

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
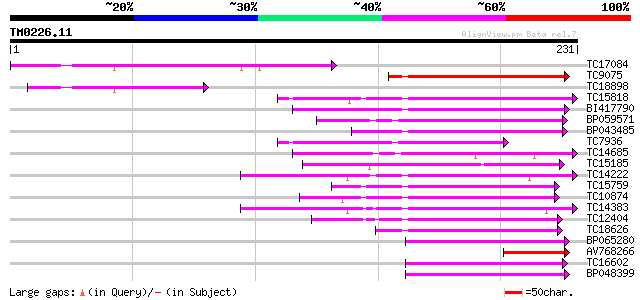
Score E
Sequences producing significant alignments: (bits) Value
TC17084 157 2e-39
TC9075 similar to UP|Q93VH8 (Q93VH8) 2-oxoglutarate-dependent di... 124 1e-29
TC18898 weakly similar to UP|Q93WB6 (Q93WB6) 2-oxoglutarate-depe... 88 1e-18
TC15818 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyge... 71 1e-13
BI417790 68 1e-12
BP059571 67 3e-12
BP043485 67 3e-12
TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, pa... 62 6e-11
TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partia... 60 3e-10
TC15185 60 4e-10
TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carb... 59 9e-10
TC15759 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxyge... 58 1e-09
TC10874 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxyge... 58 1e-09
TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-ca... 56 6e-09
TC12404 similar to UP|FLS_CITUN (Q9ZWQ9) Flavonol synthase (FLS... 54 2e-08
TC18626 54 2e-08
BP065280 54 3e-08
AV768266 53 5e-08
TC16602 weakly similar to UP|Q7X9N5 (Q7X9N5) Gibberellin 2-oxida... 53 5e-08
BP048399 52 9e-08
>TC17084
Length = 597
Score = 157 bits (396), Expect = 2e-39
Identities = 93/197 (47%), Positives = 110/197 (55%), Gaps = 64/197 (32%)
Frame = +1
Query: 1 MGSETTLKLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY------------------- 41
MGSETT KLPVIDFT+LN+L NWEAVKSKVH+ALVEY
Sbjct: 19 MGSETTRKLPVIDFTDLNSLA----NWEAVKSKVHEALVEYGCFEAIFDKVPLDLRKEIF 186
Query: 42 -------------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESLTKILWPN 88
LNVSKKPY GY GQ+P +PLF+SMGIDDANV EKVE++ ILWP+
Sbjct: 187 ASLEELFNLPLQTKLLNVSKKPYHGYVGQYPMVPLFESMGIDDANVSEKVEAMANILWPD 366
Query: 89 GNPSF-----------------------------RYASQPI---TYFLKTMKYKGPQTSD 116
GNPSF +Y + + Y L+ MKYKGP+TSD
Sbjct: 367 GNPSFSKTINSFSEQLSELDHIIRKMVLESLGVEKYLDEHLNSTNYLLRVMKYKGPETSD 546
Query: 117 TKVGLDTHTDTAILTIL 133
TK+GL TH+D I+TIL
Sbjct: 547 TKLGLTTHSDKNIVTIL 597
>TC9075 similar to UP|Q93VH8 (Q93VH8) 2-oxoglutarate-dependent dioxygenase
(Fragment), partial (26%)
Length = 533
Score = 124 bits (311), Expect = 1e-29
Identities = 59/74 (79%), Positives = 63/74 (84%)
Frame = +3
Query: 155 KPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEE 214
KPSSES F+VM+ D HAWSNGRLHSPFHRVMMTGNEARYS LF +PKG IIK PEE
Sbjct: 3 KPSSES--FVVMIGDSLHAWSNGRLHSPFHRVMMTGNEARYSTGLFSIPKGEYIIKTPEE 176
Query: 215 LVDEEHPLLYKPFD 228
LVDEEHPLL+KPFD
Sbjct: 177 LVDEEHPLLFKPFD 218
>TC18898 weakly similar to UP|Q93WB6 (Q93WB6) 2-oxoglutarate-dependent
dioxygenase (Fragment), partial (9%)
Length = 332
Score = 88.2 bits (217), Expect = 1e-18
Identities = 52/108 (48%), Positives = 60/108 (55%), Gaps = 34/108 (31%)
Frame = +3
Query: 8 KLPVIDFTNLNNLGGTSPNWEAVKSKVHKALVEY-------------------------- 41
KLP IDFTNLN+L NWEAVKSKVHKALV+Y
Sbjct: 21 KLPAIDFTNLNDLA----NWEAVKSKVHKALVDYGCFEAIFDTLVVPLDLRKEVFTQLED 188
Query: 42 --------MQLNVSKKPYFGYAGQFPNIPLFKSMGIDDANVYEKVESL 81
QLNVS K Y GY GQ+P IPL++SMGIDDA+V E VE++
Sbjct: 189 LFDLPLQTKQLNVSNKTYHGYIGQYPMIPLYESMGIDDADVSENVETM 332
