

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.2
(583 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
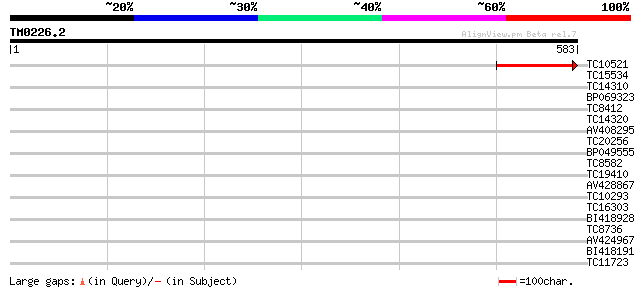
Score E
Sequences producing significant alignments: (bits) Value
TC10521 108 2e-24
TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28... 35 0.026
TC14310 homologue to UP|Q9M3H6 (Q9M3H6) Histone H2B, partial (92%) 33 0.17
BP069323 31 0.64
TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromos... 30 0.83
TC14320 similar to UP|Q41111 (Q41111) Dehydrin, partial (56%) 30 0.83
AV408295 30 1.4
TC20256 similar to UP|CYB_LAMGT (Q94T68) Cytochrome b, partial (6%) 29 2.4
BP049555 28 3.2
TC8582 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetyla... 28 3.2
TC19410 weakly similar to AAR24668 (AAR24668) At1g24040, partial... 28 4.1
AV428867 28 4.1
TC10293 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (19%) 28 4.1
TC16303 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin at... 28 4.1
BI418928 28 5.4
TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soyb... 28 5.4
AV424967 27 7.0
BI418191 27 9.2
TC11723 similar to UP|Q9SC91 (Q9SC91) Glutamine synthetase I (G... 27 9.2
>TC10521
Length = 751
Score = 108 bits (270), Expect = 2e-24
Identities = 57/84 (67%), Positives = 64/84 (75%), Gaps = 1/84 (1%)
Frame = +2
Query: 501 VHKLLGTVLLDQDAALRWKVRCAEYFPYSRYFEGSRSSASSNTALKQLSKNS-ENETLNH 559
VHKLLGTVL+D+DA LRWKVRCAE FPYS YFEG SSAS N+AL Q+ K S EN + +H
Sbjct: 5 VHKLLGTVLVDKDAXLRWKVRCAEIFPYSVYFEGRHSSASPNSALNQICKKSFENGSSSH 184
Query: 560 SVCTQNVGSITSNGKLEAFKQLAI 583
SV V SITSNGKL+ FK L I
Sbjct: 185 SVGNCKVESITSNGKLDTFKDLTI 256
>TC15534 weakly similar to GB|AAO63420.1|28950993|BT005356 At5g28040
{Arabidopsis thaliana;}, partial (8%)
Length = 937
Score = 35.4 bits (80), Expect = 0.026
Identities = 21/79 (26%), Positives = 35/79 (43%), Gaps = 1/79 (1%)
Frame = -3
Query: 286 LHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNE-ELSASGVSKSGK 344
L D P S++ E EE +Q K ++ + + + +EE+ E E +A +
Sbjct: 860 LEDPPTASSSSESEEEDDQPLSQVKHAAAEEEELSSEEEGSSEEEEEDETTAPAPPAATT 681
Query: 345 RHVKPVDPDPHGEKLLQVD 363
H KP P+P + Q D
Sbjct: 680 SHTKPPQPEPESDSATQSD 624
>TC14310 homologue to UP|Q9M3H6 (Q9M3H6) Histone H2B, partial (92%)
Length = 502
Score = 32.7 bits (73), Expect = 0.17
Identities = 18/55 (32%), Positives = 32/55 (57%), Gaps = 1/55 (1%)
Frame = +3
Query: 286 LHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEAR-AKKGAEEKNEELSASGV 339
L PP + A+ +++ ++ PA +K +K+ KAE + AK+G EK ++ S V
Sbjct: 39 LSSPPPMAPAKAEKKPAEKKPAAEKAPAEKKPKAEKKIAKEGGSEKKKKKSKKSV 203
>BP069323
Length = 404
Score = 30.8 bits (68), Expect = 0.64
Identities = 12/27 (44%), Positives = 14/27 (51%)
Frame = -2
Query: 418 PDSHRCLVCQLHEKTLFEANNSFLEKH 444
P +H CL C + EKTLF L H
Sbjct: 265 PSTHLCLACSVSEKTLFTCKTQTLPHH 185
>TC8412 weakly similar to UP|Q9LW00 (Q9LW00) Genomic DNA, chromosome 3, P1
clone: MSJ11, partial (39%)
Length = 1403
Score = 30.4 bits (67), Expect = 0.83
Identities = 12/36 (33%), Positives = 23/36 (63%)
Frame = +1
Query: 289 SPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAK 324
+P ++ A++ + + K+LP +K+K RQK K + K
Sbjct: 376 TPQRNVAKDHQLLKKMLPGKKRKRRQKMWKCKKLVK 483
