

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.6
(263 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
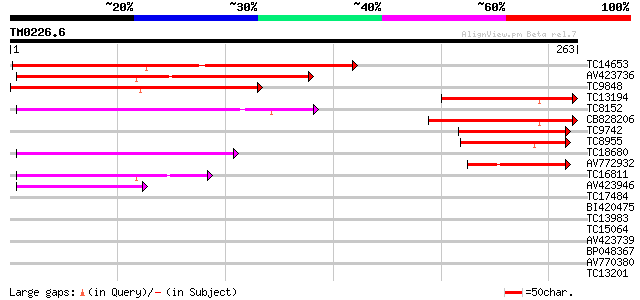
Sequences producing significant alignments: (bits) Value
TC14653 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lec... 181 1e-46
AV423736 129 7e-31
TC9848 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lect... 100 3e-22
TC13194 weakly similar to UP|Q94EW1 (Q94EW1) Nictaba, partial (29%) 79 6e-16
TC8152 73 6e-14
CB828206 72 7e-14
TC9742 similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin PP2-... 67 4e-12
TC8955 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lect... 64 2e-11
TC18680 63 5e-11
AV772932 53 6e-08
TC16811 49 9e-07
AV423946 42 1e-04
TC17484 weakly similar to UP|DBAT_TAXCU (Q9M6E2) 10-deacetylbacc... 29 0.72
BI420475 29 0.94
TC13983 homologue to UP|AAR29343 (AAR29343) Allantoinase , part... 28 1.2
TC15064 similar to UP|Q948P0 (Q948P0) Aspartic proteinase alpha,... 27 3.6
AV423739 27 3.6
BP048367 27 4.7
AV770380 27 4.7
TC13201 similar to UP|Q8LRL6 (Q8LRL6) Nam-like protein 9, partia... 26 6.1
>TC14653 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like
protein, partial (45%)
Length = 643
Score = 181 bits (458), Expect = 1e-46
Identities = 98/166 (59%), Positives = 115/166 (69%), Gaps = 6/166 (3%)
Frame = +2
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRAN 61
E LPEGCIA ILS T+P+D RL+LVS FRSAADSDAVWDRFLP D +SI+S S S A
Sbjct: 107 ENLPEGCIANILSLTSPLDVGRLALVSSAFRSAADSDAVWDRFLPDDFNSIISHSSSTAA 286
Query: 62 A------SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYR 115
A SKK YL LS +P++I +GKKSFQL +GKKC+ML ARAL I W + Y
Sbjct: 287 ADSSSSFKSKKDLYLSLSHNPLIIGDGKKSFQL--VNGKKCYMLSARALSIVWGDTPRYW 460
Query: 116 CWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFK 161
W +LPE+RF +V L + WLEI G IN LSP T YGA+LVFK
Sbjct: 461 RWISLPEARFSEVAELVSVCWLEIRGWINNKMLSPETMYGAYLVFK 598
>AV423736
Length = 454
Score = 129 bits (323), Expect = 7e-31
Identities = 71/140 (50%), Positives = 90/140 (63%), Gaps = 2/140 (1%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSL--SRAN 61
LPE C++ ILS T+P DA R SLVS RSAADSD VW FLPSD IVS+++ S
Sbjct: 38 LPEDCVSEILSHTSPPDACRFSLVSSTLRSAADSDMVWRSFLPSDYEDIVSRAVNPSALQ 217
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
SS K+ + L P+++D G KSF+LDK SGKK ++L AR L I W + + W + P
Sbjct: 218 FSSYKQLFHALCS-PLLLDGGNKSFKLDKLSGKKSYILSARDLSITWSSDPMFWSWRSNP 394
Query: 122 ESRFPDVVVLRHLLWLEIHG 141
ESRF +V LR + WLEI G
Sbjct: 395 ESRFREVAELRTVNWLEIEG 454
>TC9848 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (19%)
Length = 394
Score = 100 bits (248), Expect = 3e-22
Identities = 52/119 (43%), Positives = 74/119 (61%), Gaps = 2/119 (1%)
Frame = +1
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSR- 59
+E LPE C++ ILS T+P DA R S++S RSAA+SD +W FLPSD I+S++L+
Sbjct: 31 IETLPEECVSEILSHTSPPDACRFSMLSSTLRSAANSDMLWRSFLPSDYSDIISRALNPL 210
Query: 60 -ANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCW 117
N+SS + +P+++D G F+LDK SGKK ++L AR L I W + Y W
Sbjct: 211 FLNSSSSFKDLFKALCNPLLLDGGTMIFKLDKSSGKKSYILSARELSITWSSDPLYWTW 387
>TC13194 weakly similar to UP|Q94EW1 (Q94EW1) Nictaba, partial (29%)
Length = 471
Score = 79.3 bits (194), Expect = 6e-16
Identities = 41/64 (64%), Positives = 49/64 (76%), Gaps = 1/64 (1%)
Frame = +1
Query: 201 HDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQIDG-CFNVGLIIEGIEVRP 259