>TC15818 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (37%)
Length = 618
Score = 71.2 bits (173), Expect = 1e-13
Identities = 47/123 (38%), Positives = 65/123 (52%), Gaps = 1/123 (0%)
Frame = +2
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMN 168
K PQ D +GL H+D LTIL V GL+V + G WI+ KP + FI+ +
Sbjct: 8 KCPQP-DLTLGLSPHSDPGGLTILLPDDFVSGLQV--RKGNYWITVKPVPNA--FIINIG 172
Query: 169 DCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
D SN S HRV++ N+ R S A+F+ PK +I+ +ELV EE P LY P
Sbjct: 173 DQIQVLSNTIYKSVEHRVIVNSNQDRVSLAMFYNPKSDLLIEPAKELVTEEKPALYPPMT 352
Query: 229 RED 231
++
Sbjct: 353 YDE 361
>BI417790
Length = 610
Score = 68.2 bits (165), Expect = 1e-12
Identities = 37/113 (32%), Positives = 56/113 (48%)
Frame = +3
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D +G H D+ LTIL Q +V GL+V K +W+ P ++ +I+ + D W
Sbjct: 42 DIVLGCGRHKDSGALTILAQDEVSGLQVRRKCDGQWVRLNPLPDA--YIINVGDVIQVWC 215
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
N S HRV + +AR S F P ++ EEL +EE+P Y P++
Sbjct: 216 NDAYESVEHRVKLNPEKARLSYPFFLYPSHYTNVEPLEELTNEENPPKYTPYN 374
>BP059571
Length = 386
Score = 67.0 bits (162), Expect = 3e-12
Identities = 39/102 (38%), Positives = 54/102 (52%)
Frame = +2
Query: 126 DTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHR 185
D LTIL Q QV GL+V + +W S P + +F+V + D F A SNGR S HR
Sbjct: 77 DPTSLTILHQDQVGGLQVFVDN--EWHSISP--DLNAFVVNIGDTFMALSNGRYKSCLHR 244
Query: 186 VMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
++ + R S A F P+ ++ P ELVD +P +Y F
Sbjct: 245 AVVNSEKTRKSLAFFLCPRNDKVVTPPCELVDSCNPRIYPDF 370
>BP043485
Length = 522
Score = 66.6 bits (161), Expect = 3e-12
Identities = 31/88 (35%), Positives = 49/88 (55%)
Frame = -2
Query: 140 GLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAAL 199
GLEV K ++W+ +PS ++ +I+ + D WSN S HR M+ + RYS
Sbjct: 521 GLEVKNKANQQWVRVEPSPDA--YIINVGDIIQVWSNDAYESVEHRAMLNSEKQRYSIPF 348
Query: 200 FFVPKGGCIIKAPEELVDEEHPLLYKPF 227
FF P +K +ELV+E++P Y+P+
Sbjct: 347 FFFPSHDTEVKPLDELVNEQNPSKYRPY 264
>TC7936 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(94%)
Length = 1492
Score = 62.4 bits (150), Expect = 6e-11
Identities = 35/94 (37%), Positives = 51/94 (54%)
Frame = +3
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V + D
Sbjct: 759 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDNGKTWITVQP--VEGAFVVNLGD 929
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVP 203
H SNGR + H+ ++ N +R S A F P
Sbjct: 930 HGHYLSNGRFKNADHQAVVNSNSSRLSIATFQNP 1031
>TC14685 weakly similar to UP|Q43383 (Q43383) 2A6 protein, partial (38%)
Length = 1323
Score = 60.1 bits (144), Expect = 3e-10
Identities = 39/118 (33%), Positives = 58/118 (49%), Gaps = 2/118 (1%)
Frame = +1
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
D VGL++H D LT++ Q + GL+V T +G W+ KP + ++ + D S
Sbjct: 793 DLTVGLNSHADPGALTVVLQDHIGGLQVRTTEG--WVDVKPL--DGALVINIGDLLQIIS 960
Query: 176 NGRLHSPFHRVMM-TGNEARYSAALFFVPKGGCIIKAP-EELVDEEHPLLYKPFDRED 231
N S HRV+ + NE R S A+F P + P EL E P LY+ F ++
Sbjct: 961 NEEYKSADHRVLANSANEPRVSVAVFLNPGNREKLFGPLPELTSAEKPALYRNFTLDE 1134