>TC14320 similar to UP|Q41111 (Q41111) Dehydrin, partial (56%)
Length = 1231
Score = 30.4 bits (67), Expect = 0.83
Identities = 26/88 (29%), Positives = 37/88 (41%), Gaps = 18/88 (20%)
Frame = +3
Query: 285 KLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRK-----------------AEARAKKGA 327
KLH S S++ DEE K +KKK ++K++K A A KKG
Sbjct: 579 KLHRSDSSSSSSSDEEEGK-EGEEKKKKKKKEKKEVKSTDENTSVPVEVDPAHAEEKKGF 755
Query: 328 EEK-NEELSASGVSKSGKRHVKPVDPDP 354
+K E+L G + + P P P
Sbjct: 756 LDKIKEKLPGHGKKTTEEATTSPPPPPP 839
>AV408295
Length = 426
Score = 29.6 bits (65), Expect = 1.4
Identities = 22/73 (30%), Positives = 39/73 (53%), Gaps = 7/73 (9%)
Frame = +3
Query: 285 KLHDSPPKSTAEE----DEEMSKLLPAQKKKMRQKQRKAEAR--AKKGAEEKN-EELSAS 337
K+ + K T EE +EE + + ++KK ++++KAEA A EEKN E
Sbjct: 186 KVEEEEEKKTEEEKKPAEEEKKEEVVEEEKKPAEEEKKAEAEGEATTSGEEKNVVEAVTE 365
Query: 338 GVSKSGKRHVKPV 350
V ++G++ V+ +
Sbjct: 366 VVVEAGEKTVEAI 404
>TC20256 similar to UP|CYB_LAMGT (Q94T68) Cytochrome b, partial (6%)
Length = 543
Score = 28.9 bits (63), Expect = 2.4
Identities = 15/44 (34%), Positives = 27/44 (61%)
Frame = -1
Query: 293 STAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSA 336
S ++ED++ K K ++K++KAE+ AKK ++ E+L A
Sbjct: 543 SDSKEDDKTDK-----NNKKKKKKKKAESNAKKPKSDRFEKLEA 427
>BP049555
Length = 555
Score = 28.5 bits (62), Expect = 3.2
Identities = 20/88 (22%), Positives = 40/88 (44%)
Frame = +2
Query: 302 SKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQ 361
S L ++ +Q+Q +AEA+ ++ EEK EE + + K+ + P EK +
Sbjct: 206 STFLEQMGEEAKQEQPQAEAKPEEKTEEKAEEKAEEKPAAEEKKE----EEKPEEEKKEE 373
Query: 362 VDDPLSEAIKYLKLLQKNSPDSLETHLL 389
P + + Y+ L +E +++
Sbjct: 374 EPKPPAPCVLYVDLHCVGCAKKIERYMM 457
Score = 26.9 bits (58), Expect = 9.2
Identities = 15/59 (25%), Positives = 32/59 (53%), Gaps = 3/59 (5%)
Frame = +3
Query: 292 KSTAEEDEEMSKLLPAQKKKMRQKQRKAEARA---KKGAEEKNEELSASGVSKSGKRHV 347
K +++ +L P+Q+K+ R+KQRK + ++ +K K+++ S + + HV
Sbjct: 222 KWVRKQNRNNLRLKPSQRKRQRKKQRKKQRKSLQLRKRRRRKSQKRRRRKKSLNHQLHV 398
>TC8582 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (65%)
Length = 1341
Score = 28.5 bits (62), Expect = 3.2
Identities = 19/67 (28%), Positives = 30/67 (44%)
Frame = +1
Query: 288 DSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHV 347
DS +S E D+E PA KK+ Q +++ A K + +A+ GK++V
Sbjct: 616 DSDDESDDESDDEEETPTPA--KKVDQGKKRPNESASKTPVSGKKAKNATPEKPDGKKNV 789
Query: 348 KPVDPDP 354
P P
Sbjct: 790 HTATPHP 810
>TC19410 weakly similar to AAR24668 (AAR24668) At1g24040, partial (28%)
Length = 606
Score = 28.1 bits (61), Expect = 4.1
Identities = 17/41 (41%), Positives = 23/41 (55%), Gaps = 2/41 (4%)
Frame = -2
Query: 343 GKRHVKPVDPDPHGE--KLLQVDDPLSEAIKYLKLLQKNSP 381
G V+P DPDPHG +L+ V++ E + LKLL P
Sbjct: 416 GVHLVRPHDPDPHGSGFQLVGVEEGF-EGFELLKLLVGEKP 297
>AV428867
Length = 628
Score = 28.1 bits (61), Expect = 4.1
Identities = 12/30 (40%), Positives = 17/30 (56%)
Frame = +1
Query: 266 LHSHSYFHKAAAGAIRCYIKLHDSPPKSTA 295
L +FHK G +C++K+ PPK TA
Sbjct: 511 LRKSFFFHKGWLGVFKCFLKIW--PPKQTA 594
>TC10293 similar to UP|Q84TG3 (Q84TG3) At2g35930, partial (19%)
Length = 757
Score = 28.1 bits (61), Expect = 4.1
