

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.8
(423 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
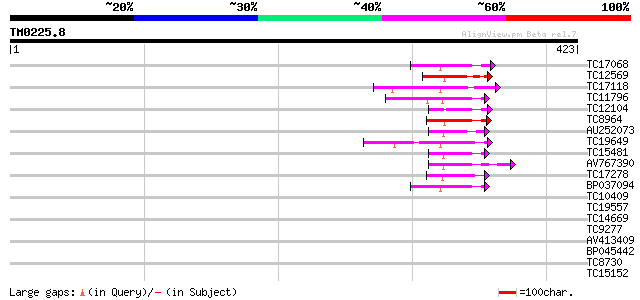
Score E
Sequences producing significant alignments: (bits) Value
TC17068 similar to UP|Q8L775 (Q8L775) RING finger-like protein, ... 44 4e-05
TC12569 UP|Q40221 (Q40221) Protein containing C-terminal RING-fi... 44 5e-05
TC17118 similar to UP|Q94CL1 (Q94CL1) RING finger-like protein (... 44 7e-05
TC11796 similar to GB|AAP12870.1|30017273|BT006221 At4g35840 {Ar... 42 2e-04
TC12104 41 4e-04
TC8964 weakly similar to UP|Q94F54 (Q94F54) AT3g15070/K15M2_22, ... 41 4e-04
AU252073 41 4e-04
TC19649 weakly similar to UP|RNF6_HUMAN (Q9Y252) RING finger pro... 40 6e-04
TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-l... 40 6e-04
AV767390 40 6e-04
TC17278 similar to UP|Q8GT74 (Q8GT74) NEP1-interacting protein 2... 40 7e-04
BP037094 40 7e-04
TC10409 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (17%) 40 0.001
TC19557 similar to PIR|E86326|E86326 protein F18O14.3 [imported]... 40 0.001
TC14669 similar to PIR|E86326|E86326 protein F18O14.3 [imported]... 39 0.002
TC9277 similar to GB|AAP13413.1|30023760|BT006305 At3g55530 {Ara... 39 0.002
AV413409 39 0.002
BP045442 39 0.002
TC8730 similar to PIR|H96764|H96764 protein RING zinc finger pro... 39 0.002
TC15152 similar to UP|Q8RZY3 (Q8RZY3) RING zinc finger protein-l... 38 0.003
>TC17068 similar to UP|Q8L775 (Q8L775) RING finger-like protein, partial
(9%)
Length = 549
Score = 44.3 bits (103), Expect = 4e-05
Identities = 22/65 (33%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Frame = +2
Query: 300 DNVAGSPISNEWSCVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLC 357
DNV + ++ C IC E++ + +G L CEH + + CIQ+W + + CP+C
Sbjct: 95 DNVTCNENKDDIKCSICQEEYVAADEVGSLKCEHKYHVVCIQQWLRLK------NWCPIC 256
Query: 358 KASFA 362
KA A
Sbjct: 257 KAPVA 271
>TC12569 UP|Q40221 (Q40221) Protein containing C-terminal RING-finger,
complete
Length = 2092
Score = 43.9 bits (102), Expect = 5e-05
Identities = 24/55 (43%), Positives = 35/55 (63%), Gaps = 3/55 (5%)
Frame = +2
Query: 309 NEWSCVICFEDFSST--LGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
+E SCVIC E++ + +G L C H + + CI+KW +SM K+ CP+CKAS
Sbjct: 1772 DEGSCVICLEEYKNMDDVGTLKTCGHDYHVNCIKKW----LSMKKL--CPICKAS 1918
>TC17118 similar to UP|Q94CL1 (Q94CL1) RING finger-like protein
(AT5g19430/F7K24_180), partial (35%)
Length = 708
Score = 43.5 bits (101), Expect = 7e-05
Identities = 30/99 (30%), Positives = 42/99 (42%), Gaps = 4/99 (4%)
Frame = +2
Query: 272 QIEDPQTVEQTSI--DLCSSDGKFIDGDQVDNVAGSPISNEWSCVICFEDF--SSTLGVL 327
QI+D + V T DL D G + G ++ C IC + T V
Sbjct: 212 QIQDEEQVVLTHDFHDLSIEDHLVEKGSDEETHKGGFGNHGGICAICLDKIVLQETALVK 391
Query: 328 PCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKI 366
CEH +C+TCI +WA + TCP CK F + +
Sbjct: 392 GCEHAYCVTCILRWA----TYNNKVTCPQCKNPFEFLNV 496
>TC11796 similar to GB|AAP12870.1|30017273|BT006221 At4g35840 {Arabidopsis
thaliana;}, partial (86%)
Length = 1114
Score = 41.6 bits (96), Expect = 2e-04
Identities = 27/90 (30%), Positives = 38/90 (42%), Gaps = 12/90 (13%)
Frame = +1
Query: 281 QTSIDLCSSDGKFIDGDQVDNVAGSPISNE---------WSCVICFEDFS--STLGVLP- 328
Q D K + GD V+ + I+ + SC +C +DF T+ LP
Sbjct: 487 QNIFDTGCGGAKGLSGDSVEKIPKIKITTDNNADASGERVSCSVCLQDFQLGETVRSLPH 666
Query: 329 CEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
C H F L CI KW + +CPLC+
Sbjct: 667 CHHMFHLPCIDKWLFRH------GSCPLCR 738