H R + +Q P+VRSDGWLEIEMGEF S LEDEE++MSV++I G GLI+EGIEVRP
Sbjct: 1 HFRVEGLQRPAVRSDGWLEIEMGEFSNSSLEDEELQMSVIEIQGYSSKSGLILEGIEVRP 180
Query: 260 KNDN 263
K N
Sbjct: 181 KEGN 192
>TC8152
Length = 715
Score = 72.8 bits (177), Expect = 6e-14
Identities = 45/142 (31%), Positives = 69/142 (47%), Gaps = 2/142 (1%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPE C+A IL P +L+ +++ FR+A+ +D VW+ LP++ +IV + + +
Sbjct: 199 LPESCVALILGYADPPQICQLATLNRAFRAASSADFVWESKLPANYRAIVRKIFTDFPSD 378
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTL--P 121
S KR +D G K LDK +GK C L + L I D R W +
Sbjct: 379 SSKRDTYAALCRLNTLDGGTKKAWLDKSTGKICMCLSFKGLSIT--GIDDRRYWKDIHTE 552
Query: 122 ESRFPDVVVLRHLLWLEIHGMI 143
ESRF V L+ W ++ G +
Sbjct: 553 ESRFGTVAYLQQTWWFQVDGEV 618
>CB828206
Length = 466
Score = 72.4 bits (176), Expect = 7e-14
Identities = 39/70 (55%), Positives = 51/70 (72%), Gaps = 1/70 (1%)
Frame = +1
Query: 195 SLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQIDG-CFNVGLIIE 253
+L H R +Q PSVRSDG LEIEMGEFF SGLEDEE+++S+L+++ + GL +E
Sbjct: 4 NLNERPHIRIQGLQRPSVRSDG*LEIEMGEFFNSGLEDEELQLSILEVNSRNWKSGLHLE 183
Query: 254 GIEVRPKNDN 263
GIEVRP+ N
Sbjct: 184 GIEVRPREVN 213
>TC9742 similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin PP2-like
protein, partial (20%)
Length = 794
Score = 66.6 bits (161), Expect = 4e-12
Identities = 31/52 (59%), Positives = 39/52 (74%)
Frame = +1
Query: 209 GPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQIDGCFNVGLIIEGIEVRPK 260
GP R DGW+EIE+GEFF G DEEV++SV+++ GLI+EGIEVRPK
Sbjct: 445 GPLKRDDGWMEIEVGEFFCDGEIDEEVKVSVMEVGYQLKGGLIVEGIEVRPK 600
Score = 36.6 bits (83), Expect = 0.005
Identities = 26/65 (40%), Positives = 32/65 (49%)
Frame = +2
Query: 86 FQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINT 145
F+LDK SGKK ++L AR L I W + Y WT+ S VV L L H T
Sbjct: 143 FKLDKSSGKKSYILSARELSITWSSDPLY--WTSS*MSH--TVVTALILRRLRYHRRWRT 310
Query: 146 FSLSP 150
S +P
Sbjct: 311 TSCTP 325
>TC8955 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like
protein, partial (12%)
Length = 693
Score = 64.3 bits (155), Expect = 2e-11
Identities = 29/52 (55%), Positives = 40/52 (76%), Gaps = 1/52 (1%)
Frame = +2
Query: 210 PSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQI-DGCFNVGLIIEGIEVRPK 260
P R+DGWLE+E+GEFF G ED EVEM+V ++ G + GL+++GIE+RPK
Sbjct: 41 PKERADGWLEMELGEFFCEGGEDREVEMAVCEVKGGDWKGGLVVQGIEIRPK 196
>TC18680
Length = 492
Score = 63.2 bits (152), Expect = 5e-11
Identities = 31/103 (30%), Positives = 53/103 (51%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE C+A + TP + L+ +++ FR AA +D+VW+ LPS+ ++ +
Sbjct: 181 IPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWETKLPSNYKDLLHALPPERFQN 360
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFI 106
K+ L P D+G K LD+ +G+ C + A+AL I
Sbjct: 361 LSKKDIFALLSRPQPFDDGNKEVWLDRVTGRVCMSISAKALSI 489
>AV772932
Length = 481
Score = 52.8 bits (125), Expect = 6e-08
Identities = 26/48 (54%), Positives = 34/48 (70%)
Frame = -3
Query: 213 RSDGWLEIEMGEFFISGLEDEEVEMSVLQIDGCFNVGLIIEGIEVRPK 260
R DGW+EIE+GEFF SG D E++M + ++ GL+ EGIEVRPK
Sbjct: 479 RDDGWMEIELGEFF-SGGGDVEIKMGLREMGYQLKGGLVFEGIEVRPK 339
>TC16811
Length = 509
Score = 48.9 bits (115), Expect = 9e-07
Identities = 28/93 (30%), Positives = 48/93 (51%), Gaps = 2/93 (2%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSL--SRAN 61
+PE CI+++L P + +L+ V+K F A+ +D +W+ LP +V++ L +
Sbjct: 233 VPENCISSLLMSLDPQEICKLARVNKAFHRASSADFLWESKLPPGYKFLVNKVLGEEKLG 412