>TC15185
Length = 876
Score = 59.7 bits (143), Expect = 4e-10
Identities = 34/108 (31%), Positives = 51/108 (46%), Gaps = 1/108 (0%)
Frame = +3
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLT-KDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
G HTD ++T+L V GL++ +D + I + +FIV + D WSNG
Sbjct: 18 GAGAHTDFGLITLLATDDVSGLQICKDRDAKPQIWEDVAPLKGAFIVNLGDMLERWSNGV 197
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
S HRV+ G E RYS A F P C+++ + +P + P
Sbjct: 198 FKSTLHRVLGNGQE-RYSIAYFLEPSHDCLVECLPTCKSDSNPPQFPP 338
>TC14222 similar to UP|Q41681 (Q41681) 1-aminocyclopropane-1-carboxylate
oxidase homolog (ACC oxidase) , partial (97%)
Length = 1597
Score = 58.5 bits (140), Expect = 9e-10
Identities = 44/139 (31%), Positives = 64/139 (45%), Gaps = 2/139 (1%)
Frame = +1
Query: 95 YASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWIS 153
Y S+ + K Y D GL HTD + +LFQ +V GL++L D +WI
Sbjct: 487 YGSKGPNFGTKVSNYPPCPKPDLIKGLRAHTDAGGIILLFQDDKVSGLQLLKDD--QWID 660
Query: 154 YKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIK-AP 212
P S +V + D +NG+ S HRV+ + AR S A F+ P +I AP
Sbjct: 661 VPPMRHS--IVVNLGDQLEVITNGKYKSVMHRVIAQTDGARMSLASFYNPGDDAVISPAP 834
Query: 213 EELVDEEHPLLYKPFDRED 231
+ ++E +Y F ED
Sbjct: 835 SLVKEDETSQVYPKFVFED 891
>TC15759 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxygenase-like
protein, partial (34%)
Length = 929
Score = 58.2 bits (139), Expect = 1e-09
Identities = 36/93 (38%), Positives = 50/93 (53%)
Frame = +2
Query: 132 ILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGN 191
+L QV GL+V + G WI+ KP + FIV + D SN S HRV++ N
Sbjct: 2 LLPDDQVTGLQV--RKGDNWITVKPVPHA--FIVNIGDQIQVLSNAIYKSVEHRVIVNSN 169
Query: 192 EARYSAALFFVPKGGCIIKAPEELVDEEHPLLY 224
+ R S A F+ P+G I+ +ELV E+ P LY
Sbjct: 170 KERVSLAFFYNPRGDIPIEPAKELVKEDKPALY 268
>TC10874 similar to UP|Q8LF06 (Q8LF06) Leucoanthocyanidin dioxygenase-like
protein, partial (38%)
Length = 723
Score = 58.2 bits (139), Expect = 1e-09
Identities = 38/107 (35%), Positives = 57/107 (52%), Gaps = 1/107 (0%)
Frame = +2
Query: 119 VGLDTHTDTAILTILF-QSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNG 177
+GL +H+D +T+L QV GL+V + G WI+ P+ + FIV + D SN
Sbjct: 8 LGLSSHSDPGGMTMLLPDDQVRGLQV--RKGDDWITVNPARHA--FIVNIGDQIQVLSNA 175
Query: 178 RLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLY 224
S HRV++ ++ R S A F+ PK I+ +ELV + P LY
Sbjct: 176 IYTSVEHRVIVNSDKERVSLAFFYNPKSDIPIEPVKELVTPDKPALY 316
>TC14383 homologue to UP|Q8W3Y3 (Q8W3Y3) 1-aminocyclopropane-1-carboxylic
acid oxidase, complete
Length = 1249
Score = 55.8 bits (133), Expect = 6e-09
Identities = 45/141 (31%), Positives = 66/141 (45%), Gaps = 4/141 (2%)
Frame = +3
Query: 95 YASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWIS 153
+ S+ T+ K Y + GL HTD + +LFQ +V GL++L KDG +W+
Sbjct: 480 HGSRGPTFGTKVANYPPCPKPNLVKGLRAHTDAGGIILLFQDDKVSGLQLL-KDG-EWVD 653
Query: 154 YKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPE 213
P S +V + D +NG+ S HRV+ + R S A F+ P +I
Sbjct: 654 VPPMRHS--IVVNLGDQLEVITNGKYKSVEHRVIAQTDGTRMSIASFYNPGSDAVIYPAP 827