>TC12104
Length = 771
Score = 40.8 bits (94), Expect = 4e-04
Identities = 19/48 (39%), Positives = 26/48 (53%)
Frame = +1
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
C IC E S + +L C+H FC C+ +W + TCPLC+AS
Sbjct: 421 CAICQEKMHSPI-LLCCKHMFCEECVSEWFERE------RTCPLCRAS 543
>TC8964 weakly similar to UP|Q94F54 (Q94F54) AT3g15070/K15M2_22, partial
(19%)
Length = 594
Score = 40.8 bits (94), Expect = 4e-04
Identities = 17/50 (34%), Positives = 30/50 (60%), Gaps = 2/50 (4%)
Frame = +1
Query: 312 SCVICFEDFSST--LGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
SC+IC ++F + +G+L CEH + C++KW + + CP+CK+
Sbjct: 193 SCMICQDEFKNQEEIGILRCEHEYHAECLRKWLLVK------NVCPICKS 324
>AU252073
Length = 258
Score = 40.8 bits (94), Expect = 4e-04
Identities = 21/49 (42%), Positives = 28/49 (56%), Gaps = 3/49 (6%)
Frame = +1
Query: 313 CVICFEDFSS--TLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
CVIC D+ L ++P C HTF L+CI W + K STCP+C+
Sbjct: 34 CVICLADYKEREVLRIMPKCGHTFHLSCIDIW------LRKQSTCPVCR 162
>TC19649 weakly similar to UP|RNF6_HUMAN (Q9Y252) RING finger protein 6
(RING-H2 protein), partial (4%)
Length = 523
Score = 40.4 bits (93), Expect = 6e-04
Identities = 27/101 (26%), Positives = 46/101 (44%), Gaps = 5/101 (4%)
Frame = +2
Query: 265 EQITDSNQIEDPQTVEQTSID--LCSSDGKFIDGDQVDNVAGSPISNEWSCVICFEDF-- 320
E + D + +P+ T +D +S K + GD+ + P ++ +C IC ++
Sbjct: 17 EAVADFEPLLNPRRTNNTGLDGPTIASYPKIVIGDEGTRL---PKPDDKACPICLSEYMP 187
Query: 321 SSTLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
T+ +P CEH F CI +W S CP+C+ S
Sbjct: 188 KETVKTIPECEHCFHAQCIDEWLPLNAS------CPICRNS 292
>TC15481 similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-like,
partial (32%)
Length = 706
Score = 40.4 bits (93), Expect = 6e-04
Identities = 21/49 (42%), Positives = 25/49 (50%), Gaps = 3/49 (6%)
Frame = +1
Query: 313 CVICFEDFSS--TLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
C +C +F S T VLP C+H F CI W H STCPLC+
Sbjct: 424 CAVCLSEFESGETGRVLPKCKHAFHTDCIDMWFHSH------STCPLCR 552
>AV767390
Length = 447
Score = 40.4 bits (93), Expect = 6e-04
Identities = 22/67 (32%), Positives = 31/67 (45%), Gaps = 2/67 (2%)
Frame = -3
Query: 313 CVICFEDFSS--TLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHA 370
C IC + L LPC H F C+ KW H +TCPLCK Y I++ +
Sbjct: 430 CCICLSSYDDGVELRELPCGHHFHCGCVDKWLHIN------ATCPLCK-----YNILKSS 284
Query: 371 ATADKEL 377
++E+
Sbjct: 283 NVGEEEV 263
>TC17278 similar to UP|Q8GT74 (Q8GT74) NEP1-interacting protein 2, partial
(62%)
Length = 610
Score = 40.0 bits (92), Expect = 7e-04
Identities = 20/50 (40%), Positives = 27/50 (54%), Gaps = 3/50 (6%)
Frame = +3
Query: 312 SCVICFEDFS--STLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
SC +C +DF T+ LP C H F L CI +W + +S CPLC+
Sbjct: 315 SCSVCLQDFQLGETVRSLPYCHHMFHLPCIDEWLSKHVS------CPLCR 446
>BP037094
Length = 502
Score = 40.0 bits (92), Expect = 7e-04
Identities = 23/62 (37%), Positives = 31/62 (49%), Gaps = 3/62 (4%)
Frame = -3
Query: 300 DNVAGSPISNEWSCVICFEDF--SSTLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPL 356
+N + SP S+ SCVIC +F + LP C H F + CI KW S+CP
Sbjct: 476 NNSSDSPNSSGSSCVICLAEFCDGEQIRFLPKCNHHFHVVCIDKWLLSH------SSCPT 315
Query: 357 CK 358
C+
Sbjct: 314 CR 309
>TC10409 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (17%)
Length = 715
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/46 (43%), Positives = 24/46 (51%)
Frame = +3
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
C+ +ED + L LPC H F TCI KW +TCPLCK
