

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.7
(399 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
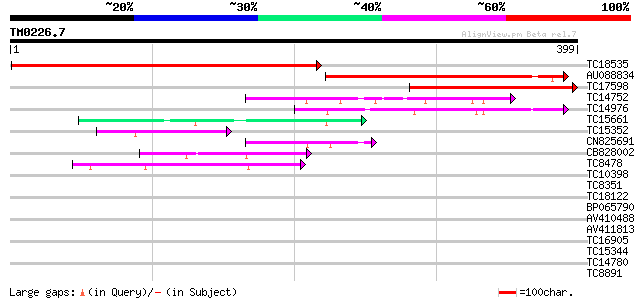
Score E
Sequences producing significant alignments: (bits) Value
TC18535 similar to GB|AAP68293.1|31711874|BT008854 At1g80360 {Ar... 434 e-122
AU088834 281 2e-76
TC17598 weakly similar to GB|AAP68293.1|31711874|BT008854 At1g80... 239 5e-64
TC14752 similar to UP|Q9LDV4 (Q9LDV4) Alanine aminotransferase ... 70 6e-13
TC14976 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26... 70 6e-13
TC15661 weakly similar to AAR99910 (AAR99910) Aminotransferase A... 54 5e-08
TC15352 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26... 49 2e-06
CN825691 48 3e-06
CB828002 40 9e-04
TC8478 similar to PIR|B86367|B86367 protein F26F24.16 [imported]... 40 9e-04
TC10398 weakly similar to UP|Q9FII5 (Q9FII5) Receptor protein ki... 32 0.14
TC8351 homologue to UP|Q41399 (Q41399) Chalcone reductase, complete 32 0.24
TC18122 similar to UP|Q8VZY4 (Q8VZY4) 1-aminocyclopropanecarboxy... 31 0.42
BP065790 30 0.71
AV410488 29 1.6
AV411813 28 2.7
TC16905 similar to UP|Q8GUE9 (Q8GUE9) Tyrosine aminotransferase ... 28 2.7
TC15344 similar to PIR|T05870|T05870 cell division protein pelot... 28 3.5
TC14780 27 4.6
TC8891 similar to GB|AAP68279.1|31711846|BT008840 At1g72680 {Ara... 27 6.0
>TC18535 similar to GB|AAP68293.1|31711874|BT008854 At1g80360 {Arabidopsis
thaliana;}, partial (47%)
Length = 751
Score = 434 bits (1116), Expect = e-122
Identities = 213/218 (97%), Positives = 213/218 (97%)
Frame = +2
Query: 2 GSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSV 61
G LARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSV
Sbjct: 98 GFVRELARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSV 277
Query: 62 SRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAP 121
SRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAP
Sbjct: 278 SRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAP 457
Query: 122 YYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTY 181
YYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESK VPKLVTVVNPGNPTGTY
Sbjct: 458 YYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKTVPKLVTVVNPGNPTGTY 637
Query: 182 IPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSC 219
IPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSC
Sbjct: 638 IPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSC 751
>AU088834
Length = 634
Score = 281 bits (718), Expect = 2e-76
Identities = 147/176 (83%), Positives = 152/176 (85%), Gaps = 5/176 (2%)
Frame = +3
Query: 223 NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLE 282
+HIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLE
Sbjct: 6 DHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLE 185
Query: 283 MGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLA 342