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGK 94
+ +KK Y L P D G K LD+ S +
Sbjct: 413 SMTKKEIYAKLC-QPNFFDGGAKEIWLDRCSAQ 508
>AV423946
Length = 426
Score = 42.0 bits (97), Expect = 1e-04
Identities = 18/61 (29%), Positives = 33/61 (53%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE CI+++ P D +L+ V++ F A+ +D VW+ LP + ++ L N +
Sbjct: 241 IPESCISSLFMNLDPPDICKLARVNRAFHRASSADFVWESKLPPSYKFLANKILGEENLA 420
Query: 64 S 64
S
Sbjct: 421 S 423
>TC17484 weakly similar to UP|DBAT_TAXCU (Q9M6E2) 10-deacetylbaccatin III
10-O-acetyltransferase (DBAT) , partial (11%)
Length = 577
Score = 29.3 bits (64), Expect = 0.72
Identities = 13/36 (36%), Positives = 18/36 (49%)
Frame = +2
Query: 18 PVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIV 53
P +A L + K F + +W FLPSD HS +
Sbjct: 383 PTEACVLRKLPKPFEPSTSQRFLWKLFLPSDDHSFM 490
>BI420475
Length = 564
Score = 28.9 bits (63), Expect = 0.94
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 5/42 (11%)
Frame = -3
Query: 113 HYRCWT-----TLPESRFPDVVVLRHLLWLEIHGMINTFSLS 149
+Y CW T+P S VV+ RHLL + + G N+ S++
Sbjct: 196 YYTCWVLKFAYTVPSSVSIQVVIKRHLLSIVLDGCSNSTSVN 71
>TC13983 homologue to UP|AAR29343 (AAR29343) Allantoinase , partial (18%)
Length = 432
Score = 28.5 bits (62), Expect = 1.2
Identities = 11/26 (42%), Positives = 13/26 (49%)
Frame = +1
Query: 97 FMLGARALFIAWDNGDHYRCWTTLPE 122
F L R + WD G +CWT L E
Sbjct: 145 FSLNIRTSLLIWDEGYPEKCWTLLSE 222
>TC15064 similar to UP|Q948P0 (Q948P0) Aspartic proteinase alpha, partial
(77%)
Length = 1444
Score = 26.9 bits (58), Expect = 3.6
Identities = 11/19 (57%), Positives = 14/19 (72%)
Frame = +2
Query: 212 VRSDGWLEIEMGEFFISGL 230
V G+ +IEMG+FFI GL
Sbjct: 431 VTKKGYWQIEMGDFFIGGL 487
>AV423739
Length = 465
Score = 26.9 bits (58), Expect = 3.6
Identities = 13/56 (23%), Positives = 26/56 (46%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSR 59
LP+ + IL R P +++ V + R + SD +W+R + ++ + R
Sbjct: 106 LPDLALECILERLPPSALCQMAGVCRSLRESCVSDHLWERHMKQKWGKVIGPAAYR 273
>BP048367
Length = 205
Score = 26.6 bits (57), Expect = 4.7
Identities = 11/12 (91%), Positives = 12/12 (99%)
Frame = -1
Query: 249 GLIIEGIEVRPK 260
GLIIEG+EVRPK
Sbjct: 184 GLIIEGLEVRPK 149
>AV770380
Length = 526
Score = 26.6 bits (57), Expect = 4.7
Identities = 18/59 (30%), Positives = 26/59 (43%)
Frame = +3
Query: 83 KKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHG 141
K+S + + C A ALF+ + WT P +RF + L HLL +HG
Sbjct: 219 KESGSVKSSNVLGCNPFFA*ALFVGY--------WTMPPFARFKEFPTLAHLLLPTLHG 371
>TC13201 similar to UP|Q8LRL6 (Q8LRL6) Nam-like protein 9, partial (22%)
Length = 465
Score = 26.2 bits (56), Expect = 6.1
Identities = 13/32 (40%), Positives = 16/32 (49%)
Frame = -2
Query: 202 DRWDRVQGPSVRSDGWLEIEMGEFFISGLEDE 233
D D V D LE+E EFF+ G+E E
Sbjct: 218 DDADGVAVEESTGDAALEVEADEFFVGGVESE 123
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.321 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,741,463
Number of Sequences: 28460
Number of extensions: 64527
Number of successful extensions: 308
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 297
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 299
length of query: 263
length of database: 4,897,600
effective HSP length: 88
effective length of query: 175
effective length of database: 2,393,120
effective search space: 418796000
effective search space used: 418796000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0226.6