Query: 214 ELVD---EEHPLLYKPFDRED 231
EL++ EE LY F ED
Sbjct: 828 ELLEKEAEEKNQLYPKFVFED 890
>TC12404 similar to UP|FLS_CITUN (Q9ZWQ9) Flavonol synthase (FLS) (CitFLS)
, partial (31%)
Length = 526
Score = 54.3 bits (129), Expect = 2e-08
Identities = 38/102 (37%), Positives = 51/102 (49%)
Frame = +2
Query: 124 HTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPF 183
HTD + LTIL ++V GL+VL KD R WI K + IV + D SNGR S
Sbjct: 5 HTDLSSLTILLPNEVPGLQVL-KDER-WIDAKYIPHA--LIVHIGDQIQIASNGRYKSVL 172
Query: 184 HRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYK 225
HR + R S +F P C + +LV +++P YK
Sbjct: 173 HRTTVDKERTRMSWPVFLEPPAECEVGPLPQLVTQDNPPKYK 298
>TC18626
Length = 469
Score = 53.9 bits (128), Expect = 2e-08
Identities = 28/76 (36%), Positives = 39/76 (50%)
Frame = +2
Query: 150 KWISYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCII 209
+WI KP S FI+ + D WSN S HRVM+ + R+S F P +
Sbjct: 11 EWIRVKPIFNS--FIINVGDMIQVWSNDAYESVEHRVMVNSEKDRFSIPFFLKPALYTDV 184
Query: 210 KAPEELVDEEHPLLYK 225
K EEL D++HP +Y+
Sbjct: 185 KPLEELTDDKHPPIYR 232
>BP065280
Length = 470
Score = 53.5 bits (127), Expect = 3e-08
Identities = 24/67 (35%), Positives = 38/67 (55%)
Frame = -1
Query: 162 SFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHP 221
+FI+ + D WSNG+ S HRV++ + R S FF+P ++K EELV+E P
Sbjct: 458 AFIINVGDITEVWSNGKYESVEHRVVVNAEKERLSYPFFFLPAHHVMVKPAEELVNEHXP 279
Query: 222 LLYKPFD 228
Y+ ++
Sbjct: 278 ARYREYN 258
>AV768266
Length = 255
Score = 52.8 bits (125), Expect = 5e-08
Identities = 21/27 (77%), Positives = 26/27 (95%)
Frame = -1
Query: 202 VPKGGCIIKAPEELVDEEHPLLYKPFD 228
+PKGG ++KAP+ELVDEEHPLL+KPFD
Sbjct: 255 IPKGGYMVKAPKELVDEEHPLLFKPFD 175
>TC16602 weakly similar to UP|Q7X9N5 (Q7X9N5) Gibberellin 2-oxidase, partial
(11%)
Length = 476
Score = 52.8 bits (125), Expect = 5e-08
Identities = 23/66 (34%), Positives = 37/66 (55%)
Frame = +1
Query: 162 SFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHP 221
++I+ + D WSN S HRVM+ + RYS FF P +K E+L++E++P
Sbjct: 7 AYIINVGDITQVWSNDAYESVEHRVMVNSEKERYSIPYFFFPAHDTEVKPLEKLINEQNP 186
Query: 222 LLYKPF 227
Y+P+
Sbjct: 187 AKYRPY 204
>BP048399
Length = 481
Score = 52.0 bits (123), Expect = 9e-08
Identities = 23/67 (34%), Positives = 37/67 (54%)
Frame = -1
Query: 162 SFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHP 221
+FI+ + D WS+ S HRVM+ + R+S FF P C +K +EL +E +P
Sbjct: 418 AFIINVCDSIQVWSHDAYESVEHRVMVNSEKERFSIPYFFNPAHNCEVKPLDELTNEPNP 239
Query: 222 LLYKPFD 228
Y+P++
Sbjct: 238 SKYRPYE 218
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,025,136
Number of Sequences: 28460
Number of extensions: 53944
Number of successful extensions: 333
Number of sequences better than 10.0: 86
Number of HSP's better than 10.0 without gapping: 312
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 314
length of query: 231
length of database: 4,897,600
effective HSP length: 87
effective length of query: 144
effective length of database: 2,421,580
effective search space: 348707520
effective search space used: 348707520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0226.11