MGPEWVR RVQTLAKNREIVS LSPLG GSVKG EGAIYLFAKLPKGGYDDFDVVRWL
Sbjct: 186 MGPEWVRXRVQTLAKNREIVSXVLSPLGXGSVKGXEGAIYLFAKLPKGGYDDFDVVRWLT 365
Query: 343 NRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERL----KKGLEEL-VANGLV 393
NRHGIAVIPGSACGAAGNL ISFG D + A R KKGL ++ ANGL+
Sbjct: 366 NRHGIAVIPGSACGAAGNLXISFGSF---DXKLIAXRRXKG*KKGLGKIWFANGLI 524
>TC17598 weakly similar to GB|AAP68293.1|31711874|BT008854 At1g80360
{Arabidopsis thaliana;}, partial (27%)
Length = 562
Score = 239 bits (611), Expect = 5e-64
Identities = 118/118 (100%), Positives = 118/118 (100%)
Frame = +1
Query: 282 EMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWL 341
EMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWL
Sbjct: 1 EMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWL 180
Query: 342 ANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEELVANGLVQDPVNF 399
ANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEELVANGLVQDPVNF
Sbjct: 181 ANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEELVANGLVQDPVNF 354
>TC14752 similar to UP|Q9LDV4 (Q9LDV4) Alanine aminotransferase , partial
(49%)
Length = 1054
Score = 70.1 bits (170), Expect = 6e-13
Identities = 69/230 (30%), Positives = 102/230 (44%), Gaps = 40/230 (17%)
Frame = +1
Query: 167 KLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTY---------EYFMFDGLKH 217
+ + V+NPGNPTG + E + I CK G LL D Y ++ F +
Sbjct: 16 RALVVINPGNPTGQVLSEENQRAIVEFCKKEGLVLLADEVYQQNIYVPDKQFHSFKKVSR 195
Query: 218 SCVEGNHIVNIFSF---SKAY-GMMGWRVGYIAYPSEVEGLSA----QLLKVQDNIPICA 269
S G+ + + SF SK Y G G R GY+ EV G SA Q+ KV ++ +C+
Sbjct: 196 SMGYGDKDITLVSFQSISKGYHGECGKRGGYM----EVTGFSADVREQIYKVA-SVNLCS 360
Query: 270 SIISQHLALHSLEMGPEWVRER------------VQTLAKNREIVSEALSPLGEGSVKGG 317
+I Q LA SL M P V + + +LA+ +I+ + + L +
Sbjct: 361 NISGQILA--SLIMSPPKVGDESYESFMAEKENILSSLARRAKILEDGFNKLEGVTCNKA 534
Query: 318 EGAIYLF--AKLPKGGY---------DDFDVVRWLANRHGIAVIPGSACG 356
EGA+YLF +LP+ D + L N G+ V+PGS G
Sbjct: 535 EGAMYLFPQIRLPQKAIKAAEAANTAPDAFYCKRLLNATGVVVVPGSGFG 684
>TC14976 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26C19.9
{Arabidopsis thaliana;}, partial (42%)
Length = 758
Score = 70.1 bits (170), Expect = 6e-13
Identities = 59/211 (27%), Positives = 98/211 (45%), Gaps = 18/211 (8%)
Frame = +1
Query: 201 LLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLS 255
++ D YE ++ H+ + + + FSK++ M GWR+GYIA P
Sbjct: 1 VISDEIYENIIYAPATHTSFASLPGMWDRTLTVNGFSKSFAMTGWRLGYIAGPKH---FV 171
Query: 256 AQLLKVQDNIPICASIISQHLALHSLEM---GPEWVRERVQTLAKNREIVSEALSPLGEG 312
A K+Q AS ISQ A+ +L + G E V V+ + R+ + ++ +
Sbjct: 172 AACGKIQSQFTSGASSISQKAAVAALGLGYAGGEAVSTMVKAFRERRDFLVKSFGEIEGV 351
Query: 313 SVKGGEGAIYLFAKL------PKGGY----DDFDVVRWLANRHGIAVIPGSACGAAGNLR 362
+ +GA YLF G+ D + R+L ++ +A++PGSA G +R
