEEAVVLPLWMPEYFQGIRRPW-KGVLMFGP 275
+E + ++ +W E + G TEA L E ++ P + + W +G+L++GP
Sbjct: 155 MESSMSSSNSNNQWRAEEAIGGNTEALEALRELIIFPQRFSHEARKLGLKWPRGLLLYGP 334
Query: 276 PGTGKTLLAKAVATECGTTFFNVSSATLASKWRGESERMVRCLF 319
PGTGKT L +AV ECG +S ++ GESER++R F
Sbjct: 335 PGTGKTSLVRAVVQECGAHLTIISPHSVHRAHAGESERILREAF 466
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.313 0.131 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,977,404
Number of Sequences: 28460
Number of extensions: 105371
Number of successful extensions: 689
Number of sequences better than 10.0: 96
Number of HSP's better than 10.0 without gapping: 625
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 665
length of query: 518
length of database: 4,897,600
effective HSP length: 94
effective length of query: 424
effective length of database: 2,222,360
effective search space: 942280640
effective search space used: 942280640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0226.8