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = -2
Query: 411 LRLDAEHPDSHRCLVCQLH 429
LRL +HP RC+VC H
Sbjct: 543 LRLKTQHPQQLRCIVCLFH 487
>TC16303 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment
subunit precursor, partial (3%)
Length = 1235
Score = 28.1 bits (61), Expect = 4.1
Identities = 23/85 (27%), Positives = 39/85 (45%), Gaps = 9/85 (10%)
Frame = +1
Query: 288 DSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAE---------ARAKKGAEEKNEELSASG 338
++P EED+E++ L KKK + ++ AE A + AEE + EL+ G
Sbjct: 727 EAPQAEEGEEDDEINDLFKMGKKKKKNERSPAEIALLVENVMAELEVTAEE-DAELNRQG 903
Query: 339 VSKSGKRHVKPVDPDPHGEKLLQVD 363
K P+ + +K LQ++
Sbjct: 904 KPAINKLKKLPLLTEVLSKKQLQLE 978
>BI418928
Length = 496
Score = 27.7 bits (60), Expect = 5.4
Identities = 23/104 (22%), Positives = 46/104 (44%)
Frame = +3
Query: 221 RALKKFLGVEKHYADINEDQFDFHSYCLRKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAI 280
RAL+ F GVE AD++ + K+T+ ++ K +D+L + + +
Sbjct: 105 RALRNFNGVEDVKADLSGN----------KLTVIGKVDPTKVRDKLAEKTNKNVELVSSP 254
Query: 281 RCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAK 324
D PP E++ + +++K + ++KAE + K
Sbjct: 255 PKKDAAGDKPPPEKKPEEKPKTDDKKPEEEKTKPDEKKAEEKTK 386
>TC8736 similar to PIR|T06396|T06396 isoprenylated protein - soybean
(fragment) {Glycine max;}, partial (84%)
Length = 1268
Score = 27.7 bits (60), Expect = 5.4
Identities = 21/68 (30%), Positives = 29/68 (41%), Gaps = 4/68 (5%)
Frame = +2
Query: 291 PKSTAE----EDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRH 346
PK+ AE +D E K +KKK K++ KKG E+ ++ G
Sbjct: 551 PKAAAEPEKKKDSEKPKAAEPEKKKDGDKEKPKSDGPKKGDEKPKDK------PAPGPIP 712
Query: 347 VKPVDPDP 354
V PV P P
Sbjct: 713 VHPVGPPP 736
>AV424967
Length = 344
Score = 27.3 bits (59), Expect = 7.0
Identities = 10/35 (28%), Positives = 18/35 (50%)
Frame = +1
Query: 493 WKLKDCIAVHKLLGTVLLDQDAALRWKVRCAEYFP 527
WKLK+ A +LG+ + W ++ ++FP
Sbjct: 1 WKLKEIFATGVVLGSYMALMTVVFFWLIKDTDFFP 105
>BI418191
Length = 652
Score = 26.9 bits (58), Expect = 9.2
Identities = 12/28 (42%), Positives = 16/28 (56%)
Frame = -1
Query: 520 VRCAEYFPYSRYFEGSRSSASSNTALKQ 547
+RC F YSR S S+ SSN + K+
Sbjct: 403 IRCRMVFKYSRSLGSSESNKSSNLSTKE 320
>TC11723 similar to UP|Q9SC91 (Q9SC91) Glutamine synthetase I
(Glutamate--ammonia ligase) , partial (17%)
Length = 528
Score = 26.9 bits (58), Expect = 9.2
Identities = 18/62 (29%), Positives = 32/62 (51%)
Frame = +3
Query: 348 KPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAV 407
+PVD DP+ E L ++ LSE+++ L N D L+ + K+L F+A+
Sbjct: 21 EPVDADPNPENLQRLPKSLSESLEAL-----NKDDFLKEFI--------DDKLLTAFKAI 161
Query: 408 KQ 409
++
Sbjct: 162 RK 167
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,400,021
Number of Sequences: 28460
Number of extensions: 124109
Number of successful extensions: 676
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 668
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 675
length of query: 583
length of database: 4,897,600
effective HSP length: 95
effective length of query: 488
effective length of database: 2,193,900
effective search space: 1070623200
effective search space used: 1070623200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0226.2