Sbjct: 352 KISEPQGAFYLFIDFSFCYGREAEGFGKIEDSESLCRYLLDKGQVALVPGSAFGDDTCVR 531
Query: 363 ISFGGLTESDCRAAAERLKKGLEELVANGLV 393
IS+ + + +AA ER+KK L L + LV
Sbjct: 532 ISYAA-SLTTLQAAVERIKKALIPLSSAALV 621
>TC15661 weakly similar to AAR99910 (AAR99910) Aminotransferase ALD1,
partial (59%)
Length = 988
Score = 53.9 bits (128), Expect = 5e-08
Identities = 53/221 (23%), Positives = 87/221 (38%), Gaps = 18/221 (8%)
Frame = +2
Query: 49 LDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLT 108
+D V L E YG ++G LR + + ++ + S V V+ GA L L
Sbjct: 350 VDFVHGLSTEKGYKGYGPEQGEKALRKEIADTIYKDLGIKPSEVFVSDGAQCDISRLQLL 529
Query: 109 LCDAGDSVVMFAPY-YFNAYMS------------FQMTGITNIHVGPGSPDT-LYPDVDW 154
+ G ++ +F P F AY+ ++ NI P T +PD+
Sbjct: 530 M---GPNLKIFVPDPSFPAYIDSSVIIGQAGEFVYETGKYKNIEYMTCGPQTNFFPDLHT 700
Query: 155 LERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDG 214
R ++ +P NPTG L ++ + K GS ++ D+ Y ++ D
Sbjct: 701 ASRA--------DIIFFCSPNNPTGHAATRKQLDQLVDFAKENGSIIIFDSAYSAYITDD 856
Query: 215 LKHSCVE----GNHIVNIFSFSKAYGMMGWRVGYIAYPSEV 251
S E + + SFSK G G R+G+ P E+
Sbjct: 857 SPKSIFEISGAREVAIEVSSFSKFAGFTGVRLGWTVVPEEL 979
>TC15352 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26C19.9
{Arabidopsis thaliana;}, partial (33%)
Length = 1061
Score = 48.5 bits (114), Expect = 2e-06
Identities = 30/97 (30%), Positives = 48/97 (48%), Gaps = 2/97 (2%)
Frame = +2
Query: 62 SRYGADEGLPELRAALVKKLRQENNL--HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
+RY + G ELR A+ KL++EN + V+V+ GA Q+ VL +C GD V++
Sbjct: 770 TRYTPNAGTLELRQAICHKLKEENGITYTPDQVVVSNGAKQSIAQAVLAVCSPGDEVIIP 949
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLE 156
AP++ + +M T + + D D LE
Sbjct: 950 APFWVSYPEMARMADATPVILPTSISDNFLLDPKRLE 1060
>CN825691
Length = 590
Score = 48.1 bits (113), Expect = 3e-06
Identities = 36/105 (34%), Positives = 50/105 (47%), Gaps = 13/105 (12%)
Frame = +2
Query: 167 KLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE---------YFMFDGLKH 217
+++ V+NPGNPTG + E + I CK G LL D Y+ + F +
Sbjct: 221 RVLVVINPGNPTGQVLYEENQRDIVKFCKQEGLVLLADEVYQENVYVVEKKFQSFKKVSR 400
Query: 218 SCVEGNH---IVNIFSFSKA-YGMMGWRVGYIAYPSEVEGLSAQL 258
S G + IV++ S SK YG G R GY+ EV G SA +
Sbjct: 401 SMGYGENDISIVSLHSISKGYYGECGKRGGYM----EVTGFSADV 523
>CB828002
Length = 489
Score = 39.7 bits (91), Expect = 9e-04
Identities = 31/126 (24%), Positives = 57/126 (44%), Gaps = 5/126 (3%)
Frame = +1
Query: 92 VMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLY 149
+++TAGA A L+ L + G++ ++ PYY F+ + ++ TG+ + + S +
Sbjct: 1 ILLTAGATSANETLMFCLAEPGEAFLLPTPYYPGFDRDLKWR-TGVEIVPIQCNSSNNFQ 177
Query: 150 PDVDWLERILTESKPV---PKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNT 206
L + ++K K V V NP NP T + L + + K+ L+ D
Sbjct: 178 ITKSALHQAHEQAKKQNLRVKGVLVTNPSNPLATTMSRGKLDLLLDFIKDKDMHLISDEI 357
Query: 207 YEYFMF 212
Y +F
Sbjct: 358 YSGTVF 375