Sbjct: 54 CISSYED-GAELHSLPCNHHFHSTCIVKWLKMN------ATCPLCK 170
>TC19557 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(24%)
Length = 457
Score = 39.7 bits (91), Expect = 0.001
Identities = 18/52 (34%), Positives = 26/52 (49%)
Frame = +2
Query: 308 SNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
+ ++ C ICFE + L C H FC C+ +W H + CP+CKA
Sbjct: 296 AGDFECNICFELAQDPVITL-CGHLFCWPCLYRWLHHH---SQSQECPVCKA 439
>TC14669 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(77%)
Length = 1120
Score = 38.9 bits (89), Expect = 0.002
Identities = 20/60 (33%), Positives = 28/60 (46%)
Frame = +2
Query: 308 SNEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIV 367
+ + C ICF D + + C H FC C+ KW H + CP+CKA K+V
Sbjct: 113 AGNFECNICF-DLAQDPIITLCGHLFCWPCLYKWLHFH---SQSRECPVCKALVEEEKLV 280
>TC9277 similar to GB|AAP13413.1|30023760|BT006305 At3g55530 {Arabidopsis
thaliana;}, partial (79%)
Length = 1297
Score = 38.9 bits (89), Expect = 0.002
Identities = 22/65 (33%), Positives = 31/65 (46%), Gaps = 3/65 (4%)
Frame = +3
Query: 297 DQVDNVAGSPISN-EWSCVICFEDFS--STLGVLPCEHTFCLTCIQKWAHQRISMGKIST 353
D+ + V +S+ E +C +C E + L LPC H F CI W Q+ T
Sbjct: 831 DKSNAVGSMKVSDDELTCSVCLEQVNVGDILRSLPCLHQFHANCIDPWLRQQ------GT 992
Query: 354 CPLCK 358
CP+CK
Sbjct: 993 CPVCK 1007
>AV413409
Length = 374
Score = 38.9 bits (89), Expect = 0.002
Identities = 18/51 (35%), Positives = 29/51 (56%), Gaps = 3/51 (5%)
Frame = +1
Query: 313 CVICFEDF--SSTLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
C +C +F + TL ++P C+H F CI +W + STCP+C+A+
Sbjct: 49 CAVCLNEFEDTETLRLIPKCDHVFHPECIDEW------LSSHSTCPVCRAN 183
>BP045442
Length = 488
Score = 38.5 bits (88), Expect = 0.002
Identities = 24/71 (33%), Positives = 32/71 (44%), Gaps = 10/71 (14%)
Frame = -1
Query: 310 EWSCVICFEDFS--STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK--------A 359
E C +C F S + LPC H F C++KW + I TCPLC+ A
Sbjct: 446 EHECSVCLTKFEPESEINCLPCGHLFHKACLEKW----LDYWNI-TCPLCRTPLMPEDDA 282
Query: 360 SFAWYKIVEHA 370
S W + +HA
Sbjct: 281 SCFW*AMSDHA 249
>TC8730 similar to PIR|H96764|H96764 protein RING zinc finger protein
F25P22.18 [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;}, partial (23%)
Length = 583
Score = 38.5 bits (88), Expect = 0.002
Identities = 19/48 (39%), Positives = 26/48 (53%), Gaps = 2/48 (4%)
Frame = +3
Query: 313 CVICFEDFSST--LGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
C IC E++ S LG L CEH F CI++W + + CP+CK
Sbjct: 207 CTICQEEYESDDELGKLKCEHCFHFQCIKQWLVLK------NFCPVCK 332
>TC15152 similar to UP|Q8RZY3 (Q8RZY3) RING zinc finger protein-like,
partial (11%)
Length = 779
Score = 38.1 bits (87), Expect = 0.003
Identities = 17/49 (34%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = +3
Query: 312 SCVICFEDFS--STLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
+C IC E++ +G L C H++ CIQ+W Q+ + CP+CK
Sbjct: 333 NCTICQEEYEVEEEIGRLNCGHSYHYQCIQQWVEQK------NFCPVCK 461
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,503,035
Number of Sequences: 28460
Number of extensions: 108292
Number of successful extensions: 553
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 543
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 544
length of query: 423
length of database: 4,897,600
effective HSP length: 93
effective length of query: 330
effective length of database: 2,250,820
effective search space: 742770600
effective search space used: 742770600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0225.8