>TC8478 similar to PIR|B86367|B86367 protein F26F24.16 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(67%)
Length = 1124
Score = 39.7 bits (91), Expect = 9e-04
Identities = 36/172 (20%), Positives = 73/172 (41%), Gaps = 8/172 (4%)
Frame = +3
Query: 45 PKPALDKVKEL--VWEPSVSRYGADEGLPELRAALVKKLRQENNLHKSSVMV--TAGANQ 100
P A+ + K+ V + Y GLP +R + + +++ + ++ T GA++
Sbjct: 390 PADAIARAKQYFSVTSGGLGAYSDSRGLPGVRQEVAEFIQRRDGYPSDPELIYLTDGASK 569
Query: 101 AFVNLVLTLCDAG-DSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERIL 159
A + ++ T+ D +++ P Y + + G T + D L R +
Sbjct: 570 AVMQILNTIIRGPEDGILVPVPQYPLYSATIALLGGTLVGYYLEETANWGLDTKELRRSV 749
Query: 160 TESKPVP---KLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE 208
+++ K + ++NPGNPTG + + L+ + C LL D Y+
Sbjct: 750 EQARYKGINVKAMVILNPGNPTGQCLSQENLREVLQFCYEENLVLLGDEVYQ 905
>TC10398 weakly similar to UP|Q9FII5 (Q9FII5) Receptor protein kinase-like
protein, partial (5%)
Length = 602
Score = 32.3 bits (72), Expect = 0.14
Identities = 21/59 (35%), Positives = 26/59 (43%), Gaps = 8/59 (13%)
Frame = -3
Query: 237 MMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALH--------SLEMGPEW 287
++GW G I S +EGLSA L VQ N + IIS H + M P W
Sbjct: 291 LLGWASGSINTTSLMEGLSAGFLLVQSNAILSIGIISSLTEAHPAPAFLSNTSSMPPSW 115
>TC8351 homologue to UP|Q41399 (Q41399) Chalcone reductase, complete
Length = 1331
Score = 31.6 bits (70), Expect = 0.24
Identities = 15/45 (33%), Positives = 24/45 (53%)
Frame = +1
Query: 277 ALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
A++ +EM P W + +++ K + I A S LG V G GA+
Sbjct: 586 AVNQVEMNPSWQQGKLREFCKQKGIHVSAWSALGAYKVTWGSGAV 720
>TC18122 similar to UP|Q8VZY4 (Q8VZY4) 1-aminocyclopropanecarboxylic acid
synthase, partial (36%)
Length = 603
Score = 30.8 bits (68), Expect = 0.42
Identities = 18/68 (26%), Positives = 31/68 (45%), Gaps = 4/68 (5%)
Frame = +3
Query: 60 SVSRYGADEGLPELRAALVKKLRQENN----LHKSSVMVTAGANQAFVNLVLTLCDAGDS 115
+++ + GLPE R A+ K + + V+++ GA A L D GD+
Sbjct: 330 AIANFQDYHGLPEFREAVAKFMARTRGNRVTFDPDRVVMSGGATGAHEVTAFCLADPGDA 509
Query: 116 VVMFAPYY 123
++ PYY
Sbjct: 510 FLVPTPYY 533
>BP065790
Length = 410
Score = 30.0 bits (66), Expect = 0.71
Identities = 15/39 (38%), Positives = 22/39 (55%)
Frame = +1
Query: 143 GSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTY 181
GSP + P + W + +L S PV ++V PG P GT+
Sbjct: 226 GSPGSNMPSILW*KLVLHISYPVHRMVITGWPGFPHGTW 342
>AV410488
Length = 428
Score = 28.9 bits (63), Expect = 1.6
Identities = 21/91 (23%), Positives = 36/91 (39%), Gaps = 12/91 (13%)
Frame = +2
Query: 167 KLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCV------ 220
K + + NP NP GT + + L+ L+ D Y +F + +
Sbjct: 131 KGLLITNPSNPLGTVMDRTTLRTTLTFINENRIHLICDEIYAATVFGHPTFTSIAEIIEQ 310
Query: 221 ------EGNHIVNIFSFSKAYGMMGWRVGYI 245
+ + + ++S SK G G+RVG I
Sbjct: 311 DTDIECDRDLVHIVYSLSKDMGFPGFRVGII 403
>AV411813
Length = 419
Score = 28.1 bits (61), Expect = 2.7
Identities = 19/86 (22%), Positives = 37/86 (42%), Gaps = 9/86 (10%)
Frame = +1
Query: 169 VTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVD-----NTYEYFMF----DGLKHSC 219
+ + NP NP G + +L + + + ++ D +TY F + L
Sbjct: 76 ILISNPSNPVGNILTPDVLFSLLDFAEEKNIHIIADEVFAGSTYGSDKFVSIAEILDSEY 255
Query: 220 VEGNHIVNIFSFSKAYGMMGWRVGYI 245
V+ + + ++ SK + G+RVG I
Sbjct: 256 VDKSRVHIVYGLSKDLSLAGFRVGVI 333
>TC16905 similar to UP|Q8GUE9 (Q8GUE9) Tyrosine aminotransferase , partial
(37%)
Length = 610
Score = 28.1 bits (61), Expect = 2.7
Identities = 17/50 (34%), Positives = 25/50 (50%)
Frame = +3
Query: 333 DDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLKK 382
DD D LA + ++PG+A G LRI+F +D A AE + +
Sbjct: 312 DDIDFSFKLAKEESVIILPGTAVGLKDWLRITFA----ADPSALAEGMDR 449
>TC15344 similar to PIR|T05870|T05870 cell division protein pelota -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(23%)
Length = 498
Score = 27.7 bits (60), Expect = 3.5
Identities = 14/41 (34%), Positives = 25/41 (60%)
Frame = +3
Query: 98 ANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNI 138
A + FVNLV ++ D+G +V +F+ + + Q++GI I
Sbjct: 111 ARKKFVNLVNSIKDSGGTVHVFSSMHVSGEQLAQISGIAAI 233
>TC14780
Length = 671
Score = 27.3 bits (59), Expect = 4.6
Identities = 11/24 (45%), Positives = 17/24 (70%), Gaps = 4/24 (16%)
Frame = +2
Query: 113 GDSVVMFAPYYFNAYM----SFQM 132
GD+V++FAP +FN ++ FQM
Sbjct: 419 GDTVILFAPLFFNCFLCHLFDFQM 490
>TC8891 similar to GB|AAP68279.1|31711846|BT008840 At1g72680 {Arabidopsis
thaliana;} , complete
Length = 1636
Score = 26.9 bits (58), Expect = 6.0
Identities = 36/155 (23%), Positives = 64/155 (41%), Gaps = 8/155 (5%)
Frame = +1
Query: 86 NLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSP 145
+LH+ +M + GA++ + T D+ + +PY F S + G +++V
Sbjct: 76 HLHRK-IMSSEGASEECLGWAAT-----DTSGVLSPYKF----SRRAVGDDDVYVKISHC 225
Query: 146 DTLYPDVDWLERILTES-------KPVPKLVTVVNPGNPTGTYIPESL-LQRIANLCKNA 197
Y DV W L S + +VT V N G + + + + N C++
Sbjct: 226 GVCYADVAWTRNTLGNSVYPCVPGHEIAGIVTKVG-SNVHGFSVGDHVGVGTYVNSCRDC 402
Query: 198 GSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFS 232
F DGL+H CV+G ++F+F+
Sbjct: 403 E-----------FCDDGLEHHCVKG----SVFTFN 462
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,919,789
Number of Sequences: 28460
Number of extensions: 96438
Number of successful extensions: 414
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 408
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 411
length of query: 399
length of database: 4,897,600
effective HSP length: 92
effective length of query: 307
effective length of database: 2,279,280
effective search space: 699738960
effective search space used: 699738960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0226.7


